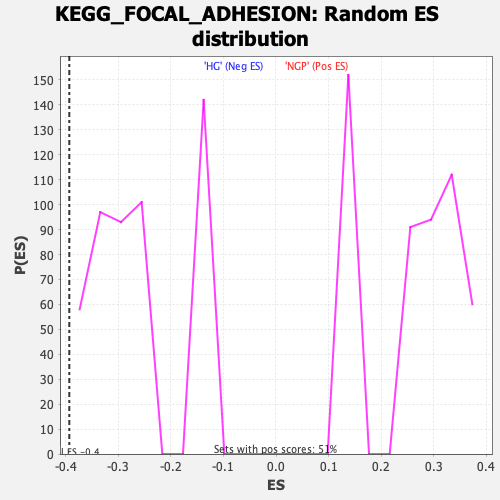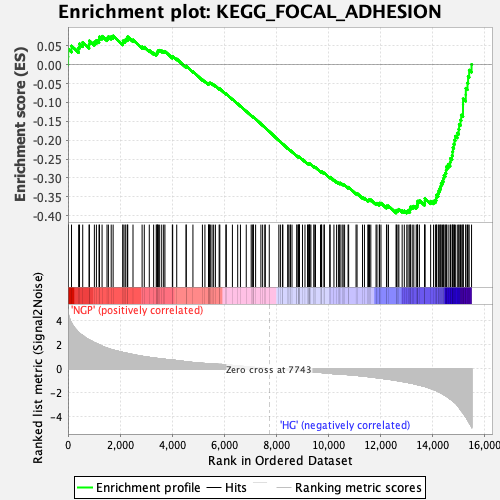
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_HG.phenotype.cls#NGP_versus_HG_repos |
| Phenotype | phenotype.cls#NGP_versus_HG_repos |
| Upregulated in class | HG |
| GeneSet | KEGG_FOCAL_ADHESION |
| Enrichment Score (ES) | -0.3931931 |
| Normalized Enrichment Score (NES) | -1.5387684 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.14743951 |
| FWER p-Value | 0.557 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | VWF | Vwf | 3 | 4.885 | 0.0208 | No | ||
| 2 | KDR | Kdr | 6 | 4.826 | 0.0415 | No | ||
| 3 | COL11A2 | Col11a2 | 135 | 3.829 | 0.0496 | No | ||
| 4 | COL4A4 | Col4a4 | 410 | 3.008 | 0.0446 | No | ||
| 5 | PARVG | Parvg | 443 | 2.954 | 0.0553 | No | ||
| 6 | CHAD | Chad | 566 | 2.774 | 0.0592 | No | ||
| 7 | ITGB7 | Itgb7 | 810 | 2.407 | 0.0538 | No | ||
| 8 | ITGA9 | Itga9 | 822 | 2.396 | 0.0633 | No | ||
| 9 | RELN | Reln | 1014 | 2.179 | 0.0603 | No | ||
| 10 | FLT4 | Flt4 | 1092 | 2.106 | 0.0643 | No | ||
| 11 | LAMB3 | Lamb3 | 1193 | 1.992 | 0.0664 | No | ||
| 12 | LAMA1 | Lama1 | 1210 | 1.976 | 0.0738 | No | ||
| 13 | ITGB5 | Itgb5 | 1310 | 1.874 | 0.0754 | No | ||
| 14 | THBS3 | Thbs3 | 1500 | 1.724 | 0.0705 | No | ||
| 15 | VEGFB | Vegfb | 1553 | 1.680 | 0.0743 | No | ||
| 16 | PIK3CD | Pik3cd | 1661 | 1.603 | 0.0743 | No | ||
| 17 | TNXB | Tnxb | 1735 | 1.558 | 0.0762 | No | ||
| 18 | LAMB2 | Lamb2 | 2107 | 1.356 | 0.0578 | No | ||
| 19 | COMP | Comp | 2116 | 1.353 | 0.0631 | No | ||
| 20 | VEGFC | Vegfc | 2190 | 1.316 | 0.0640 | No | ||
| 21 | PRKCG | Prkcg | 2224 | 1.299 | 0.0675 | No | ||
| 22 | CTNNB1 | Ctnnb1 | 2291 | 1.271 | 0.0686 | No | ||
| 23 | PDGFB | Pdgfb | 2294 | 1.269 | 0.0740 | No | ||
| 24 | ITGB6 | Itgb6 | 2499 | 1.178 | 0.0657 | No | ||
| 25 | ELK1 | Elk1 | 2849 | 1.035 | 0.0474 | No | ||
| 26 | COL2A1 | Col2a1 | 2933 | 1.004 | 0.0463 | No | ||
| 27 | ERBB2 | Erbb2 | 3125 | 0.943 | 0.0379 | No | ||
| 28 | LAMA3 | Lama3 | 3292 | 0.895 | 0.0310 | No | ||
| 29 | BCAR1 | Bcar1 | 3392 | 0.865 | 0.0282 | No | ||
| 30 | GSK3B | Gsk3b | 3418 | 0.860 | 0.0303 | No | ||
| 31 | MYL12B | Myl12b | 3439 | 0.856 | 0.0327 | No | ||
| 32 | ITGA10 | Gm42957 | 3450 | 0.853 | 0.0357 | No | ||
| 33 | FLNA | Flna | 3474 | 0.848 | 0.0378 | No | ||
