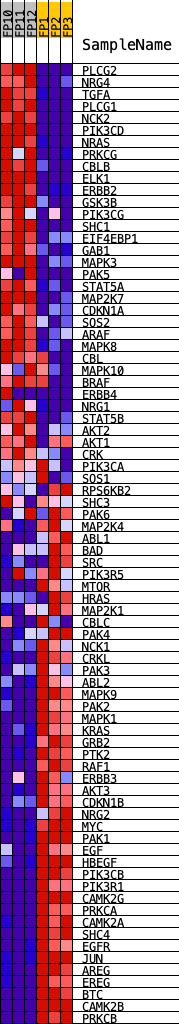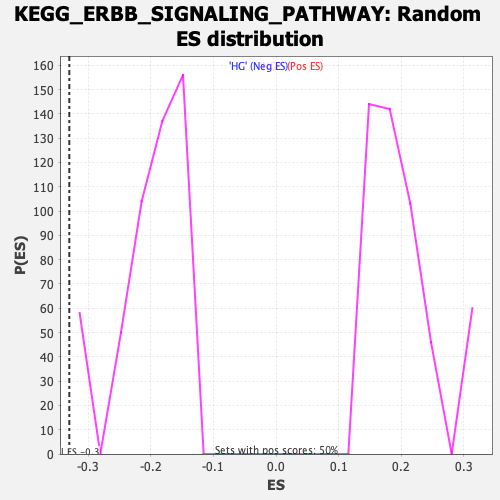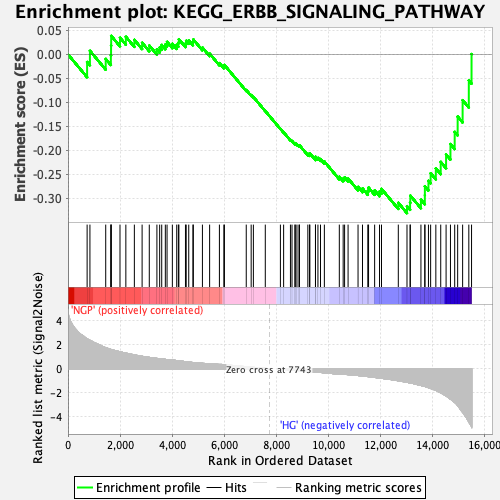
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_HG.phenotype.cls#NGP_versus_HG_repos |
| Phenotype | phenotype.cls#NGP_versus_HG_repos |
| Upregulated in class | HG |
| GeneSet | KEGG_ERBB_SIGNALING_PATHWAY |
| Enrichment Score (ES) | -0.33062512 |
| Normalized Enrichment Score (NES) | -1.6448158 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.081694156 |
| FWER p-Value | 0.211 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PLCG2 | Plcg2 | 735 | 2.492 | -0.0158 | No | ||
| 2 | NRG4 | Nrg4 | 843 | 2.371 | 0.0075 | No | ||
| 3 | TGFA | Tgfa | 1450 | 1.754 | -0.0094 | No | ||
| 4 | PLCG1 | Plcg1 | 1648 | 1.613 | -0.0016 | No | ||
| 5 | NCK2 | Nck2 | 1656 | 1.606 | 0.0184 | No | ||
| 6 | PIK3CD | Pik3cd | 1661 | 1.603 | 0.0386 | No | ||
| 7 | NRAS | Nras | 1999 | 1.405 | 0.0347 | No | ||
| 8 | PRKCG | Prkcg | 2224 | 1.299 | 0.0367 | No | ||
| 9 | CBLB | Cblb | 2551 | 1.153 | 0.0303 | No | ||
| 10 | ELK1 | Elk1 | 2849 | 1.035 | 0.0243 | No | ||
| 11 | ERBB2 | Erbb2 | 3125 | 0.943 | 0.0185 | No | ||
| 12 | GSK3B | Gsk3b | 3418 | 0.860 | 0.0105 | No | ||
| 13 | PIK3CG | Pik3cg | 3524 | 0.831 | 0.0143 | No | ||
| 14 | SHC1 | Shc1 | 3603 | 0.815 | 0.0197 | No | ||
| 15 | EIF4EBP1 | Eif4ebp1 | 3742 | 0.777 | 0.0206 | No | ||
| 16 | GAB1 | Gab1 | 3808 | 0.759 | 0.0261 | No | ||
| 17 | MAPK3 | Mapk3 | 4012 | 0.720 | 0.0221 | No | ||
| 18 | PAK5 | Pak7 | 4180 | 0.678 | 0.0200 | No | ||
| 19 | STAT5A | Stat5a | 4244 | 0.660 | 0.0243 | No | ||
| 20 | MAP2K7 | Map2k7 | 4263 | 0.656 | 0.0315 | No | ||
| 21 | CDKN1A | Cdkn1a | 4516 | 0.587 | 0.0227 | No | ||
| 22 | SOS2 | Sos2 | 4544 | 0.582 | 0.0283 | No | ||
| 23 | ARAF | Araf | 4642 | 0.557 | 0.0292 | No | ||
| 24 | MAPK8 | Mapk8 | 4803 | 0.518 | 0.0254 | No | ||
| 25 | CBL | Cbl | 4818 | 0.516 | 0.0311 | No | ||
| 26 | MAPK10 | Mapk10 | 5168 | 0.449 | 0.0142 | No | ||
| 27 | BRAF | Braf | 5445 | 0.408 | 0.0015 | No | ||
| 28 | ERBB4 | Erbb4 | 5821 | 0.356 | -0.0182 | No | ||
| 29 | NRG1 | Nrg1 | 5992 | 0.338 | -0.0249 | No | ||
| 30 | STAT5B | Stat5b | 6014 | 0.334 | -0.0220 | No | ||
| 31 | AKT2 | Akt2 | 6853 | 0.164 | -0.0742 | No | ||
| 32 | AKT1 | Akt1 | 7042 | 0.129 | -0.0847 | No | ||
| 33 | CRK | Crk | 7127 | 0.113 | -0.0887 | No | ||
| 34 | PIK3CA | Pik3ca | 7585 | 0.030 | -0.1179 | No | ||
| 35 | SOS1 | Sos1 | 8166 | -0.030 | -0.1551 | No | ||
| 36 | RPS6KB2 | Rps6kb2 | 8290 | -0.055 | -0.1623 | No | ||
