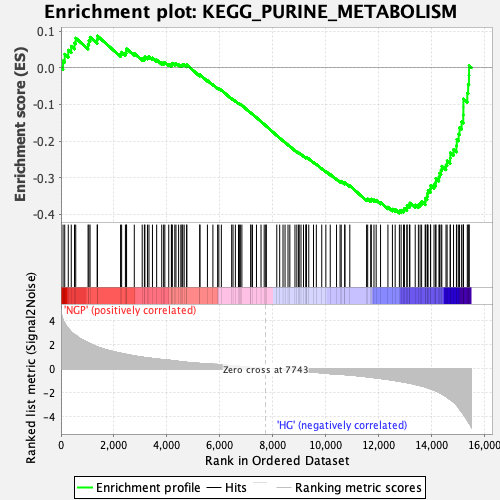
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_HG.phenotype.cls#NGP_versus_HG_repos |
| Phenotype | phenotype.cls#NGP_versus_HG_repos |
| Upregulated in class | HG |
| GeneSet | KEGG_PURINE_METABOLISM |
| Enrichment Score (ES) | -0.39678466 |
| Normalized Enrichment Score (NES) | -1.6479135 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.085078955 |
| FWER p-Value | 0.211 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PDE6B | Pde6b | 78 | 4.079 | 0.0189 | No | ||
| 2 | PDE5A | Pde5a | 141 | 3.802 | 0.0372 | No | ||
| 3 | AMPD3 | Ampd3 | 275 | 3.347 | 0.0482 | No | ||
| 4 | ADCY1 | Adcy1 | 388 | 3.055 | 0.0589 | No | ||
| 5 | NPR1 | Npr1 | 511 | 2.839 | 0.0676 | No | ||
| 6 | PDE3A | Pde3a | 557 | 2.787 | 0.0810 | No | ||
| 7 | GUCY1A2 | Gucy1a2 | 1024 | 2.169 | 0.0635 | No | ||
| 8 | PAPSS2 | Papss2 | 1055 | 2.136 | 0.0741 | No | ||
| 9 | PDE8A | Pde8a | 1099 | 2.096 | 0.0836 | No | ||
| 10 | NPR2 | Npr2 | 1368 | 1.829 | 0.0769 | No | ||
| 11 | AMPD1 | Ampd1 | 1379 | 1.818 | 0.0870 | No | ||
| 12 | ADA | Ada | 2253 | 1.288 | 0.0378 | No | ||
| 13 | ADPRM | Adprm | 2293 | 1.271 | 0.0427 | No | ||
| 14 | POLR3A | Polr3a | 2436 | 1.208 | 0.0406 | No | ||
| 15 | PDE10A | Pde10a | 2466 | 1.193 | 0.0457 | No | ||
| 16 | ALLC | Allc | 2480 | 1.187 | 0.0518 | No | ||
| 17 | NME2 | Nme2 | 2772 | 1.066 | 0.0392 | No | ||
| 18 | PNP | Pnp | 3071 | 0.961 | 0.0254 | No | ||
| 19 | GUCY2C | Gucy2c | 3154 | 0.935 | 0.0256 | No | ||
| 20 | NME4 | Nme4 | 3173 | 0.927 | 0.0299 | No | ||
| 21 | POLR2A | Polr2a | 3270 | 0.899 | 0.0289 | No | ||
| 22 | ATIC | Atic | 3327 | 0.884 | 0.0305 | No | ||
| 23 | ADCY5 | Adcy5 | 3459 | 0.852 | 0.0270 | No | ||
| 24 | PRPS1L1 | Prps1l1 | 3618 | 0.811 | 0.0214 | No | ||
| 25 | XDH | Xdh | 3810 | 0.758 | 0.0135 | No | ||
| 26 | ENTPD3 | Entpd3 | 3882 | 0.741 | 0.0132 | No | ||
| 27 | NME5 | Nme5 | 3927 | 0.740 | 0.0147 | No | ||
| 28 | POLR3F | Polr3f | 4075 | 0.706 | 0.0093 | No | ||
| 29 | AK7 | Ak7 | 4175 | 0.679 | 0.0068 | No | ||
| 30 | ENTPD5 | Entpd5 | 4183 | 0.678 | 0.0104 | No | ||
| 31 | NME6 | Nme6 | 4211 | 0.668 | 0.0125 | No | ||
