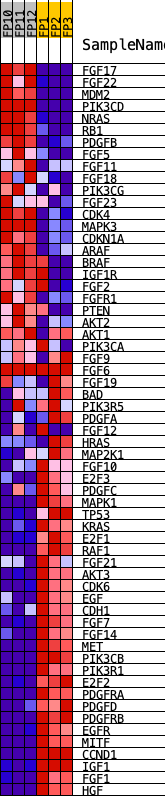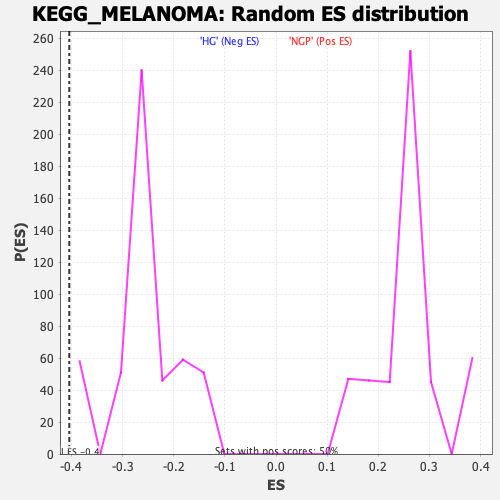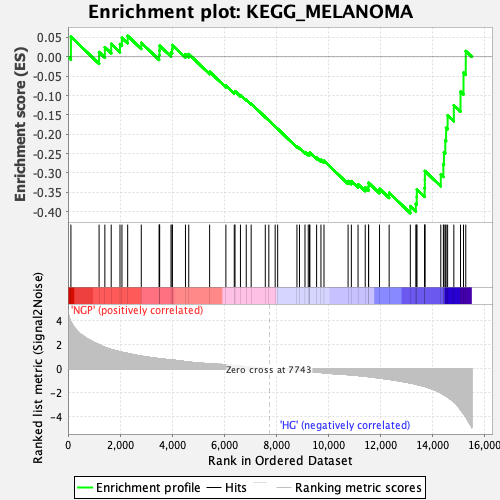
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_HG.phenotype.cls#NGP_versus_HG_repos |
| Phenotype | phenotype.cls#NGP_versus_HG_repos |
| Upregulated in class | HG |
| GeneSet | KEGG_MELANOMA |
| Enrichment Score (ES) | -0.40398303 |
| Normalized Enrichment Score (NES) | -1.5346752 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.14105089 |
| FWER p-Value | 0.557 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | FGF17 | Fgf17 | 112 | 3.899 | 0.0519 | No | ||
| 2 | FGF22 | Fgf22 | 1195 | 1.992 | 0.0122 | No | ||
| 3 | MDM2 | Mdm2 | 1417 | 1.780 | 0.0249 | No | ||
| 4 | PIK3CD | Pik3cd | 1661 | 1.603 | 0.0335 | No | ||
| 5 | NRAS | Nras | 1999 | 1.405 | 0.0330 | No | ||
| 6 | RB1 | Rb1 | 2074 | 1.372 | 0.0491 | No | ||
| 7 | PDGFB | Pdgfb | 2294 | 1.269 | 0.0542 | No | ||
| 8 | FGF5 | Fgf5 | 2817 | 1.046 | 0.0363 | No | ||
| 9 | FGF11 | Fgf11 | 3502 | 0.839 | 0.0048 | No | ||
| 10 | FGF18 | Fgf18 | 3517 | 0.834 | 0.0165 | No | ||
| 11 | PIK3CG | Pik3cg | 3524 | 0.831 | 0.0287 | No | ||
| 12 | FGF23 | Fgf23 | 3960 | 0.732 | 0.0117 | No | ||
| 13 | CDK4 | Cdk4 | 4004 | 0.722 | 0.0199 | No | ||
| 14 | MAPK3 | Mapk3 | 4012 | 0.720 | 0.0304 | No | ||
| 15 | CDKN1A | Cdkn1a | 4516 | 0.587 | 0.0067 | No | ||
| 16 | ARAF | Araf | 4642 | 0.557 | 0.0071 | No | ||
| 17 | BRAF | Braf | 5445 | 0.408 | -0.0386 | No | ||
| 18 | IGF1R | Igf1r | 6070 | 0.322 | -0.0740 | No | ||
| 19 | FGF2 | Fgf2 | 6392 | 0.258 | -0.0908 | No | ||
| 20 | FGFR1 | Fgfr1 | 6421 | 0.253 | -0.0888 | No | ||
| 21 | PTEN | Pten | 6630 | 0.208 | -0.0991 | No | ||
| 22 | AKT2 | Akt2 | 6853 | 0.164 | -0.1110 | No | ||
| 23 | AKT1 | Akt1 | 7042 | 0.129 | -0.1212 | No | ||
| 24 | PIK3CA | Pik3ca | 7585 | 0.030 | -0.1558 | No | ||
| 25 | FGF9 | Fgf9 | 7721 | 0.004 | -0.1644 | No | ||
| 26 | FGF6 | Fgf6 | 7965 | 0.000 | -0.1801 | No | ||
