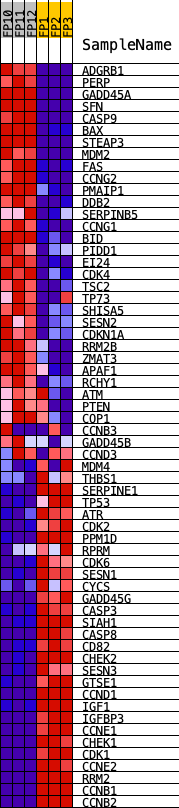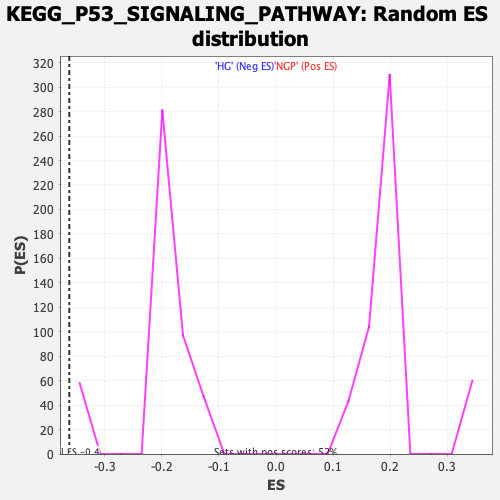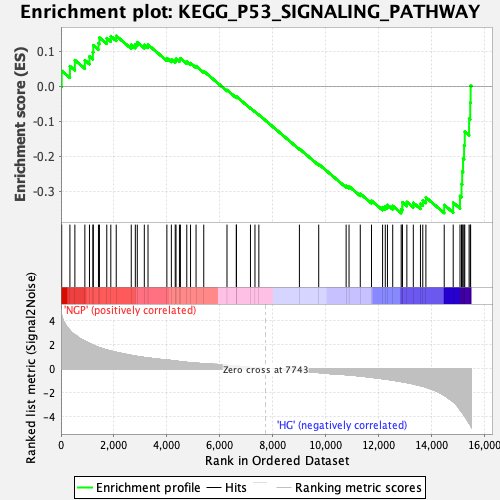
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_HG.phenotype.cls#NGP_versus_HG_repos |
| Phenotype | phenotype.cls#NGP_versus_HG_repos |
| Upregulated in class | HG |
| GeneSet | KEGG_P53_SIGNALING_PATHWAY |
| Enrichment Score (ES) | -0.3623384 |
| Normalized Enrichment Score (NES) | -1.752646 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.058000036 |
| FWER p-Value | 0.058 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ADGRB1 | Adgrb1 | 26 | 4.445 | 0.0446 | No | ||
| 2 | PERP | Perp | 337 | 3.201 | 0.0579 | No | ||
| 3 | GADD45A | Gadd45a | 525 | 2.819 | 0.0752 | No | ||
| 4 | SFN | Sfn | 902 | 2.312 | 0.0749 | No | ||
| 5 | CASP9 | Casp9 | 1077 | 2.120 | 0.0858 | No | ||
| 6 | BAX | Bax | 1205 | 1.979 | 0.0982 | No | ||
| 7 | STEAP3 | Steap3 | 1222 | 1.969 | 0.1176 | No | ||
| 8 | MDM2 | Mdm2 | 1417 | 1.780 | 0.1236 | No | ||
| 9 | FAS | Fas | 1454 | 1.752 | 0.1396 | No | ||
| 10 | CCNG2 | Ccng2 | 1731 | 1.561 | 0.1380 | No | ||
| 11 | PMAIP1 | Pmaip1 | 1884 | 1.476 | 0.1435 | No | ||
| 12 | DDB2 | Ddb2 | 2089 | 1.365 | 0.1445 | No | ||
| 13 | SERPINB5 | Serpinb5 | 2657 | 1.110 | 0.1194 | No | ||
| 14 | CCNG1 | Ccng1 | 2811 | 1.048 | 0.1205 | No | ||
| 15 | BID | Bid | 2885 | 1.022 | 0.1264 | No | ||
| 16 | PIDD1 | Pidd1 | 3151 | 0.935 | 0.1190 | No | ||
| 17 | EI24 | Ei24 | 3291 | 0.895 | 0.1193 | No | ||
| 18 | CDK4 | Cdk4 | 4004 | 0.722 | 0.0808 | No | ||
| 19 | TSC2 | Tsc2 | 4174 | 0.679 | 0.0769 | No | ||
| 20 | TP73 | Trp73 | 4320 | 0.640 | 0.0742 | No | ||
| 21 | SHISA5 | Shisa5 | 4350 | 0.633 | 0.0789 | No | ||
| 22 | SESN2 | Sesn2 | 4480 | 0.596 | 0.0768 | No | ||
| 23 | CDKN1A | Cdkn1a | 4516 | 0.587 | 0.0807 | No | ||
| 24 | RRM2B | Rrm2b | 4757 | 0.528 | 0.0706 | No | ||
| 25 | ZMAT3 | Zmat3 | 4899 | 0.499 | 0.0667 | No | ||
| 26 | APAF1 | Apaf1 | 5108 | 0.461 | 0.0581 | No | ||
| 27 | RCHY1 | Rchy1 | 5399 | 0.418 | 0.0437 | No | ||
