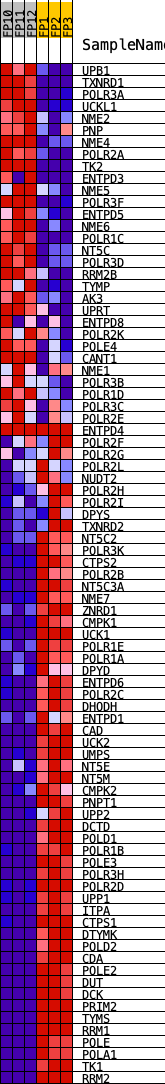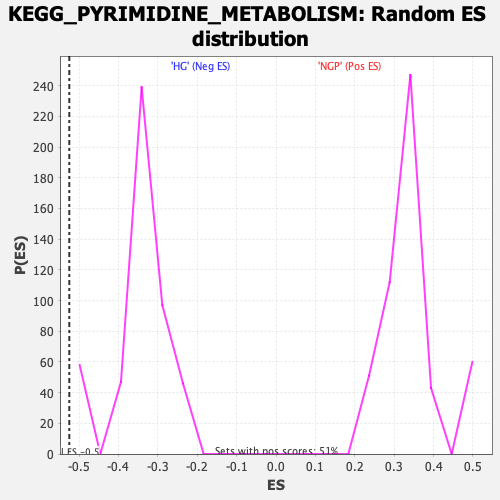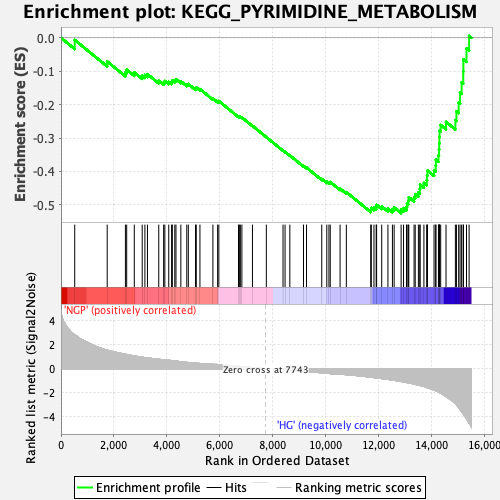
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_HG.phenotype.cls#NGP_versus_HG_repos |
| Phenotype | phenotype.cls#NGP_versus_HG_repos |
| Upregulated in class | HG |
| GeneSet | KEGG_PYRIMIDINE_METABOLISM |
| Enrichment Score (ES) | -0.5247066 |
| Normalized Enrichment Score (NES) | -1.4982964 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.14822891 |
| FWER p-Value | 0.765 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | UPB1 | Upb1 | 519 | 2.824 | -0.0063 | No | ||
| 2 | TXNRD1 | Txnrd1 | 1747 | 1.554 | -0.0708 | No | ||
| 3 | POLR3A | Polr3a | 2436 | 1.208 | -0.1037 | No | ||
| 4 | UCKL1 | Uckl1 | 2484 | 1.186 | -0.0953 | No | ||
| 5 | NME2 | Nme2 | 2772 | 1.066 | -0.1036 | No | ||
| 6 | PNP | Pnp | 3071 | 0.961 | -0.1136 | No | ||
| 7 | NME4 | Nme4 | 3173 | 0.927 | -0.1112 | No | ||
| 8 | POLR2A | Polr2a | 3270 | 0.899 | -0.1087 | No | ||
| 9 | TK2 | Tk2 | 3697 | 0.790 | -0.1287 | No | ||
| 10 | ENTPD3 | Entpd3 | 3882 | 0.741 | -0.1335 | No | ||
| 11 | NME5 | Nme5 | 3927 | 0.740 | -0.1292 | No | ||
| 12 | POLR3F | Polr3f | 4075 | 0.706 | -0.1319 | No | ||
| 13 | ENTPD5 | Entpd5 | 4183 | 0.678 | -0.1322 | No | ||
| 14 | NME6 | Nme6 | 4211 | 0.668 | -0.1275 | No | ||
| 15 | POLR1C | Polr1c | 4304 | 0.644 | -0.1273 | No | ||
| 16 | NT5C | Nt5c | 4348 | 0.634 | -0.1239 | No | ||
| 17 | POLR3D | Polr3d | 4533 | 0.585 | -0.1302 | No | ||
| 18 | RRM2B | Rrm2b | 4757 | 0.528 | -0.1396 | No | ||
| 19 | TYMP | Tymp | 4816 | 0.516 | -0.1383 | No | ||
| 20 | AK3 | Ak3 | 5088 | 0.467 | -0.1514 | No | ||
| 21 | UPRT | Uprt | 5116 | 0.460 | -0.1487 | No | ||
| 22 | ENTPD8 | Entpd8 | 5255 | 0.438 | -0.1534 | No | ||
| 23 | POLR2K | Polr2k | 5745 | 0.365 | -0.1815 | No | ||
| 24 | POLE4 | Pole4 | 5924 | 0.353 | -0.1897 | No | ||
| 25 | CANT1 | Cant1 | 5970 | 0.343 | -0.1892 | No | ||
| 26 | NME1 | Nme1 | 6713 | 0.196 | -0.2354 | No | ||
| 27 | POLR3B | Polr3b | 6746 | 0.188 | -0.2357 | No | ||
| 28 | POLR1D | Polr1d | 6783 | 0.180 | -0.2363 | No | ||
| 29 | POLR3C | Polr3c | 6841 | 0.167 | -0.2383 | No | ||
| 30 | POLR2E | Polr2e | 7241 | 0.091 | -0.2633 | No | ||
| 31 | ENTPD4 | Entpd4b | 7766 | 0.000 | -0.2972 | No | ||
| 32 | POLR2F | Polr2f | 8397 | -0.073 | -0.3373 | No | ||
| 33 | POLR2G | Polr2g | 8474 | -0.089 | -0.3414 | No | ||
| 34 | POLR2L | Polr2l | 8655 | -0.121 | -0.3519 | No | ||
| 35 | NUDT2 | Nudt2 | 9172 | -0.209 | -0.3833 | No | ||
| 36 | POLR2H | Polr2h | 9286 | -0.230 | -0.3884 | No | ||
| 37 | POLR2I | Polr2i | 9862 | -0.341 | -0.4223 | No | ||
| 38 | DPYS | Dpys | 10053 | -0.381 | -0.4310 | No | ||
