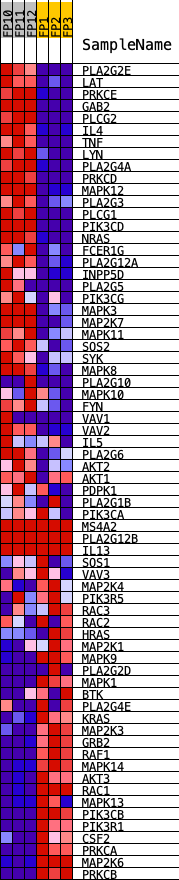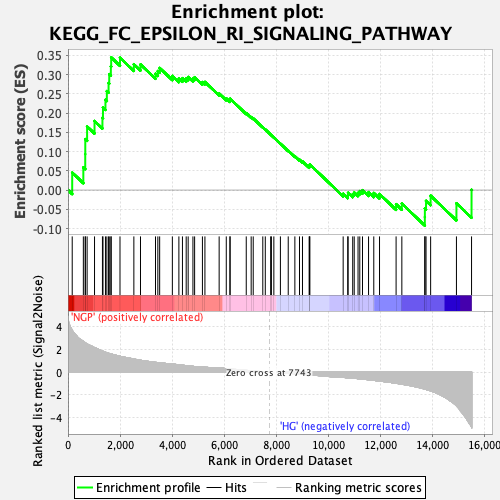
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_HG.phenotype.cls#NGP_versus_HG_repos |
| Phenotype | phenotype.cls#NGP_versus_HG_repos |
| Upregulated in class | NGP |
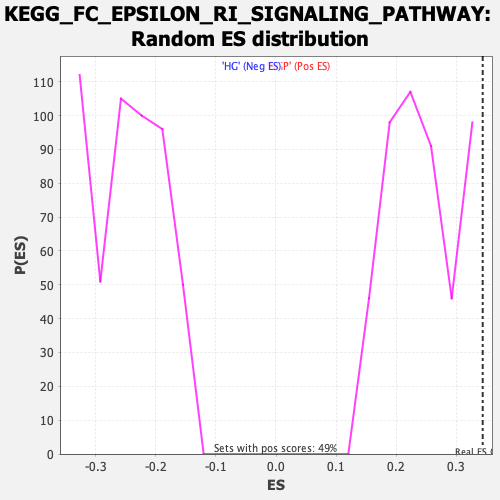
| GeneSet | KEGG_FC_EPSILON_RI_SIGNALING_PATHWAY |
| Enrichment Score (ES) | 0.34436792 |
| Normalized Enrichment Score (NES) | 1.4185051 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.28737456 |
| FWER p-Value | 0.856 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PLA2G2E | Pla2g2e | 162 | 3.725 | 0.0454 | Yes | ||
| 2 | LAT | Lat | 590 | 2.745 | 0.0589 | Yes | ||
| 3 | PRKCE | Prkce | 663 | 2.600 | 0.0933 | Yes | ||
| 4 | GAB2 | Gab2 | 665 | 2.598 | 0.1322 | Yes | ||
| 5 | PLCG2 | Plcg2 | 735 | 2.492 | 0.1651 | Yes | ||
| 6 | IL4 | Il4 | 1020 | 2.171 | 0.1793 | Yes | ||
| 7 | TNF | Tnf | 1324 | 1.864 | 0.1876 | Yes | ||
| 8 | LYN | Lyn | 1345 | 1.850 | 0.2141 | Yes | ||
| 9 | PLA2G4A | Pla2g4a | 1440 | 1.760 | 0.2344 | Yes | ||
| 10 | PRKCD | Prkcd | 1494 | 1.727 | 0.2568 | Yes | ||
| 11 | MAPK12 | Mapk12 | 1560 | 1.674 | 0.2777 | Yes | ||
| 12 | PLA2G3 | Pla2g3 | 1586 | 1.650 | 0.3009 | Yes | ||
| 13 | PLCG1 | Plcg1 | 1648 | 1.613 | 0.3211 | Yes | ||
| 14 | PIK3CD | Pik3cd | 1661 | 1.603 | 0.3444 | Yes | ||
| 15 | NRAS | Nras | 1999 | 1.405 | 0.3436 | No | ||
| 16 | FCER1G | Fcer1g | 2532 | 1.162 | 0.3266 | No | ||
| 17 | PLA2G12A | Pla2g12a | 2787 | 1.061 | 0.3261 | No | ||
| 18 | INPP5D | Inpp5d | 3364 | 0.870 | 0.3019 | No | ||
| 19 | PLA2G5 | Pla2g5 | 3454 | 0.853 | 0.3089 | No | ||
| 20 | PIK3CG | Pik3cg | 3524 | 0.831 | 0.3169 | No | ||
| 21 | MAPK3 | Mapk3 | 4012 | 0.720 | 0.2962 | No | ||
| 22 | MAP2K7 | Map2k7 | 4263 | 0.656 | 0.2899 | No | ||
| 23 | MAPK11 | Mapk11 | 4400 | 0.621 | 0.2904 | No | ||
| 24 | SOS2 | Sos2 | 4544 | 0.582 | 0.2899 | No | ||
| 25 | SYK | Syk | 4620 | 0.565 | 0.2935 | No | ||
| 26 | MAPK8 | Mapk8 | 4803 | 0.518 | 0.2895 | No | ||
| 27 | PLA2G10 | Pla2g10 | 4868 | 0.503 | 0.2929 | No | ||
| 28 | MAPK10 | Mapk10 | 5168 | 0.449 | 0.2803 | No | ||
| 29 | FYN | Fyn | 5264 | 0.436 | 0.2807 | No | ||
