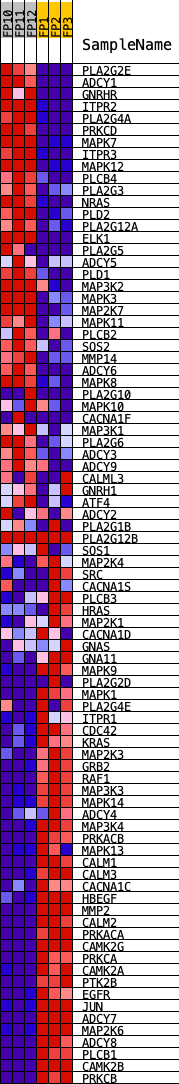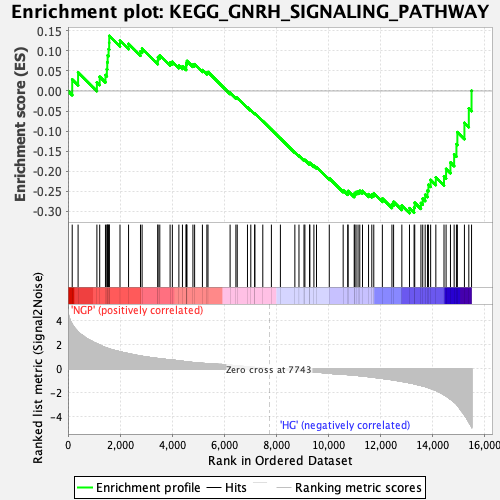
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_HG.phenotype.cls#NGP_versus_HG_repos |
| Phenotype | phenotype.cls#NGP_versus_HG_repos |
| Upregulated in class | HG |
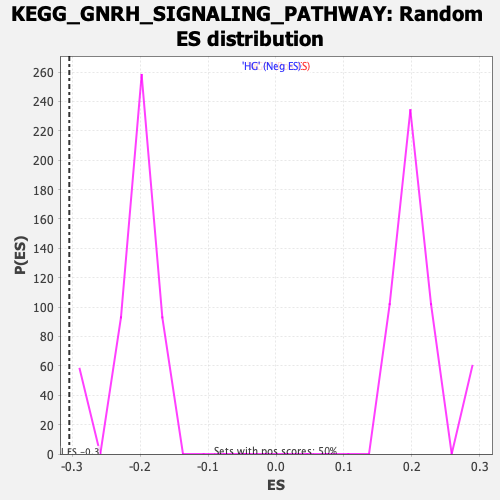
| GeneSet | KEGG_GNRH_SIGNALING_PATHWAY |
| Enrichment Score (ES) | -0.30442905 |
| Normalized Enrichment Score (NES) | -1.480067 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.15819505 |
| FWER p-Value | 0.81 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PLA2G2E | Pla2g2e | 162 | 3.725 | 0.0288 | No | ||
| 2 | ADCY1 | Adcy1 | 388 | 3.055 | 0.0464 | No | ||
| 3 | GNRHR | Gnrhr | 1111 | 2.088 | 0.0216 | No | ||
| 4 | ITPR2 | Itpr2 | 1219 | 1.970 | 0.0355 | No | ||
| 5 | PLA2G4A | Pla2g4a | 1440 | 1.760 | 0.0397 | No | ||
| 6 | PRKCD | Prkcd | 1494 | 1.727 | 0.0545 | No | ||
| 7 | MAPK7 | Mapk7 | 1510 | 1.714 | 0.0716 | No | ||
| 8 | ITPR3 | Itpr3 | 1524 | 1.703 | 0.0887 | No | ||
| 9 | MAPK12 | Mapk12 | 1560 | 1.674 | 0.1041 | No | ||
| 10 | PLCB4 | Plcb4 | 1583 | 1.655 | 0.1201 | No | ||
| 11 | PLA2G3 | Pla2g3 | 1586 | 1.650 | 0.1373 | No | ||
| 12 | NRAS | Nras | 1999 | 1.405 | 0.1255 | No | ||
| 13 | PLD2 | Pld2 | 2328 | 1.252 | 0.1174 | No | ||
| 14 | PLA2G12A | Pla2g12a | 2787 | 1.061 | 0.0989 | No | ||
| 15 | ELK1 | Elk1 | 2849 | 1.035 | 0.1059 | No | ||
| 16 | PLA2G5 | Pla2g5 | 3454 | 0.853 | 0.0758 | No | ||
| 17 | ADCY5 | Adcy5 | 3459 | 0.852 | 0.0845 | No | ||
| 18 | PLD1 | Pld1 | 3529 | 0.830 | 0.0888 | No | ||
| 19 | MAP3K2 | Map3k2 | 3925 | 0.740 | 0.0710 | No | ||
| 20 | MAPK3 | Mapk3 | 4012 | 0.720 | 0.0730 | No | ||
| 21 | MAP2K7 | Map2k7 | 4263 | 0.656 | 0.0637 | No | ||
| 22 | MAPK11 | Mapk11 | 4400 | 0.621 | 0.0614 | No | ||
| 23 | PLCB2 | Plcb2 | 4543 | 0.582 | 0.0584 | No | ||