| 32 | POLR1C | Polr1c | 4304 | 0.644 | 0.0103 | No | ||
| 33 | NT5C | Nt5c | 4348 | 0.634 | 0.0113 | No | ||
| 34 | PRUNE1 | Prune1 | 4448 | 0.606 | 0.0084 | No | ||
| 35 | POLR3D | Polr3d | 4533 | 0.585 | 0.0064 | No | ||
| 36 | ADCY6 | Adcy6 | 4574 | 0.576 | 0.0072 | No | ||
| 37 | PDE9A | Pde9a | 4604 | 0.569 | 0.0086 | No | ||
| 38 | NUDT9 | Nudt9 | 4659 | 0.551 | 0.0083 | No | ||
| 39 | PRPS2 | Prps2 | 4753 | 0.529 | 0.0054 | No | ||
| 40 | RRM2B | Rrm2b | 4757 | 0.528 | 0.0083 | No | ||
| 41 | ENTPD2 | Entpd2 | 5248 | 0.439 | -0.0210 | No | ||
| 42 | ENTPD8 | Entpd8 | 5255 | 0.438 | -0.0188 | No | ||
| 43 | PDE2A | Gm45837 | 5538 | 0.401 | -0.0348 | No | ||
| 44 | POLR2K | Polr2k | 5745 | 0.365 | -0.0460 | No | ||
| 45 | POLE4 | Pole4 | 5924 | 0.353 | -0.0555 | No | ||
| 46 | CANT1 | Cant1 | 5970 | 0.343 | -0.0564 | No | ||
| 47 | PDE6G | Pde6g | 6065 | 0.324 | -0.0606 | No | ||
| 48 | ADCY3 | Adcy3 | 6455 | 0.245 | -0.0845 | No | ||
| 49 | ADCY9 | Adcy9 | 6506 | 0.234 | -0.0863 | No | ||
| 50 | FHIT | Fhit | 6595 | 0.216 | -0.0908 | No | ||
| 51 | NME1 | Nme1 | 6713 | 0.196 | -0.0972 | No | ||
| 52 | POLR3B | Polr3b | 6746 | 0.188 | -0.0982 | No | ||
| 53 | POLR1D | Polr1d | 6783 | 0.180 | -0.0995 | No | ||
| 54 | POLR3C | Polr3c | 6841 | 0.167 | -0.1022 | No | ||
| 55 | AK1 | Ak1 | 7178 | 0.103 | -0.1235 | No | ||
| 56 | ADCY2 | Adcy2 | 7183 | 0.102 | -0.1231 | No | ||
| 57 | POLR2E | Polr2e | 7241 | 0.091 | -0.1263 | No | ||
| 58 | AK5 | Ak5 | 7393 | 0.062 | -0.1357 | No | ||
| 59 | AMPD2 | Ampd2 | 7565 | 0.035 | -0.1467 | No | ||
| 60 | PDE11A | Pde11a | 7685 | 0.010 | -0.1543 | No | ||
| 61 | PKLR | Pklr | 7743 | 0.000 | -0.1580 | No | ||
| 62 | ENTPD4 | Entpd4b | 7766 | 0.000 | -0.1595 | No | ||
| 63 | GUK1 | Guk1 | 8161 | -0.029 | -0.1849 | No | ||
| 64 | PDE1C | Pde1c | 8271 | -0.051 | -0.1917 | No | ||
| 65 | POLR2F | Polr2f | 8397 | -0.073 | -0.1994 | No | ||
| 66 | POLR2G | Polr2g | 8474 | -0.089 | -0.2038 | No | ||
| 67 | IMPDH1 | Impdh1 | 8586 | -0.109 | -0.2104 | No | ||
| 68 | POLR2L | Polr2l | 8655 | -0.121 | -0.2141 | No | ||
| 69 | GUCY2F | Gucy2f | 8849 | -0.153 | -0.2257 | No | ||