| 30 | VAV1 | Vav1 | 5811 | 0.356 | 0.2507 | No | ||
| 31 | VAV2 | Vav2 | 6085 | 0.320 | 0.2378 | No | ||
| 32 | IL5 | Il5 | 6226 | 0.293 | 0.2332 | No | ||
| 33 | PLA2G6 | Pla2g6 | 6232 | 0.291 | 0.2372 | No | ||
| 34 | AKT2 | Akt2 | 6853 | 0.164 | 0.1996 | No | ||
| 35 | AKT1 | Akt1 | 7042 | 0.129 | 0.1893 | No | ||
| 36 | PDPK1 | Pdpk1 | 7121 | 0.114 | 0.1860 | No | ||
| 37 | PLA2G1B | Pla2g1b | 7489 | 0.048 | 0.1630 | No | ||
| 38 | PIK3CA | Pik3ca | 7585 | 0.030 | 0.1573 | No | ||
| 39 | MS4A2 | Ms4a2 | 7792 | 0.000 | 0.1440 | No | ||
| 40 | PLA2G12B | Pla2g12b | 7820 | 0.000 | 0.1422 | No | ||
| 41 | IL13 | Il13 | 7914 | 0.000 | 0.1362 | No | ||
| 42 | SOS1 | Sos1 | 8166 | -0.030 | 0.1204 | No | ||
| 43 | VAV3 | Vav3 | 8467 | -0.087 | 0.1023 | No | ||
| 44 | MAP2K4 | Map2k4 | 8722 | -0.131 | 0.0879 | No | ||
| 45 | PIK3R5 | Pik3r5 | 8900 | -0.160 | 0.0788 | No | ||
| 46 | RAC3 | Rac3 | 9015 | -0.180 | 0.0741 | No | ||
| 47 | RAC2 | Rac2 | 9271 | -0.227 | 0.0610 | No | ||
| 48 | HRAS | Hras | 9287 | -0.231 | 0.0635 | No | ||
| 49 | MAP2K1 | Map2k1 | 9298 | -0.232 | 0.0664 | No | ||
| 50 | MAPK9 | Mapk9 | 10579 | -0.451 | -0.0097 | No | ||
| 51 | PLA2G2D | Pla2g2d | 10756 | -0.489 | -0.0137 | No | ||
| 52 | MAPK1 | Mapk1 | 10768 | -0.489 | -0.0071 | No | ||
| 53 | BTK | Btk | 10946 | -0.521 | -0.0107 | No | ||
| 54 | PLA2G4E | Pla2g4e | 11004 | -0.534 | -0.0064 | No | ||
| 55 | KRAS | Kras | 11151 | -0.567 | -0.0074 | No | ||
| 56 | MAP2K3 | Map2k3 | 11218 | -0.583 | -0.0029 | No | ||
| 57 | GRB2 | Grb2 | 11322 | -0.609 | -0.0004 | No | ||
| 58 | RAF1 | Raf1 | 11555 | -0.662 | -0.0055 | No | ||
| 59 | MAPK14 | Mapk14 | 11756 | -0.714 | -0.0077 | No | ||
| 60 | AKT3 | Akt3 | 11977 | -0.777 | -0.0103 | No | ||
| 61 | RAC1 | Rac1 | 12614 | -0.976 | -0.0368 | No | ||
| 62 | MAPK13 | Mapk13 | 12835 | -1.057 | -0.0352 | No | ||
| 63 | PIK3CB | Pik3cb | 13719 | -1.479 | -0.0702 | No | ||
| 64 | PIK3R1 | Pik3r1 | 13721 | -1.480 | -0.0480 | No | ||
| 65 | CSF2 | Csf2 | 13765 | -1.517 | -0.0280 | No | ||
| 66 | PRKCA | Prkca | 13942 | -1.648 | -0.0147 | No | ||
| 67 | MAP2K6 | Map2k6 | 14934 | -2.963 | -0.0344 | No | ||
| 68 | PRKCB | Prkcb | 15515 | -4.838 | 0.0006 | No |