| 37 | SHC3 | Shc3 | 8552 | -0.102 | -0.1779 | No | ||
| 38 | PAK6 | Pak6 | 8612 | -0.113 | -0.1803 | No | ||
| 39 | MAP2K4 | Map2k4 | 8722 | -0.131 | -0.1857 | No | ||
| 40 | ABL1 | Abl1 | 8739 | -0.135 | -0.1850 | No | ||
| 41 | BAD | Bad | 8802 | -0.146 | -0.1871 | No | ||
| 42 | SRC | Src | 8880 | -0.156 | -0.1901 | No | ||
| 43 | PIK3R5 | Pik3r5 | 8900 | -0.160 | -0.1893 | No | ||
| 44 | MTOR | Mtor | 9221 | -0.219 | -0.2073 | No | ||
| 45 | HRAS | Hras | 9287 | -0.231 | -0.2085 | No | ||
| 46 | MAP2K1 | Map2k1 | 9298 | -0.232 | -0.2062 | No | ||
| 47 | CBLC | Cblc | 9514 | -0.271 | -0.2167 | No | ||
| 48 | PAK4 | Pak4 | 9515 | -0.272 | -0.2132 | No | ||
| 49 | NCK1 | Nck1 | 9609 | -0.291 | -0.2155 | No | ||
| 50 | CRKL | Crkl | 9708 | -0.310 | -0.2179 | No | ||
| 51 | PAK3 | Pak3 | 9856 | -0.340 | -0.2231 | No | ||
| 52 | ABL2 | Abl2 | 10431 | -0.432 | -0.2548 | No | ||
| 53 | MAPK9 | Mapk9 | 10579 | -0.451 | -0.2585 | No | ||
| 54 | PAK2 | Pak2 | 10634 | -0.463 | -0.2561 | No | ||
| 55 | MAPK1 | Mapk1 | 10768 | -0.489 | -0.2585 | No | ||
| 56 | KRAS | Kras | 11151 | -0.567 | -0.2760 | No | ||
| 57 | GRB2 | Grb2 | 11322 | -0.609 | -0.2792 | No | ||
| 58 | PTK2 | Ptk2 | 11527 | -0.655 | -0.2841 | No | ||
| 59 | RAF1 | Raf1 | 11555 | -0.662 | -0.2774 | No | ||
| 60 | ERBB3 | Erbb3 | 11789 | -0.724 | -0.2833 | No | ||
| 61 | AKT3 | Akt3 | 11977 | -0.777 | -0.2855 | No | ||
| 62 | CDKN1B | Cdkn1b | 12053 | -0.801 | -0.2801 | No | ||
| 63 | NRG2 | Nrg2 | 12699 | -1.006 | -0.3091 | No | ||
| 64 | MYC | Myc | 13033 | -1.135 | -0.3162 | Yes | ||
| 65 | PAK1 | Pak1 | 13152 | -1.182 | -0.3087 | Yes | ||
| 66 | EGF | Egf | 13161 | -1.188 | -0.2941 | Yes | ||
| 67 | HBEGF | Hbegf | 13569 | -1.400 | -0.3026 | Yes | ||
| 68 | PIK3CB | Pik3cb | 13719 | -1.479 | -0.2934 | Yes | ||
| 69 | PIK3R1 | Pik3r1 | 13721 | -1.480 | -0.2746 | Yes | ||
| 70 | CAMK2G | Camk2g | 13858 | -1.580 | -0.2633 | Yes | ||
| 71 | PRKCA | Prkca | 13942 | -1.648 | -0.2476 | Yes | ||
| 72 | CAMK2A | Camk2a | 14142 | -1.811 | -0.2375 | Yes | ||
| 73 | SHC4 | Shc4 | 14330 | -2.013 | -0.2239 | Yes | ||
| 74 | EGFR | Egfr | 14536 | -2.270 | -0.2082 | Yes | ||
| 75 | JUN | Jun | 14704 | -2.538 | -0.1867 | Yes | ||
| 76 | AREG | Areg | 14867 | -2.807 | -0.1614 | Yes | ||
| 77 | EREG | Ereg | 14981 | -3.080 | -0.1295 | Yes | ||
| 78 | BTC | Btc | 15172 | -3.638 | -0.0954 | Yes | ||
| 79 | CAMK2B | Camk2b | 15410 | -4.432 | -0.0543 | Yes | ||
| 80 | PRKCB | Prkcb | 15515 | -4.838 | 0.0006 | Yes |