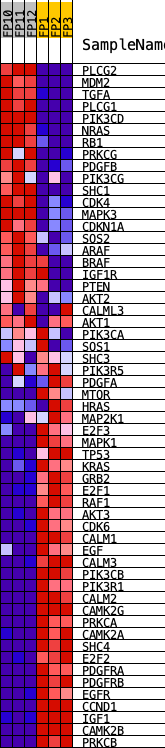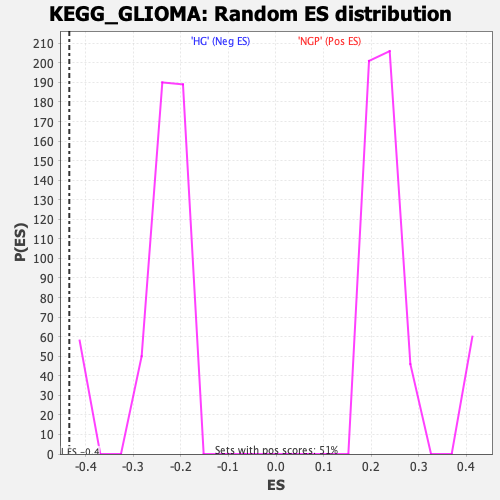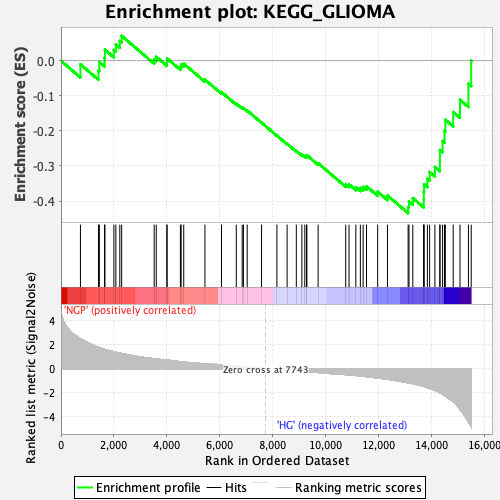
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_HG.phenotype.cls#NGP_versus_HG_repos |
| Phenotype | phenotype.cls#NGP_versus_HG_repos |
| Upregulated in class | HG |
| GeneSet | KEGG_GLIOMA |
| Enrichment Score (ES) | -0.43449438 |
| Normalized Enrichment Score (NES) | -1.7256079 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.06757407 |
| FWER p-Value | 0.105 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PLCG2 | Plcg2 | 735 | 2.492 | -0.0101 | No | ||
| 2 | MDM2 | Mdm2 | 1417 | 1.780 | -0.0274 | No | ||
| 3 | TGFA | Tgfa | 1450 | 1.754 | -0.0032 | No | ||
| 4 | PLCG1 | Plcg1 | 1648 | 1.613 | 0.0083 | No | ||
| 5 | PIK3CD | Pik3cd | 1661 | 1.603 | 0.0316 | No | ||
| 6 | NRAS | Nras | 1999 | 1.405 | 0.0309 | No | ||
| 7 | RB1 | Rb1 | 2074 | 1.372 | 0.0467 | No | ||
| 8 | PRKCG | Prkcg | 2224 | 1.299 | 0.0565 | No | ||
| 9 | PDGFB | Pdgfb | 2294 | 1.269 | 0.0711 | No | ||
| 10 | PIK3CG | Pik3cg | 3524 | 0.831 | 0.0042 | No | ||
| 11 | SHC1 | Shc1 | 3603 | 0.815 | 0.0113 | No | ||
| 12 | CDK4 | Cdk4 | 4004 | 0.722 | -0.0037 | No | ||
| 13 | MAPK3 | Mapk3 | 4012 | 0.720 | 0.0067 | No | ||
| 14 | CDKN1A | Cdkn1a | 4516 | 0.587 | -0.0170 | No | ||
| 15 | SOS2 | Sos2 | 4544 | 0.582 | -0.0100 | No | ||
| 16 | ARAF | Araf | 4642 | 0.557 | -0.0080 | No | ||
| 17 | BRAF | Braf | 5445 | 0.408 | -0.0537 | No | ||
| 18 | IGF1R | Igf1r | 6070 | 0.322 | -0.0892 | No | ||
| 19 | PTEN | Pten | 6630 | 0.208 | -0.1222 | No | ||
| 20 | AKT2 | Akt2 | 6853 | 0.164 | -0.1341 | No | ||
| 21 | CALML3 | Calml3 | 6900 | 0.155 | -0.1347 | No | ||
| 22 | AKT1 | Akt1 | 7042 | 0.129 | -0.1419 | No | ||
| 23 | PIK3CA | Pik3ca | 7585 | 0.030 | -0.1765 | No | ||
| 24 | SOS1 | Sos1 | 8166 | -0.030 | -0.2135 | No | ||