| 34 | PIK3CG | Pik3cg | 3524 | 0.831 | 0.0382 | No | ||
| 35 | SHC1 | Shc1 | 3603 | 0.815 | 0.0366 | No | ||
| 36 | FN1 | Fn1 | 3678 | 0.794 | 0.0352 | No | ||
| 37 | VEGFA | Vegfa | 3726 | 0.782 | 0.0355 | No | ||
| 38 | MAPK3 | Mapk3 | 4012 | 0.720 | 0.0200 | No | ||
| 39 | PARVB | Parvb | 4025 | 0.717 | 0.0224 | No | ||
| 40 | PAK5 | Pak7 | 4180 | 0.678 | 0.0152 | No | ||
| 41 | ARHGAP35 | Arhgap35 | 4541 | 0.582 | -0.0057 | No | ||
| 42 | SOS2 | Sos2 | 4544 | 0.582 | -0.0034 | No | ||
| 43 | MAPK8 | Mapk8 | 4803 | 0.518 | -0.0180 | No | ||
| 44 | MAPK10 | Mapk10 | 5168 | 0.449 | -0.0398 | No | ||
| 45 | FYN | Fyn | 5264 | 0.436 | -0.0441 | No | ||
| 46 | ITGB8 | Itgb8 | 5402 | 0.417 | -0.0512 | No | ||
| 47 | MYL10 | Myl10 | 5411 | 0.415 | -0.0500 | No | ||
| 48 | DIAPH1 | Diaph1 | 5425 | 0.412 | -0.0491 | No | ||
| 49 | BRAF | Braf | 5445 | 0.408 | -0.0485 | No | ||
| 50 | LAMC3 | Lamc3 | 5471 | 0.404 | -0.0484 | No | ||
| 51 | VTN | Vtn | 5537 | 0.401 | -0.0509 | No | ||
| 52 | ITGA2B | Itga2b | 5599 | 0.396 | -0.0532 | No | ||
| 53 | ITGA3 | Itga3 | 5667 | 0.382 | -0.0559 | No | ||
| 54 | VAV1 | Vav1 | 5811 | 0.356 | -0.0637 | No | ||
| 55 | IBSP | Ibsp | 5837 | 0.356 | -0.0638 | No | ||
| 56 | IGF1R | Igf1r | 6070 | 0.322 | -0.0776 | No | ||
| 57 | VAV2 | Vav2 | 6085 | 0.320 | -0.0771 | No | ||
| 58 | LAMA5 | Lama5 | 6321 | 0.271 | -0.0913 | No | ||
| 59 | LAMB1 | Lamb1 | 6524 | 0.230 | -0.1035 | No | ||
| 60 | PTEN | Pten | 6630 | 0.208 | -0.1094 | No | ||
| 61 | AKT2 | Akt2 | 6853 | 0.164 | -0.1232 | No | ||
| 62 | AKT1 | Akt1 | 7042 | 0.129 | -0.1349 | No | ||
| 63 | ROCK1 | Rock1 | 7085 | 0.119 | -0.1371 | No | ||
| 64 | PDPK1 | Pdpk1 | 7121 | 0.114 | -0.1389 | No | ||
| 65 | CRK | Crk | 7127 | 0.113 | -0.1387 | No | ||
| 66 | CAPN2 | Capn2 | 7213 | 0.097 | -0.1439 | No | ||
| 67 | PXN | Pxn | 7416 | 0.059 | -0.1568 | No | ||
| 68 | CCND3 | Ccnd3 | 7483 | 0.049 | -0.1609 | No | ||
| 69 | PIP5K1C | Pip5k1c | 7567 | 0.035 | -0.1661 | No | ||
| 70 | PIK3CA | Pik3ca | 7585 | 0.030 | -0.1671 | No | ||
| 71 | LAMA2 | Lama2 | 7738 | 0.001 | -0.1770 | No | ||
| 72 | MYLK3 | Mylk3 | 8106 | -0.017 | -0.2009 | No | ||
| 73 | SOS1 | Sos1 | 8166 | -0.030 | -0.2046 | No | ||
| 74 | XIAP | Xiap | 8244 | -0.044 | -0.2094 | No | ||