| 70 | APRT | Aprt | 8920 | -0.165 | -0.2293 | No | ||
| 71 | PDE6A | Pde6a | 8987 | -0.175 | -0.2326 | No | ||
| 72 | PDE7A | Pde7a | 9008 | -0.179 | -0.2328 | No | ||
| 73 | ADCY10 | Adcy10 | 9077 | -0.192 | -0.2361 | No | ||
| 74 | NUDT2 | Nudt2 | 9172 | -0.209 | -0.2410 | No | ||
| 75 | AK4 | Ak4 | 9261 | -0.225 | -0.2454 | No | ||
| 76 | GUCY1B1 | Gucy1b1 | 9272 | -0.227 | -0.2447 | No | ||
| 77 | POLR2H | Polr2h | 9286 | -0.230 | -0.2442 | No | ||
| 78 | DGUOK | Dguok | 9371 | -0.247 | -0.2482 | No | ||
| 79 | PDE6D | Pde6d | 9550 | -0.279 | -0.2582 | No | ||
| 80 | PDE4A | Pde4a | 9657 | -0.299 | -0.2633 | No | ||
| 81 | POLR2I | Polr2i | 9862 | -0.341 | -0.2746 | No | ||
| 82 | PDE6H | Pde6h | 10017 | -0.372 | -0.2824 | No | ||
| 83 | NT5C2 | Nt5c2 | 10183 | -0.406 | -0.2907 | No | ||
| 84 | PKM | Pkm | 10421 | -0.430 | -0.3036 | No | ||
| 85 | POLR3K | Polr3k | 10555 | -0.447 | -0.3096 | No | ||
| 86 | PDE1B | Pde1b | 10598 | -0.456 | -0.3097 | No | ||
| 87 | GUCY2D | Gucy2e | 10730 | -0.485 | -0.3154 | No | ||
| 88 | NUDT5 | Nudt5 | 10739 | -0.487 | -0.3130 | No | ||
| 89 | PDE4B | Pde4b | 10923 | -0.517 | -0.3219 | No | ||
| 90 | PRPS1 | Prps1 | 11558 | -0.662 | -0.3592 | No | ||
| 91 | HPRT1 | Hprt | 11599 | -0.673 | -0.3578 | No | ||
| 92 | POLR2B | Polr2b | 11715 | -0.703 | -0.3612 | No | ||
| 93 | NT5C3A | Nt5c3 | 11736 | -0.708 | -0.3583 | No | ||
| 94 | NME7 | Nme7 | 11837 | -0.738 | -0.3605 | No | ||
| 95 | ZNRD1 | Znrd1 | 11920 | -0.760 | -0.3614 | No | ||
| 96 | ADCY4 | Adcy4 | 12086 | -0.808 | -0.3673 | No | ||
| 97 | POLR1E | Polr1e | 12365 | -0.888 | -0.3802 | No | ||
| 98 | POLR1A | Polr1a | 12535 | -0.950 | -0.3856 | No | ||
| 99 | PAICS | Paics | 12639 | -0.985 | -0.3865 | No | ||
| 100 | GUCY1A1 | Gucy1a1 | 12798 | -1.044 | -0.3907 | Yes | ||
| 101 | ENTPD6 | Entpd6 | 12864 | -1.069 | -0.3886 | Yes | ||
| 102 | POLR2C | Polr2c | 12955 | -1.103 | -0.3880 | Yes | ||
| 103 | ADSL | Adsl | 12989 | -1.117 | -0.3836 | Yes | ||
| 104 | GART | Gart | 13085 | -1.156 | -0.3829 | Yes | ||
| 105 | ENTPD1 | Entpd1 | 13088 | -1.158 | -0.3763 | Yes | ||
| 106 | PFAS | Pfas | 13178 | -1.195 | -0.3750 | Yes | ||
