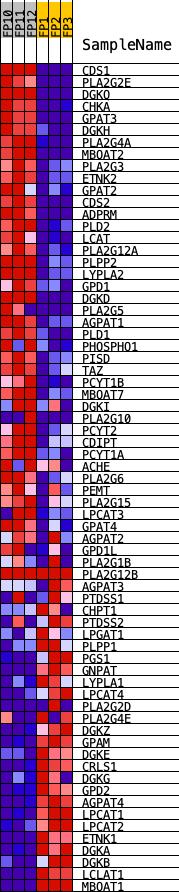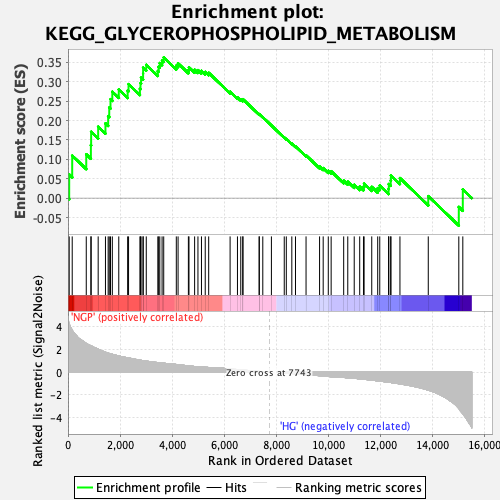
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_HG.phenotype.cls#NGP_versus_HG_repos |
| Phenotype | phenotype.cls#NGP_versus_HG_repos |
| Upregulated in class | NGP |
| GeneSet | KEGG_GLYCEROPHOSPHOLIPID_METABOLISM |
| Enrichment Score (ES) | 0.36288822 |
| Normalized Enrichment Score (NES) | 1.3492635 |
| Nominal p-value | 0.19917865 |
| FDR q-value | 0.2876003 |
| FWER p-Value | 0.899 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CDS1 | Cds1 | 46 | 4.242 | 0.0609 | Yes | ||
| 2 | PLA2G2E | Pla2g2e | 162 | 3.725 | 0.1096 | Yes | ||
| 3 | DGKQ | Dgkq | 702 | 2.535 | 0.1129 | Yes | ||
| 4 | CHKA | Chka | 880 | 2.334 | 0.1367 | Yes | ||
| 5 | GPAT3 | Gpat3 | 890 | 2.323 | 0.1711 | Yes | ||
| 6 | DGKH | Dgkh | 1160 | 2.032 | 0.1843 | Yes | ||
| 7 | PLA2G4A | Pla2g4a | 1440 | 1.760 | 0.1927 | Yes | ||
| 8 | MBOAT2 | Mboat2 | 1544 | 1.687 | 0.2115 | Yes | ||
| 9 | PLA2G3 | Pla2g3 | 1586 | 1.650 | 0.2337 | Yes | ||
| 10 | ETNK2 | Etnk2 | 1634 | 1.620 | 0.2551 | Yes | ||
| 11 | GPAT2 | Gpat2 | 1704 | 1.581 | 0.2744 | Yes | ||
| 12 | CDS2 | Cds2 | 1952 | 1.439 | 0.2801 | Yes | ||
| 13 | ADPRM | Adprm | 2293 | 1.271 | 0.2773 | Yes | ||
| 14 | PLD2 | Pld2 | 2328 | 1.252 | 0.2939 | Yes | ||
| 15 | LCAT | Lcat | 2765 | 1.069 | 0.2818 | Yes | ||
| 16 | PLA2G12A | Pla2g12a | 2787 | 1.061 | 0.2965 | Yes | ||
| 17 | PLPP2 | Plpp2 | 2809 | 1.049 | 0.3109 | Yes | ||
| 18 | LYPLA2 | Lypla2 | 2887 | 1.022 | 0.3213 | Yes | ||
| 19 | GPD1 | Gpd1 | 2889 | 1.021 | 0.3366 | Yes | ||
| 20 | DGKD | Dgkd | 3009 | 0.978 | 0.3437 | Yes | ||
| 21 | PLA2G5 | Pla2g5 | 3454 | 0.853 | 0.3278 | Yes | ||
| 22 | AGPAT1 | Agpat1 | 3485 | 0.846 | 0.3386 | Yes | ||
| 23 | PLD1 | Pld1 | 3529 | 0.830 | 0.3483 | Yes | ||
| 24 | PHOSPHO1 | Phospho1 | 3617 | 0.812 | 0.3549 | Yes | ||
| 25 | PISD | Pisd | 3680 | 0.794 | 0.3629 | Yes | ||
| 26 | TAZ | Taz | 4164 | 0.681 | 0.3419 | No | ||
| 27 | PCYT1B | Pcyt1b | 4232 | 0.662 | 0.3475 | No | ||
| 28 | MBOAT7 | Mboat7 | 4628 | 0.562 | 0.3305 | No | ||
| 29 | DGKI | Dgki | 4651 | 0.553 | 0.3374 | No | ||
| 30 | PLA2G10 | Pla2g10 | 4868 | 0.503 | 0.3310 | No | ||