| 75 | CAV3 | Cav3 | 8261 | -0.048 | -0.2103 | No | ||
| 76 | MYLK2 | Mylk2 | 8447 | -0.083 | -0.2220 | No | ||
| 77 | VAV3 | Vav3 | 8467 | -0.087 | -0.2229 | No | ||
| 78 | MYL2 | Myl2 | 8535 | -0.099 | -0.2268 | No | ||
| 79 | SHC3 | Shc3 | 8552 | -0.102 | -0.2274 | No | ||
| 80 | PAK6 | Pak6 | 8612 | -0.113 | -0.2308 | No | ||
| 81 | ITGA5 | Itga5 | 8800 | -0.145 | -0.2423 | No | ||
| 82 | BAD | Bad | 8802 | -0.146 | -0.2418 | No | ||
| 83 | FLNC | Flnc | 8860 | -0.154 | -0.2448 | No | ||
| 84 | COL5A2 | Col5a2 | 8874 | -0.155 | -0.2450 | No | ||
| 85 | SRC | Src | 8880 | -0.156 | -0.2447 | No | ||
| 86 | VASP | Vasp | 8892 | -0.158 | -0.2447 | No | ||
| 87 | PIK3R5 | Pik3r5 | 8900 | -0.160 | -0.2445 | No | ||
| 88 | RAC3 | Rac3 | 9015 | -0.180 | -0.2511 | No | ||
| 89 | PDGFA | Pdgfa | 9111 | -0.196 | -0.2565 | No | ||
| 90 | ACTN2 | Actn2 | 9209 | -0.215 | -0.2619 | No | ||
| 91 | RHOA | Rhoa | 9229 | -0.220 | -0.2622 | No | ||
| 92 | FLNB | Flnb | 9256 | -0.224 | -0.2629 | No | ||
| 93 | RAC2 | Rac2 | 9271 | -0.227 | -0.2628 | No | ||
| 94 | HRAS | Hras | 9287 | -0.231 | -0.2628 | No | ||
| 95 | MAP2K1 | Map2k1 | 9298 | -0.232 | -0.2625 | No | ||
| 96 | RAPGEF1 | Rapgef1 | 9338 | -0.241 | -0.2640 | No | ||
| 97 | RAP1B | Rap1b | 9457 | -0.264 | -0.2705 | No | ||
| 98 | ACTN1 | Actn1 | 9458 | -0.264 | -0.2694 | No | ||
| 99 | PAK4 | Pak4 | 9515 | -0.272 | -0.2719 | No | ||
| 100 | CRKL | Crkl | 9708 | -0.310 | -0.2831 | No | ||
| 101 | THBS1 | Thbs1 | 9747 | -0.318 | -0.2842 | No | ||
| 102 | PPP1R12A | Ppp1r12a | 9753 | -0.318 | -0.2831 | No | ||
| 103 | PDGFC | Pdgfc | 9842 | -0.336 | -0.2874 | No | ||
| 104 | PAK3 | Pak3 | 9856 | -0.340 | -0.2868 | No | ||
| 105 | COL6A6 | Col6a6 | 10063 | -0.382 | -0.2986 | No | ||
| 106 | ACTB | Actb | 10078 | -0.384 | -0.2979 | No | ||
| 107 | ITGB1 | Itgb1 | 10224 | -0.414 | -0.3056 | No | ||
| 108 | ACTG1 | Actg1 | 10326 | -0.421 | -0.3103 | No | ||
| 109 | ITGA11 | Itga11 | 10396 | -0.426 | -0.3130 | No | ||
| 110 | PPP1CB | Ppp1cb | 10446 | -0.435 | -0.3143 | No | ||
| 111 | BIRC2 | Birc2 | 10458 | -0.437 | -0.3132 | No | ||
| 112 | ROCK2 | Rock2 | 10537 | -0.442 | -0.3163 | No | ||
| 113 | MAPK9 | Mapk9 | 10579 | -0.451 | -0.3171 | No | ||
| 114 | PAK2 | Pak2 | 10634 | -0.463 | -0.3186 | No | ||
| 115 | MAPK1 | Mapk1 | 10768 | -0.489 | -0.3252 | No | ||
