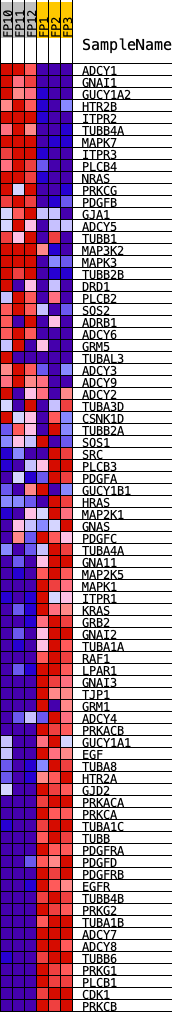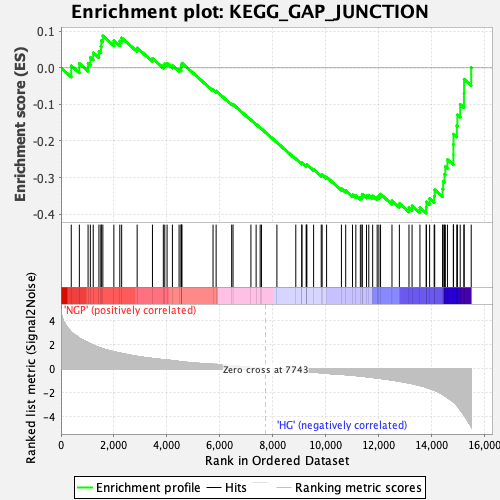
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_HG.phenotype.cls#NGP_versus_HG_repos |
| Phenotype | phenotype.cls#NGP_versus_HG_repos |
| Upregulated in class | HG |
| GeneSet | KEGG_GAP_JUNCTION |
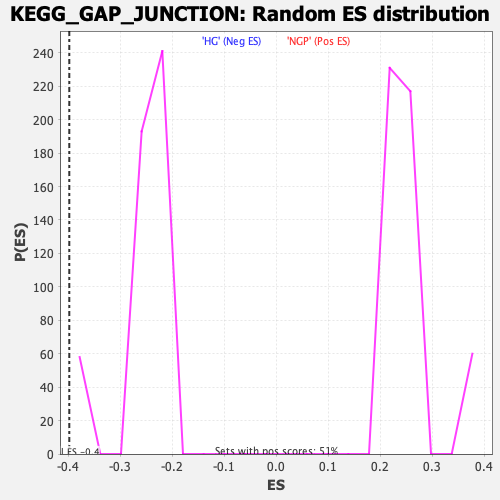
| Enrichment Score (ES) | -0.39754316 |
| Normalized Enrichment Score (NES) | -1.5557219 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.13185835 |
| FWER p-Value | 0.463 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ADCY1 | Adcy1 | 388 | 3.055 | 0.0059 | No | ||
| 2 | GNAI1 | Gnai1 | 694 | 2.552 | 0.0121 | No | ||
| 3 | GUCY1A2 | Gucy1a2 | 1024 | 2.169 | 0.0128 | No | ||
| 4 | HTR2B | Htr2b | 1110 | 2.089 | 0.0285 | No | ||
| 5 | ITPR2 | Itpr2 | 1219 | 1.970 | 0.0415 | No | ||
| 6 | TUBB4A | Tubb4a | 1435 | 1.765 | 0.0455 | No | ||
| 7 | MAPK7 | Mapk7 | 1510 | 1.714 | 0.0581 | No | ||
| 8 | ITPR3 | Itpr3 | 1524 | 1.703 | 0.0745 | No | ||
| 9 | PLCB4 | Plcb4 | 1583 | 1.655 | 0.0876 | No | ||
| 10 | NRAS | Nras | 1999 | 1.405 | 0.0750 | No | ||
| 11 | PRKCG | Prkcg | 2224 | 1.299 | 0.0737 | No | ||
| 12 | PDGFB | Pdgfb | 2294 | 1.269 | 0.0821 | No | ||
| 13 | GJA1 | Gja6 | 2880 | 1.025 | 0.0546 | No | ||
| 14 | ADCY5 | Adcy5 | 3459 | 0.852 | 0.0259 | No | ||
| 15 | TUBB1 | Tubb1 | 3870 | 0.744 | 0.0069 | No | ||
| 16 | MAP3K2 | Map3k2 | 3925 | 0.740 | 0.0109 | No | ||
| 17 | MAPK3 | Mapk3 | 4012 | 0.720 | 0.0126 | No | ||
| 18 | TUBB2B | Tubb2b | 4214 | 0.667 | 0.0064 | No | ||
| 19 | DRD1 | Drd1 | 4466 | 0.601 | -0.0038 | No | ||
| 20 | PLCB2 | Plcb2 | 4543 | 0.582 | -0.0028 | No | ||
| 21 | SOS2 | Sos2 | 4544 | 0.582 | 0.0031 | No | ||
| 22 | ADRB1 | Adrb1 | 4560 | 0.579 | 0.0080 | No | ||
| 23 | ADCY6 | Adcy6 | 4574 | 0.576 | 0.0130 | No | ||
| 24 | GRM5 | Grm5 | 5751 | 0.363 | -0.0594 | No | ||
| 25 | TUBAL3 | Tubal3 | 5867 | 0.356 | -0.0632 | No | ||
| 26 | ADCY3 | Adcy3 | 6455 | 0.245 | -0.0987 | No | ||
| 27 | ADCY9 | Adcy9 | 6506 | 0.234 | -0.0996 | No | ||
| 28 | ADCY2 | Adcy2 | 7183 | 0.102 | -0.1423 | No | ||
| 29 | TUBA3D | Tuba3b | 7380 | 0.064 | -0.1544 | No | ||
| 30 | CSNK1D | Csnk1d | 7533 | 0.040 | -0.1638 | No | ||
| 31 | TUBB2A | Tubb2a | 7582 | 0.031 | -0.1666 | No | ||
| 32 | SOS1 | Sos1 | 8166 | -0.030 | -0.2040 | No | ||
| 33 | SRC | Src | 8880 | -0.156 | -0.2486 | No | ||
| 34 | PLCB3 | Plcb3 | 9108 | -0.196 | -0.2613 | No | ||
| 35 | PDGFA | Pdgfa | 9111 | -0.196 | -0.2595 | No | ||
| 36 | GUCY1B1 | Gucy1b1 | 9272 | -0.227 | -0.2675 | No | ||