| 107 | PDE4D | Pde4d | 13195 | -1.205 | -0.3690 | Yes | ||
| 108 | NT5E | Nt5e | 13397 | -1.305 | -0.3744 | Yes | ||
| 109 | NT5M | Nt5m | 13517 | -1.366 | -0.3741 | Yes | ||
| 110 | PNPT1 | Pnpt1 | 13581 | -1.406 | -0.3699 | Yes | ||
| 111 | GMPS | Gmps | 13639 | -1.438 | -0.3652 | Yes | ||
| 112 | ADSS2 | Adss | 13778 | -1.528 | -0.3652 | Yes | ||
| 113 | IMPDH2 | Impdh2 | 13780 | -1.528 | -0.3563 | Yes | ||
| 114 | POLD1 | Pold1 | 13846 | -1.575 | -0.3513 | Yes | ||
| 115 | POLR1B | Polr1b | 13861 | -1.583 | -0.3429 | Yes | ||
| 116 | ADK | Adk | 13883 | -1.600 | -0.3348 | Yes | ||
| 117 | GMPR2 | Gmpr2 | 13974 | -1.671 | -0.3309 | Yes | ||
| 118 | GMPR | Gmpr | 13981 | -1.678 | -0.3214 | Yes | ||
| 119 | POLE3 | Pole3 | 14109 | -1.780 | -0.3192 | Yes | ||
| 120 | POLR3H | Polr3h | 14173 | -1.839 | -0.3125 | Yes | ||
| 121 | POLR2D | Polr2d | 14179 | -1.845 | -0.3020 | Yes | ||
| 122 | ITPA | Itpa | 14295 | -1.974 | -0.2979 | Yes | ||
| 123 | POLD2 | Pold2 | 14317 | -1.999 | -0.2875 | Yes | ||
| 124 | PDE3B | Pde3b | 14373 | -2.067 | -0.2789 | Yes | ||
| 125 | ADSS1 | Adssl1 | 14403 | -2.096 | -0.2685 | Yes | ||
| 126 | POLE2 | Pole2 | 14562 | -2.309 | -0.2652 | Yes | ||
| 127 | PAPSS1 | Papss1 | 14603 | -2.365 | -0.2539 | Yes | ||
| 128 | AK2 | Ak2 | 14719 | -2.556 | -0.2464 | Yes | ||
| 129 | PPAT | Ppat | 14726 | -2.566 | -0.2317 | Yes | ||
| 130 | ADCY7 | Adcy7 | 14846 | -2.774 | -0.2232 | Yes | ||
| 131 | DCK | Dck | 14956 | -3.010 | -0.2126 | Yes | ||
| 132 | ADCY8 | Adcy8 | 14970 | -3.056 | -0.1955 | Yes | ||
| 133 | PRIM2 | Prim2 | 15041 | -3.242 | -0.1810 | Yes | ||
| 134 | PDE7B | Pde7b | 15074 | -3.359 | -0.1633 | Yes | ||
| 135 | RRM1 | Rrm1 | 15152 | -3.581 | -0.1473 | Yes | ||
| 136 | POLE | Pole | 15214 | -3.772 | -0.1291 | Yes | ||
| 137 | POLA1 | Pola1 | 15219 | -3.784 | -0.1071 | Yes | ||
| 138 | GDA | Gda | 15220 | -3.787 | -0.0849 | Yes | ||
| 139 | ENPP3 | Enpp3 | 15367 | -4.299 | -0.0691 | Yes | ||
| 140 | ENPP1 | Enpp1 | 15399 | -4.404 | -0.0453 | Yes | ||
| 141 | PDE1A | Pde1a | 15430 | -4.525 | -0.0207 | Yes | ||
| 142 | RRM2 | Rrm2 | 15435 | -4.557 | 0.0059 | Yes |