| 25 | SHC3 | Shc3 | 8552 | -0.102 | -0.2369 | No | ||
| 26 | PIK3R5 | Pik3r5 | 8900 | -0.160 | -0.2569 | No | ||
| 27 | PDGFA | Pdgfa | 9111 | -0.196 | -0.2675 | No | ||
| 28 | MTOR | Mtor | 9221 | -0.219 | -0.2713 | No | ||
| 29 | HRAS | Hras | 9287 | -0.231 | -0.2720 | No | ||
| 30 | MAP2K1 | Map2k1 | 9298 | -0.232 | -0.2692 | No | ||
| 31 | E2F3 | E2f3 | 9725 | -0.313 | -0.2921 | No | ||
| 32 | MAPK1 | Mapk1 | 10768 | -0.489 | -0.3521 | No | ||
| 33 | TP53 | Trp53 | 10894 | -0.512 | -0.3525 | No | ||
| 34 | KRAS | Kras | 11151 | -0.567 | -0.3605 | No | ||
| 35 | GRB2 | Grb2 | 11322 | -0.609 | -0.3624 | No | ||
| 36 | E2F1 | E2f1 | 11427 | -0.633 | -0.3596 | No | ||
| 37 | RAF1 | Raf1 | 11555 | -0.662 | -0.3579 | No | ||
| 38 | AKT3 | Akt3 | 11977 | -0.777 | -0.3734 | No | ||
| 39 | CDK6 | Cdk6 | 12347 | -0.884 | -0.3840 | No | ||
| 40 | CALM1 | Calm1 | 13129 | -1.174 | -0.4169 | Yes | ||
| 41 | EGF | Egf | 13161 | -1.188 | -0.4011 | Yes | ||
| 42 | CALM3 | Calm3 | 13306 | -1.252 | -0.3916 | Yes | ||
| 43 | PIK3CB | Pik3cb | 13719 | -1.479 | -0.3960 | Yes | ||
| 44 | PIK3R1 | Pik3r1 | 13721 | -1.480 | -0.3739 | Yes | ||
| 45 | CALM2 | Calm2 | 13732 | -1.487 | -0.3522 | Yes | ||
| 46 | CAMK2G | Camk2g | 13858 | -1.580 | -0.3365 | Yes | ||
| 47 | PRKCA | Prkca | 13942 | -1.648 | -0.3172 | Yes | ||
| 48 | CAMK2A | Camk2a | 14142 | -1.811 | -0.3029 | Yes | ||
| 49 | SHC4 | Shc4 | 14330 | -2.013 | -0.2847 | Yes | ||
| 50 | E2F2 | E2f2 | 14332 | -2.016 | -0.2545 | Yes | ||
| 51 | PDGFRA | Pdgfra | 14432 | -2.130 | -0.2290 | Yes | ||
| 52 | PDGFRB | Pdgfrb | 14505 | -2.226 | -0.2002 | Yes | ||
| 53 | EGFR | Egfr | 14536 | -2.270 | -0.1681 | Yes | ||
| 54 | CCND1 | Ccnd1 | 14834 | -2.752 | -0.1460 | Yes | ||
| 55 | IGF1 | Igf1 | 15091 | -3.421 | -0.1112 | Yes | ||
| 56 | CAMK2B | Camk2b | 15410 | -4.432 | -0.0652 | Yes | ||
| 57 | PRKCB | Prkcb | 15515 | -4.838 | 0.0006 | Yes |