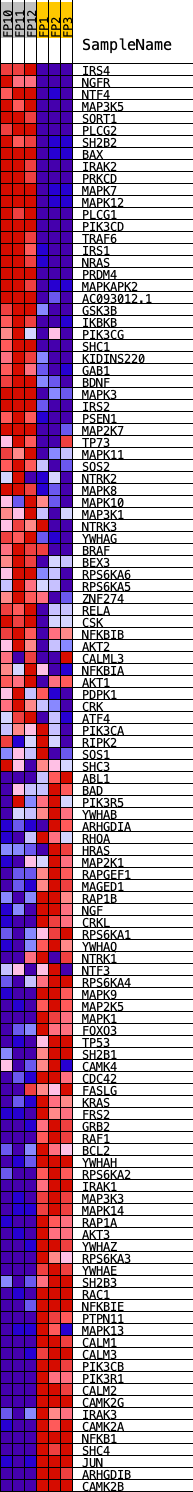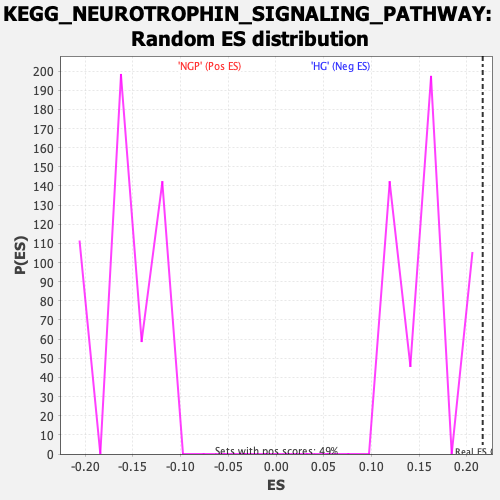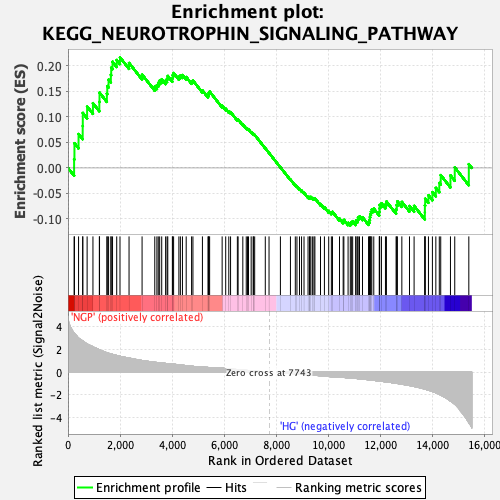
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_HG.phenotype.cls#NGP_versus_HG_repos |
| Phenotype | phenotype.cls#NGP_versus_HG_repos |
| Upregulated in class | NGP |
| GeneSet | KEGG_NEUROTROPHIN_SIGNALING_PATHWAY |
| Enrichment Score (ES) | 0.21657312 |
| Normalized Enrichment Score (NES) | 1.3585553 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.2935603 |
| FWER p-Value | 0.899 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | IRS4 | Irs4 | 235 | 3.470 | 0.0172 | Yes | ||
| 2 | NGFR | Ngfr | 248 | 3.430 | 0.0485 | Yes | ||
| 3 | NTF4 | Ntf5 | 406 | 3.015 | 0.0664 | Yes | ||
| 4 | MAP3K5 | Map3k5 | 565 | 2.774 | 0.0821 | Yes | ||
| 5 | SORT1 | Sort1 | 569 | 2.771 | 0.1078 | Yes | ||
| 6 | PLCG2 | Plcg2 | 735 | 2.492 | 0.1204 | Yes | ||
| 7 | SH2B2 | Sh2b2 | 959 | 2.247 | 0.1269 | Yes | ||
| 8 | BAX | Bax | 1205 | 1.979 | 0.1295 | Yes | ||
| 9 | IRAK2 | Irak2 | 1212 | 1.975 | 0.1476 | Yes | ||
| 10 | PRKCD | Prkcd | 1494 | 1.727 | 0.1455 | Yes | ||
| 11 | MAPK7 | Mapk7 | 1510 | 1.714 | 0.1605 | Yes | ||
| 12 | MAPK12 | Mapk12 | 1560 | 1.674 | 0.1730 | Yes | ||
| 13 | PLCG1 | Plcg1 | 1648 | 1.613 | 0.1824 | Yes | ||
| 14 | PIK3CD | Pik3cd | 1661 | 1.603 | 0.1966 | Yes | ||
| 15 | TRAF6 | Traf6 | 1712 | 1.577 | 0.2081 | Yes | ||
| 16 | IRS1 | Irs1 | 1873 | 1.480 | 0.2116 | Yes | ||
| 17 | NRAS | Nras | 1999 | 1.405 | 0.2166 | Yes | ||
| 18 | PRDM4 | Prdm4 | 2347 | 1.248 | 0.2057 | No | ||
| 19 | MAPKAPK2 | Mapkapk2 | 2848 | 1.035 | 0.1829 | No | ||
| 20 | AC093012.1 | Irak4 | 3336 | 0.879 | 0.1595 | No | ||
| 21 | GSK3B | Gsk3b | 3418 | 0.860 | 0.1623 | No | ||
| 22 | IKBKB | Ikbkb | 3470 | 0.850 | 0.1670 | No | ||
| 23 | PIK3CG | Pik3cg | 3524 | 0.831 | 0.1713 | No | ||
| 24 | SHC1 | Shc1 | 3603 | 0.815 | 0.1738 | No | ||
| 25 | KIDINS220 | Kidins220 | 3753 | 0.775 | 0.1714 | No | ||
| 26 | GAB1 | Gab1 | 3808 | 0.759 | 0.1750 | No | ||
| 27 | BDNF | Bdnf | 3836 | 0.752 | 0.1803 | No | ||
| 28 | MAPK3 | Mapk3 | 4012 | 0.720 | 0.1756 | No | ||
| 29 | IRS2 | Irs2 | 4023 | 0.718 | 0.1817 | No | ||
| 30 | PSEN1 | Psen1 | 4060 | 0.711 | 0.1860 | No | ||
| 31 | MAP2K7 | Map2k7 | 4263 | 0.656 | 0.1790 | No | ||
| 32 | TP73 | Trp73 | 4320 | 0.640 | 0.1814 | No | ||
| 33 | MAPK11 | Mapk11 | 4400 | 0.621 | 0.1820 | No | ||
| 34 | SOS2 | Sos2 | 4544 | 0.582 | 0.1782 | No | ||