| 28 | ATM | Atm | 6278 | 0.282 | -0.0102 | No | ||
| 29 | PTEN | Pten | 6630 | 0.208 | -0.0307 | No | ||
| 30 | COP1 | Cop1 | 6636 | 0.208 | -0.0289 | No | ||
| 31 | CCNB3 | Ccnb3 | 7166 | 0.106 | -0.0620 | No | ||
| 32 | GADD45B | Gadd45b | 7336 | 0.072 | -0.0721 | No | ||
| 33 | CCND3 | Ccnd3 | 7483 | 0.049 | -0.0811 | No | ||
| 34 | MDM4 | Mdm4 | 9017 | -0.180 | -0.1783 | No | ||
| 35 | THBS1 | Thbs1 | 9747 | -0.318 | -0.2222 | No | ||
| 36 | SERPINE1 | Serpine1 | 10784 | -0.493 | -0.2840 | No | ||
| 37 | TP53 | Trp53 | 10894 | -0.512 | -0.2857 | No | ||
| 38 | ATR | Atr | 11319 | -0.608 | -0.3068 | No | ||
| 39 | CDK2 | Cdk2 | 11744 | -0.710 | -0.3268 | No | ||
| 40 | PPM1D | Ppm1d | 12163 | -0.832 | -0.3452 | No | ||
| 41 | RPRM | Rprm | 12265 | -0.858 | -0.3428 | No | ||
| 42 | CDK6 | Cdk6 | 12347 | -0.884 | -0.3388 | No | ||
| 43 | SESN1 | Sesn1 | 12546 | -0.953 | -0.3417 | No | ||
| 44 | CYCS | Cycs | 12866 | -1.070 | -0.3512 | Yes | ||
| 45 | GADD45G | Gadd45g | 12910 | -1.085 | -0.3427 | Yes | ||
| 46 | CASP3 | Casp3 | 12911 | -1.085 | -0.3314 | Yes | ||
| 47 | SIAH1 | Siah1a | 13081 | -1.155 | -0.3303 | Yes | ||
| 48 | CASP8 | Casp8 | 13325 | -1.265 | -0.3328 | Yes | ||
| 49 | CD82 | Cd82 | 13595 | -1.413 | -0.3355 | Yes | ||
| 50 | CHEK2 | Chek2 | 13683 | -1.463 | -0.3259 | Yes | ||
| 51 | SESN3 | Sesn3 | 13801 | -1.546 | -0.3173 | Yes | ||
| 52 | GTSE1 | Gtse1 | 14494 | -2.215 | -0.3390 | Yes | ||
| 53 | CCND1 | Ccnd1 | 14834 | -2.752 | -0.3323 | Yes | ||
| 54 | IGF1 | Igf1 | 15091 | -3.421 | -0.3132 | Yes | ||
| 55 | IGFBP3 | Igfbp3 | 15148 | -3.569 | -0.2796 | Yes | ||
| 56 | CCNE1 | Ccne1 | 15174 | -3.647 | -0.2433 | Yes | ||
| 57 | CHEK1 | Chek1 | 15207 | -3.730 | -0.2065 | Yes | ||
| 58 | CDK1 | Cdk1 | 15246 | -3.862 | -0.1687 | Yes | ||
| 59 | CCNE2 | Ccne2 | 15271 | -3.945 | -0.1292 | Yes | ||
| 60 | RRM2 | Rrm2 | 15435 | -4.557 | -0.0923 | Yes | ||
| 61 | CCNB1 | Ccnb1 | 15478 | -4.669 | -0.0464 | Yes | ||
| 62 | CCNB2 | Ccnb2 | 15493 | -4.736 | 0.0021 | Yes |