| 31 | PCYT2 | Pcyt2 | 4996 | 0.481 | 0.3300 | No | ||
| 32 | CDIPT | Cdipt | 5131 | 0.457 | 0.3282 | No | ||
| 33 | PCYT1A | Pcyt1a | 5276 | 0.433 | 0.3254 | No | ||
| 34 | ACHE | Ache | 5409 | 0.415 | 0.3231 | No | ||
| 35 | PLA2G6 | Pla2g6 | 6232 | 0.291 | 0.2744 | No | ||
| 36 | PEMT | Pemt | 6515 | 0.232 | 0.2596 | No | ||
| 37 | PLA2G15 | Pla2g15 | 6642 | 0.207 | 0.2546 | No | ||
| 38 | LPCAT3 | Lpcat3 | 6719 | 0.195 | 0.2526 | No | ||
| 39 | GPAT4 | Gpat4 | 6727 | 0.192 | 0.2550 | No | ||
| 40 | AGPAT2 | Agpat2 | 7349 | 0.069 | 0.2159 | No | ||
| 41 | GPD1L | Gpd1l | 7354 | 0.068 | 0.2167 | No | ||
| 42 | PLA2G1B | Pla2g1b | 7489 | 0.048 | 0.2087 | No | ||
| 43 | PLA2G12B | Pla2g12b | 7820 | 0.000 | 0.1874 | No | ||
| 44 | AGPAT3 | Agpat3 | 8314 | -0.059 | 0.1564 | No | ||
| 45 | PTDSS1 | Ptdss1 | 8393 | -0.073 | 0.1524 | No | ||
| 46 | CHPT1 | Chpt1 | 8605 | -0.112 | 0.1405 | No | ||
| 47 | PTDSS2 | Ptdss2 | 8746 | -0.137 | 0.1335 | No | ||
| 48 | LPGAT1 | Lpgat1 | 9152 | -0.205 | 0.1103 | No | ||
| 49 | PLPP1 | Plpp1 | 9672 | -0.303 | 0.0813 | No | ||
| 50 | PGS1 | Pgs1 | 9811 | -0.331 | 0.0774 | No | ||
| 51 | GNPAT | Gnpat | 10007 | -0.370 | 0.0704 | No | ||
| 52 | LYPLA1 | Lypla1 | 10117 | -0.392 | 0.0692 | No | ||
| 53 | LPCAT4 | Lpcat4 | 10599 | -0.456 | 0.0450 | No | ||
| 54 | PLA2G2D | Pla2g2d | 10756 | -0.489 | 0.0422 | No | ||
| 55 | PLA2G4E | Pla2g4e | 11004 | -0.534 | 0.0343 | No | ||
| 56 | DGKZ | Dgkz | 11215 | -0.582 | 0.0295 | No | ||
| 57 | GPAM | Gpam | 11360 | -0.617 | 0.0295 | No | ||
| 58 | DGKE | Dgke | 11384 | -0.625 | 0.0374 | No | ||
| 59 | CRLS1 | Crls1 | 11678 | -0.693 | 0.0289 | No | ||
| 60 | DGKG | Dgkg | 11907 | -0.757 | 0.0255 | No | ||
| 61 | GPD2 | Gpd2 | 11986 | -0.780 | 0.0322 | No | ||
| 62 | AGPAT4 | Agpat4 | 12325 | -0.875 | 0.0236 | No | ||
| 63 | LPCAT1 | Lpcat1 | 12334 | -0.879 | 0.0363 | No | ||
| 64 | LPCAT2 | Lpcat2 | 12407 | -0.903 | 0.0452 | No | ||
| 65 | ETNK1 | Etnk1 | 12415 | -0.906 | 0.0584 | No | ||
| 66 | DGKA | Dgka | 12762 | -1.030 | 0.0516 | No | ||
| 67 | DGKB | Dgkb | 13851 | -1.578 | 0.0050 | No | ||
| 68 | LCLAT1 | Lclat1 | 15025 | -3.191 | -0.0228 | No | ||
| 69 | MBOAT1 | Mboat1 | 15179 | -3.659 | 0.0224 | No |