| 116 | MYLPF | Mylpf | 10790 | -0.494 | -0.3244 | No | ||
| 117 | CDC42 | Cdc42 | 11074 | -0.550 | -0.3405 | No | ||
| 118 | VCL | Vcl | 11113 | -0.558 | -0.3406 | No | ||
| 119 | GRB2 | Grb2 | 11322 | -0.609 | -0.3515 | No | ||
| 120 | TLN1 | Tln1 | 11397 | -0.628 | -0.3537 | No | ||
| 121 | PTK2 | Ptk2 | 11527 | -0.655 | -0.3592 | No | ||
| 122 | RAF1 | Raf1 | 11555 | -0.662 | -0.3582 | No | ||
| 123 | BCL2 | Bcl2 | 11593 | -0.671 | -0.3577 | No | ||
| 124 | MYL12A | Myl12a | 11642 | -0.682 | -0.3579 | No | ||
| 125 | ITGA7 | Itga7 | 11843 | -0.740 | -0.3677 | No | ||
| 126 | THBS4 | Thbs4 | 11879 | -0.752 | -0.3668 | No | ||
| 127 | RAP1A | Rap1a | 11967 | -0.775 | -0.3691 | No | ||
| 128 | AKT3 | Akt3 | 11977 | -0.777 | -0.3664 | No | ||
| 129 | ITGAV | Itgav | 12032 | -0.796 | -0.3665 | No | ||
| 130 | LAMC2 | Lamc2 | 12244 | -0.851 | -0.3766 | No | ||
| 131 | PPP1CA | Ppp1ca | 12259 | -0.857 | -0.3738 | No | ||
| 132 | LAMC1 | Lamc1 | 12320 | -0.874 | -0.3740 | No | ||
| 133 | RAC1 | Rac1 | 12614 | -0.976 | -0.3889 | Yes | ||
| 134 | FLT1 | Flt1 | 12627 | -0.980 | -0.3854 | Yes | ||
| 135 | TLN2 | Tln2 | 12685 | -1.001 | -0.3848 | Yes | ||
| 136 | PPP1CC | Ppp1cc | 12721 | -1.015 | -0.3828 | Yes | ||
| 137 | MYL9 | Myl9 | 12847 | -1.060 | -0.3863 | Yes | ||
| 138 | ZYX | Zyx | 12930 | -1.092 | -0.3870 | Yes | ||
| 139 | DOCK1 | Dock1 | 13026 | -1.132 | -0.3883 | Yes | ||
| 140 | ARHGAP5 | Arhgap5 | 13074 | -1.152 | -0.3864 | Yes | ||
| 141 | PGF | Pgf | 13145 | -1.179 | -0.3859 | Yes | ||
| 142 | PAK1 | Pak1 | 13152 | -1.182 | -0.3812 | Yes | ||
| 143 | EGF | Egf | 13161 | -1.188 | -0.3766 | Yes | ||
| 144 | BIRC3 | Birc3 | 13233 | -1.225 | -0.3760 | Yes | ||
| 145 | MYL7 | Myl7 | 13295 | -1.248 | -0.3746 | Yes | ||
| 146 | ACTN4 | Actn4 | 13388 | -1.302 | -0.3750 | Yes | ||
| 147 | COL4A6 | Col4a6 | 13425 | -1.324 | -0.3717 | Yes | ||
| 148 | ITGA8 | Itga8 | 13437 | -1.329 | -0.3667 | Yes | ||
| 149 | COL4A1 | Col4a1 | 13442 | -1.331 | -0.3612 | Yes | ||
| 150 | COL4A2 | Col4a2 | 13510 | -1.361 | -0.3597 | Yes | ||
| 151 | MET | Met | 13710 | -1.474 | -0.3664 | Yes | ||
| 152 | PIK3CB | Pik3cb | 13719 | -1.479 | -0.3605 | Yes | ||
| 153 | PIK3R1 | Pik3r1 | 13721 | -1.480 | -0.3542 | Yes | ||