| 24 | SOS2 | Sos2 | 4544 | 0.582 | 0.0645 | No | ||
| 25 | MMP14 | Mmp14 | 4555 | 0.580 | 0.0700 | No | ||
| 26 | ADCY6 | Adcy6 | 4574 | 0.576 | 0.0749 | No | ||
| 27 | MAPK8 | Mapk8 | 4803 | 0.518 | 0.0656 | No | ||
| 28 | PLA2G10 | Pla2g10 | 4868 | 0.503 | 0.0667 | No | ||
| 29 | MAPK10 | Mapk10 | 5168 | 0.449 | 0.0521 | No | ||
| 30 | CACNA1F | Cacna1f | 5339 | 0.428 | 0.0456 | No | ||
| 31 | MAP3K1 | Map3k1 | 5382 | 0.422 | 0.0473 | No | ||
| 32 | PLA2G6 | Pla2g6 | 6232 | 0.291 | -0.0046 | No | ||
| 33 | ADCY3 | Adcy3 | 6455 | 0.245 | -0.0164 | No | ||
| 34 | ADCY9 | Adcy9 | 6506 | 0.234 | -0.0171 | No | ||
| 35 | CALML3 | Calml3 | 6900 | 0.155 | -0.0410 | No | ||
| 36 | GNRH1 | Gnrh1 | 7028 | 0.131 | -0.0478 | No | ||
| 37 | ATF4 | Atf4 | 7177 | 0.103 | -0.0563 | No | ||
| 38 | ADCY2 | Adcy2 | 7183 | 0.102 | -0.0555 | No | ||
| 39 | PLA2G1B | Pla2g1b | 7489 | 0.048 | -0.0748 | No | ||
| 40 | PLA2G12B | Pla2g12b | 7820 | 0.000 | -0.0962 | No | ||
| 41 | SOS1 | Sos1 | 8166 | -0.030 | -0.1182 | No | ||
| 42 | MAP2K4 | Map2k4 | 8722 | -0.131 | -0.1528 | No | ||
| 43 | SRC | Src | 8880 | -0.156 | -0.1613 | No | ||
| 44 | CACNA1S | Cacna1s | 9073 | -0.191 | -0.1717 | No | ||
| 45 | PLCB3 | Plcb3 | 9108 | -0.196 | -0.1718 | No | ||
| 46 | HRAS | Hras | 9287 | -0.231 | -0.1809 | No | ||
| 47 | MAP2K1 | Map2k1 | 9298 | -0.232 | -0.1791 | No | ||
| 48 | CACNA1D | Cacna1d | 9454 | -0.263 | -0.1864 | No | ||
| 49 | GNAS | Gnas | 9554 | -0.280 | -0.1899 | No | ||
| 50 | GNA11 | Gna11 | 10045 | -0.380 | -0.2176 | No | ||
| 51 | MAPK9 | Mapk9 | 10579 | -0.451 | -0.2474 | No | ||
| 52 | PLA2G2D | Pla2g2d | 10756 | -0.489 | -0.2536 | No | ||
| 53 | MAPK1 | Mapk1 | 10768 | -0.489 | -0.2492 | No | ||
| 54 | PLA2G4E | Pla2g4e | 11004 | -0.534 | -0.2588 | No | ||