| 27 | FGF19 | Fgf15 | 8061 | -0.009 | -0.1861 | No | ||
| 28 | BAD | Bad | 8802 | -0.146 | -0.2318 | No | ||
| 29 | PIK3R5 | Pik3r5 | 8900 | -0.160 | -0.2356 | No | ||
| 30 | PDGFA | Pdgfa | 9111 | -0.196 | -0.2462 | No | ||
| 31 | FGF12 | Fgf12 | 9237 | -0.221 | -0.2509 | No | ||
| 32 | HRAS | Hras | 9287 | -0.231 | -0.2506 | No | ||
| 33 | MAP2K1 | Map2k1 | 9298 | -0.232 | -0.2477 | No | ||
| 34 | FGF10 | Fgf10 | 9558 | -0.280 | -0.2602 | No | ||
| 35 | E2F3 | E2f3 | 9725 | -0.313 | -0.2662 | No | ||
| 36 | PDGFC | Pdgfc | 9842 | -0.336 | -0.2686 | No | ||
| 37 | MAPK1 | Mapk1 | 10768 | -0.489 | -0.3210 | No | ||
| 38 | TP53 | Trp53 | 10894 | -0.512 | -0.3213 | No | ||
| 39 | KRAS | Kras | 11151 | -0.567 | -0.3293 | No | ||
| 40 | E2F1 | E2f1 | 11427 | -0.633 | -0.3375 | No | ||
| 41 | RAF1 | Raf1 | 11555 | -0.662 | -0.3356 | No | ||
| 42 | FGF21 | Fgf21 | 11557 | -0.662 | -0.3257 | No | ||
| 43 | AKT3 | Akt3 | 11977 | -0.777 | -0.3410 | No | ||
| 44 | CDK6 | Cdk6 | 12347 | -0.884 | -0.3514 | No | ||
| 45 | EGF | Egf | 13161 | -1.188 | -0.3860 | Yes | ||
| 46 | CDH1 | Cdh1 | 13373 | -1.294 | -0.3800 | Yes | ||
| 47 | FGF7 | Fgf7 | 13408 | -1.311 | -0.3623 | Yes | ||
| 48 | FGF14 | Fgf14 | 13412 | -1.313 | -0.3426 | Yes | ||
| 49 | MET | Met | 13710 | -1.474 | -0.3394 | Yes | ||
| 50 | PIK3CB | Pik3cb | 13719 | -1.479 | -0.3175 | Yes | ||
| 51 | PIK3R1 | Pik3r1 | 13721 | -1.480 | -0.2951 | Yes | ||
| 52 | E2F2 | E2f2 | 14332 | -2.016 | -0.3039 | Yes | ||
| 53 | PDGFRA | Pdgfra | 14432 | -2.130 | -0.2780 | Yes | ||
| 54 | PDGFD | Pdgfd | 14454 | -2.166 | -0.2465 | Yes | ||
| 55 | PDGFRB | Pdgfrb | 14505 | -2.226 | -0.2160 | Yes | ||
| 56 | EGFR | Egfr | 14536 | -2.270 | -0.1835 | Yes | ||
| 57 | MITF | Mitf | 14595 | -2.354 | -0.1515 | Yes | ||
| 58 | CCND1 | Ccnd1 | 14834 | -2.752 | -0.1251 | Yes | ||
| 59 | IGF1 | Igf1 | 15091 | -3.421 | -0.0898 | Yes | ||
| 60 | FGF1 | Fgf1 | 15206 | -3.722 | -0.0407 | Yes | ||
| 61 | HGF | Hgf | 15293 | -4.036 | 0.0150 | Yes |