| 37 | HRAS | Hras | 9287 | -0.231 | -0.2661 | No | ||
| 38 | MAP2K1 | Map2k1 | 9298 | -0.232 | -0.2644 | No | ||
| 39 | GNAS | Gnas | 9554 | -0.280 | -0.2780 | No | ||
| 40 | PDGFC | Pdgfc | 9842 | -0.336 | -0.2932 | No | ||
| 41 | TUBA4A | Tuba4a | 9876 | -0.345 | -0.2918 | No | ||
| 42 | GNA11 | Gna11 | 10045 | -0.380 | -0.2989 | No | ||
| 43 | MAP2K5 | Map2k5 | 10606 | -0.458 | -0.3305 | No | ||
| 44 | MAPK1 | Mapk1 | 10768 | -0.489 | -0.3359 | No | ||
| 45 | ITPR1 | Itpr1 | 11025 | -0.538 | -0.3470 | No | ||
| 46 | KRAS | Kras | 11151 | -0.567 | -0.3494 | No | ||
| 47 | GRB2 | Grb2 | 11322 | -0.609 | -0.3542 | No | ||
| 48 | GNAI2 | Gnai2 | 11388 | -0.627 | -0.3520 | No | ||
| 49 | TUBA1A | Tuba1a | 11389 | -0.627 | -0.3457 | No | ||
| 50 | RAF1 | Raf1 | 11555 | -0.662 | -0.3496 | No | ||
| 51 | LPAR1 | Lpar1 | 11641 | -0.681 | -0.3482 | No | ||
| 52 | GNAI3 | Gnai3 | 11788 | -0.724 | -0.3503 | No | ||
| 53 | TJP1 | Tjp1 | 11958 | -0.773 | -0.3534 | No | ||
| 54 | GRM1 | Grm1 | 12023 | -0.793 | -0.3495 | No | ||
| 55 | ADCY4 | Adcy4 | 12086 | -0.808 | -0.3453 | No | ||
| 56 | PRKACB | Prkacb | 12518 | -0.946 | -0.3636 | No | ||
| 57 | GUCY1A1 | Gucy1a1 | 12798 | -1.044 | -0.3711 | No | ||
| 58 | EGF | Egf | 13161 | -1.188 | -0.3825 | Yes | ||
| 59 | TUBA8 | Tuba8 | 13278 | -1.240 | -0.3774 | Yes | ||
| 60 | HTR2A | Htr2a | 13576 | -1.403 | -0.3824 | Yes | ||
| 61 | GJD2 | Gjd2 | 13811 | -1.552 | -0.3818 | Yes | ||
| 62 | PRKACA | Prkaca | 13821 | -1.556 | -0.3666 | Yes | ||
| 63 | PRKCA | Prkca | 13942 | -1.648 | -0.3576 | Yes | ||
| 64 | TUBA1C | Tuba1c | 14114 | -1.784 | -0.3506 | Yes | ||
| 65 | TUBB | Tubb5 | 14134 | -1.803 | -0.3335 | Yes | ||
| 66 | PDGFRA | Pdgfra | 14432 | -2.130 | -0.3311 | Yes | ||
| 67 | PDGFD | Pdgfd | 14454 | -2.166 | -0.3105 | Yes | ||
| 68 | PDGFRB | Pdgfrb | 14505 | -2.226 | -0.2911 | Yes | ||
| 69 | EGFR | Egfr | 14536 | -2.270 | -0.2700 | Yes | ||
| 70 | TUBB4B | Tubb4b | 14616 | -2.385 | -0.2509 | Yes | ||
| 71 | PRKG2 | Prkg2 | 14841 | -2.765 | -0.2374 | Yes | ||
| 72 | TUBA1B | Tuba1b | 14842 | -2.768 | -0.2093 | Yes | ||
| 73 | ADCY7 | Adcy7 | 14846 | -2.774 | -0.1813 | Yes | ||
| 74 | ADCY8 | Adcy8 | 14970 | -3.056 | -0.1582 | Yes | ||
| 75 | TUBB6 | Tubb6 | 14993 | -3.101 | -0.1282 | Yes | ||
| 76 | PRKG1 | Prkg1 | 15099 | -3.440 | -0.1001 | Yes | ||
| 77 | PLCB1 | Plcb1 | 15241 | -3.854 | -0.0701 | Yes | ||
| 78 | CDK1 | Cdk1 | 15246 | -3.862 | -0.0311 | Yes | ||
| 79 | PRKCB | Prkcb | 15515 | -4.838 | 0.0006 | Yes |