| 35 | NTRK2 | Ntrk2 | 4751 | 0.529 | 0.1698 | No | ||
| 36 | MAPK8 | Mapk8 | 4803 | 0.518 | 0.1713 | No | ||
| 37 | MAPK10 | Mapk10 | 5168 | 0.449 | 0.1519 | No | ||
| 38 | MAP3K1 | Map3k1 | 5382 | 0.422 | 0.1420 | No | ||
| 39 | NTRK3 | Ntrk3 | 5403 | 0.417 | 0.1446 | No | ||
| 40 | YWHAG | Ywhag | 5416 | 0.413 | 0.1477 | No | ||
| 41 | BRAF | Braf | 5445 | 0.408 | 0.1497 | No | ||
| 42 | BEX3 | Bex3 | 5926 | 0.353 | 0.1218 | No | ||
| 43 | RPS6KA6 | Rps6ka6 | 6063 | 0.324 | 0.1160 | No | ||
| 44 | RPS6KA5 | Rps6ka5 | 6182 | 0.301 | 0.1112 | No | ||
| 45 | ZNF274 | Zfp110 | 6250 | 0.287 | 0.1095 | No | ||
| 46 | RELA | Rela | 6511 | 0.233 | 0.0948 | No | ||
| 47 | CSK | Csk | 6540 | 0.226 | 0.0951 | No | ||
| 48 | NFKBIB | Nfkbib | 6721 | 0.193 | 0.0852 | No | ||
| 49 | AKT2 | Akt2 | 6853 | 0.164 | 0.0783 | No | ||
| 50 | CALML3 | Calml3 | 6900 | 0.155 | 0.0767 | No | ||
| 51 | NFKBIA | Nfkbia | 6940 | 0.148 | 0.0756 | No | ||
| 52 | AKT1 | Akt1 | 7042 | 0.129 | 0.0702 | No | ||
| 53 | PDPK1 | Pdpk1 | 7121 | 0.114 | 0.0662 | No | ||
| 54 | CRK | Crk | 7127 | 0.113 | 0.0670 | No | ||
| 55 | ATF4 | Atf4 | 7177 | 0.103 | 0.0648 | No | ||
| 56 | PIK3CA | Pik3ca | 7585 | 0.030 | 0.0386 | No | ||
| 57 | RIPK2 | Ripk2 | 7729 | 0.003 | 0.0294 | No | ||
| 58 | SOS1 | Sos1 | 8166 | -0.030 | 0.0014 | No | ||
| 59 | SHC3 | Shc3 | 8552 | -0.102 | -0.0227 | No | ||
| 60 | ABL1 | Abl1 | 8739 | -0.135 | -0.0335 | No | ||
| 61 | BAD | Bad | 8802 | -0.146 | -0.0362 | No | ||
| 62 | PIK3R5 | Pik3r5 | 8900 | -0.160 | -0.0409 | No | ||
| 63 | YWHAB | Ywhab | 8979 | -0.173 | -0.0444 | No | ||
| 64 | ARHGDIA | Arhgdia | 9074 | -0.191 | -0.0487 | No | ||
| 65 | RHOA | Rhoa | 9229 | -0.220 | -0.0567 | No | ||
| 66 | HRAS | Hras | 9287 | -0.231 | -0.0582 | No | ||
| 67 | MAP2K1 | Map2k1 | 9298 | -0.232 | -0.0567 | No | ||
| 68 | RAPGEF1 | Rapgef1 | 9338 | -0.241 | -0.0570 | No | ||
| 69 | MAGED1 | Maged1 | 9405 | -0.254 | -0.0589 | No | ||
| 70 | RAP1B | Rap1b | 9457 | -0.264 | -0.0597 | No | ||
| 71 | NGF | Ngf | 9492 | -0.269 | -0.0594 | No | ||
| 72 | CRKL | Crkl | 9708 | -0.310 | -0.0705 | No | ||
| 73 | RPS6KA1 | Rps6ka1 | 9858 | -0.341 | -0.0770 | No | ||
| 74 | YWHAQ | Ywhaq | 10023 | -0.373 | -0.0841 | No | ||
| 75 | NTRK1 | Ntrk1 | 10125 | -0.394 | -0.0870 | No | ||
| 76 | NTF3 | Ntf3 | 10168 | -0.404 | -0.0859 | No | ||
| 77 | RPS6KA4 | Rps6ka4 | 10438 | -0.434 | -0.0993 | No | ||