| 39 | TXNRD2 | Txnrd2 | 10134 | -0.395 | -0.4323 | No | ||
| 40 | NT5C2 | Nt5c2 | 10183 | -0.406 | -0.4315 | No | ||
| 41 | POLR3K | Polr3k | 10555 | -0.447 | -0.4512 | No | ||
| 42 | CTPS2 | Ctps2 | 10792 | -0.495 | -0.4617 | No | ||
| 43 | POLR2B | Polr2b | 11715 | -0.703 | -0.5147 | Yes | ||
| 44 | NT5C3A | Nt5c3 | 11736 | -0.708 | -0.5091 | Yes | ||
| 45 | NME7 | Nme7 | 11837 | -0.738 | -0.5085 | Yes | ||
| 46 | ZNRD1 | Znrd1 | 11920 | -0.760 | -0.5064 | Yes | ||
| 47 | CMPK1 | Cmpk1 | 11937 | -0.768 | -0.5001 | Yes | ||
| 48 | UCK1 | Uck1 | 12130 | -0.821 | -0.5046 | Yes | ||
| 49 | POLR1E | Polr1e | 12365 | -0.888 | -0.5111 | Yes | ||
| 50 | POLR1A | Polr1a | 12535 | -0.950 | -0.5129 | Yes | ||
| 51 | DPYD | Dpyd | 12600 | -0.972 | -0.5077 | Yes | ||
| 52 | ENTPD6 | Entpd6 | 12864 | -1.069 | -0.5144 | Yes | ||
| 53 | POLR2C | Polr2c | 12955 | -1.103 | -0.5096 | Yes | ||
| 54 | DHODH | Dhodh | 13067 | -1.148 | -0.5057 | Yes | ||
| 55 | ENTPD1 | Entpd1 | 13088 | -1.158 | -0.4958 | Yes | ||
| 56 | CAD | Cad | 13135 | -1.176 | -0.4874 | Yes | ||
| 57 | UCK2 | Uck2 | 13154 | -1.182 | -0.4772 | Yes | ||
| 58 | UMPS | Umps | 13354 | -1.281 | -0.4777 | Yes | ||
| 59 | NT5E | Nt5e | 13397 | -1.305 | -0.4678 | Yes | ||
| 60 | NT5M | Nt5m | 13517 | -1.366 | -0.4623 | Yes | ||
| 61 | CMPK2 | Cmpk2 | 13571 | -1.401 | -0.4522 | Yes | ||
| 62 | PNPT1 | Pnpt1 | 13581 | -1.406 | -0.4392 | Yes | ||
| 63 | UPP2 | Upp2 | 13726 | -1.484 | -0.4342 | Yes | ||
| 64 | DCTD | Dctd | 13834 | -1.564 | -0.4261 | Yes | ||
| 65 | POLD1 | Pold1 | 13846 | -1.575 | -0.4116 | Yes | ||
| 66 | POLR1B | Polr1b | 13861 | -1.583 | -0.3972 | Yes | ||
| 67 | POLE3 | Pole3 | 14109 | -1.780 | -0.3960 | Yes | ||
| 68 | POLR3H | Polr3h | 14173 | -1.839 | -0.3823 | Yes | ||
| 69 | POLR2D | Polr2d | 14179 | -1.845 | -0.3648 | Yes | ||
| 70 | UPP1 | Upp1 | 14276 | -1.950 | -0.3522 | Yes | ||
| 71 | ITPA | Itpa | 14295 | -1.974 | -0.3343 | Yes | ||
| 72 | CTPS1 | Ctps | 14305 | -1.984 | -0.3157 | Yes | ||
| 73 | DTYMK | Dtymk | 14312 | -1.993 | -0.2969 | Yes | ||
| 74 | POLD2 | Pold2 | 14317 | -1.999 | -0.2778 | Yes | ||
| 75 | CDA | Cda | 14360 | -2.046 | -0.2608 | Yes | ||
| 76 | POLE2 | Pole2 | 14562 | -2.309 | -0.2515 | Yes | ||
| 77 | DUT | Dut | 14916 | -2.906 | -0.2463 | Yes | ||
| 78 | DCK | Dck | 14956 | -3.010 | -0.2198 | Yes | ||
| 79 | PRIM2 | Prim2 | 15041 | -3.242 | -0.1939 | Yes | ||
| 80 | TYMS | Tyms | 15093 | -3.428 | -0.1641 | Yes | ||
| 81 | RRM1 | Rrm1 | 15152 | -3.581 | -0.1333 | Yes | ||
| 82 | POLE | Pole | 15214 | -3.772 | -0.1008 | Yes | ||
| 83 | POLA1 | Pola1 | 15219 | -3.784 | -0.0646 | Yes | ||
| 84 | TK1 | Tk1 | 15337 | -4.168 | -0.0319 | Yes | ||
| 85 | RRM2 | Rrm2 | 15435 | -4.557 | 0.0058 | Yes |