| 55 | ITPR1 | Itpr1 | 11025 | -0.538 | -0.2544 | No | ||
| 56 | CDC42 | Cdc42 | 11074 | -0.550 | -0.2517 | No | ||
| 57 | KRAS | Kras | 11151 | -0.567 | -0.2506 | No | ||
| 58 | MAP2K3 | Map2k3 | 11218 | -0.583 | -0.2488 | No | ||
| 59 | GRB2 | Grb2 | 11322 | -0.609 | -0.2490 | No | ||
| 60 | RAF1 | Raf1 | 11555 | -0.662 | -0.2571 | No | ||
| 61 | MAP3K3 | Map3k3 | 11680 | -0.693 | -0.2578 | No | ||
| 62 | MAPK14 | Mapk14 | 11756 | -0.714 | -0.2551 | No | ||
| 63 | ADCY4 | Adcy4 | 12086 | -0.808 | -0.2679 | No | ||
| 64 | MAP3K4 | Map3k4 | 12454 | -0.920 | -0.2820 | No | ||
| 65 | PRKACB | Prkacb | 12518 | -0.946 | -0.2761 | No | ||
| 66 | MAPK13 | Mapk13 | 12835 | -1.057 | -0.2855 | No | ||
| 67 | CALM1 | Calm1 | 13129 | -1.174 | -0.2921 | Yes | ||
| 68 | CALM3 | Calm3 | 13306 | -1.252 | -0.2903 | Yes | ||
| 69 | CACNA1C | Cacna1c | 13331 | -1.267 | -0.2785 | Yes | ||
| 70 | HBEGF | Hbegf | 13569 | -1.400 | -0.2791 | Yes | ||
| 71 | MMP2 | Mmp2 | 13638 | -1.437 | -0.2683 | Yes | ||
| 72 | CALM2 | Calm2 | 13732 | -1.487 | -0.2587 | Yes | ||
| 73 | PRKACA | Prkaca | 13821 | -1.556 | -0.2480 | Yes | ||
| 74 | CAMK2G | Camk2g | 13858 | -1.580 | -0.2336 | Yes | ||
| 75 | PRKCA | Prkca | 13942 | -1.648 | -0.2217 | Yes | ||
| 76 | CAMK2A | Camk2a | 14142 | -1.811 | -0.2155 | Yes | ||
| 77 | PTK2B | Ptk2b | 14456 | -2.167 | -0.2129 | Yes | ||
| 78 | EGFR | Egfr | 14536 | -2.270 | -0.1941 | Yes | ||
| 79 | JUN | Jun | 14704 | -2.538 | -0.1782 | Yes | ||
| 80 | ADCY7 | Adcy7 | 14846 | -2.774 | -0.1581 | Yes | ||
| 81 | MAP2K6 | Map2k6 | 14934 | -2.963 | -0.1325 | Yes | ||
| 82 | ADCY8 | Adcy8 | 14970 | -3.056 | -0.1025 | Yes | ||
| 83 | PLCB1 | Plcb1 | 15241 | -3.854 | -0.0794 | Yes | ||
| 84 | CAMK2B | Camk2b | 15410 | -4.432 | -0.0436 | Yes | ||
| 85 | PRKCB | Prkcb | 15515 | -4.838 | 0.0006 | Yes |