| 78 | MAPK9 | Mapk9 | 10579 | -0.451 | -0.1042 | No | ||
| 79 | MAP2K5 | Map2k5 | 10606 | -0.458 | -0.1016 | No | ||
| 80 | MAPK1 | Mapk1 | 10768 | -0.489 | -0.1075 | No | ||
| 81 | FOXO3 | Foxo3 | 10853 | -0.502 | -0.1083 | No | ||
| 82 | TP53 | Trp53 | 10894 | -0.512 | -0.1061 | No | ||
| 83 | SH2B1 | Sh2b1 | 10944 | -0.521 | -0.1044 | No | ||
| 84 | CAMK4 | Camk4 | 11062 | -0.547 | -0.1069 | No | ||
| 85 | CDC42 | Cdc42 | 11074 | -0.550 | -0.1024 | No | ||
| 86 | FASLG | Fasl | 11139 | -0.565 | -0.1013 | No | ||
| 87 | KRAS | Kras | 11151 | -0.567 | -0.0967 | No | ||
| 88 | FRS2 | Frs2 | 11207 | -0.580 | -0.0949 | No | ||
| 89 | GRB2 | Grb2 | 11322 | -0.609 | -0.0966 | No | ||
| 90 | RAF1 | Raf1 | 11555 | -0.662 | -0.1055 | No | ||
| 91 | BCL2 | Bcl2 | 11593 | -0.671 | -0.1016 | No | ||
| 92 | YWHAH | Ywhah | 11608 | -0.674 | -0.0962 | No | ||
| 93 | RPS6KA2 | Rps6ka2 | 11620 | -0.677 | -0.0906 | No | ||
| 94 | IRAK1 | Irak1 | 11647 | -0.683 | -0.0859 | No | ||
| 95 | MAP3K3 | Map3k3 | 11680 | -0.693 | -0.0815 | No | ||
| 96 | MAPK14 | Mapk14 | 11756 | -0.714 | -0.0797 | No | ||
| 97 | RAP1A | Rap1a | 11967 | -0.775 | -0.0861 | No | ||
| 98 | AKT3 | Akt3 | 11977 | -0.777 | -0.0794 | No | ||
| 99 | YWHAZ | Ywhaz | 11982 | -0.778 | -0.0724 | No | ||
| 100 | RPS6KA3 | Rps6ka3 | 12050 | -0.800 | -0.0692 | No | ||
| 101 | YWHAE | Ywhae | 12205 | -0.844 | -0.0714 | No | ||
| 102 | SH2B3 | Sh2b3 | 12242 | -0.851 | -0.0657 | No | ||
| 103 | RAC1 | Rac1 | 12614 | -0.976 | -0.0807 | No | ||
| 104 | NFKBIE | Nfkbie | 12634 | -0.984 | -0.0727 | No | ||
| 105 | PTPN11 | Ptpn11 | 12664 | -0.992 | -0.0653 | No | ||
| 106 | MAPK13 | Mapk13 | 12835 | -1.057 | -0.0665 | No | ||
| 107 | CALM1 | Calm1 | 13129 | -1.174 | -0.0745 | No | ||
| 108 | CALM3 | Calm3 | 13306 | -1.252 | -0.0743 | No | ||
| 109 | PIK3CB | Pik3cb | 13719 | -1.479 | -0.0872 | No | ||
| 110 | PIK3R1 | Pik3r1 | 13721 | -1.480 | -0.0734 | No | ||
| 111 | CALM2 | Calm2 | 13732 | -1.487 | -0.0602 | No | ||
| 112 | CAMK2G | Camk2g | 13858 | -1.580 | -0.0535 | No | ||
| 113 | IRAK3 | Irak3 | 14010 | -1.702 | -0.0474 | No | ||
| 114 | CAMK2A | Camk2a | 14142 | -1.811 | -0.0390 | No | ||
| 115 | NFKB1 | Nfkb1 | 14275 | -1.949 | -0.0293 | No | ||
| 116 | SHC4 | Shc4 | 14330 | -2.013 | -0.0140 | No | ||
| 117 | JUN | Jun | 14704 | -2.538 | -0.0145 | No | ||
| 118 | ARHGDIB | Arhgdib | 14869 | -2.810 | 0.0011 | No | ||
| 119 | CAMK2B | Camk2b | 15410 | -4.432 | 0.0075 | No |