| 154 | PRKCA | Prkca | 13942 | -1.648 | -0.3615 | Yes | ||
| 155 | COL1A2 | Col1a2 | 14054 | -1.731 | -0.3613 | Yes | ||
| 156 | ITGB4 | Itgb4 | 14125 | -1.795 | -0.3581 | Yes | ||
| 157 | ITGA6 | Itga6 | 14158 | -1.823 | -0.3523 | Yes | ||
| 158 | COL5A3 | Col5a3 | 14168 | -1.835 | -0.3450 | Yes | ||
| 159 | PARVA | Parva | 14233 | -1.907 | -0.3410 | Yes | ||
| 160 | CAV2 | Cav2 | 14255 | -1.925 | -0.3341 | Yes | ||
| 161 | ILK | Ilk | 14293 | -1.971 | -0.3280 | Yes | ||
| 162 | SHC4 | Shc4 | 14330 | -2.013 | -0.3217 | Yes | ||
| 163 | COL11A1 | Col11a1 | 14344 | -2.028 | -0.3138 | Yes | ||
| 164 | ITGA4 | Itga4 | 14396 | -2.091 | -0.3082 | Yes | ||
| 165 | PDGFRA | Pdgfra | 14432 | -2.130 | -0.3013 | Yes | ||
| 166 | PDGFD | Pdgfd | 14454 | -2.166 | -0.2933 | Yes | ||
| 167 | PDGFRB | Pdgfrb | 14505 | -2.226 | -0.2870 | Yes | ||
| 168 | EGFR | Egfr | 14536 | -2.270 | -0.2792 | Yes | ||
| 169 | VEGFD | Vegfd | 14548 | -2.290 | -0.2701 | Yes | ||
| 170 | ITGA1 | Itga1 | 14614 | -2.383 | -0.2641 | Yes | ||
| 171 | ITGA2 | Itga2 | 14691 | -2.524 | -0.2581 | Yes | ||
| 172 | JUN | Jun | 14704 | -2.538 | -0.2480 | Yes | ||
| 173 | ACTN3 | Actn3 | 14766 | -2.627 | -0.2407 | Yes | ||
| 174 | LAMA4 | Lama4 | 14786 | -2.655 | -0.2305 | Yes | ||
| 175 | MYLK | Mylk | 14799 | -2.685 | -0.2197 | Yes | ||
| 176 | CCND1 | Ccnd1 | 14834 | -2.752 | -0.2101 | Yes | ||
| 177 | COL6A3 | Col6a3 | 14860 | -2.803 | -0.1997 | Yes | ||
| 178 | COL5A1 | Col5a1 | 14890 | -2.862 | -0.1893 | Yes | ||
| 179 | CAV1 | Cav1 | 14966 | -3.049 | -0.1810 | Yes | ||
| 180 | COL6A1 | Col6a1 | 15022 | -3.185 | -0.1709 | Yes | ||
| 181 | SPP1 | Spp1 | 15038 | -3.234 | -0.1580 | Yes | ||
| 182 | IGF1 | Igf1 | 15091 | -3.421 | -0.1466 | Yes | ||
| 183 | TNR | Tnr | 15117 | -3.492 | -0.1333 | Yes | ||
| 184 | COL6A2 | Col6a2 | 15186 | -3.673 | -0.1219 | Yes | ||
| 185 | COL1A1 | Col1a1 | 15187 | -3.673 | -0.1061 | Yes | ||
| 186 | RASGRF1 | Rasgrf1 | 15189 | -3.674 | -0.0903 | Yes | ||
| 187 | HGF | Hgf | 15293 | -4.036 | -0.0797 | Yes | ||
| 188 | TNN | Tnn | 15295 | -4.041 | -0.0624 | Yes | ||
| 189 | COL3A1 | Col3a1 | 15359 | -4.249 | -0.0482 | Yes | ||
| 190 | THBS2 | Thbs2 | 15382 | -4.337 | -0.0310 | Yes | ||
| 191 | TNC | Tnc | 15423 | -4.499 | -0.0142 | Yes | ||
| 192 | PRKCB | Prkcb | 15515 | -4.838 | 0.0007 | Yes |