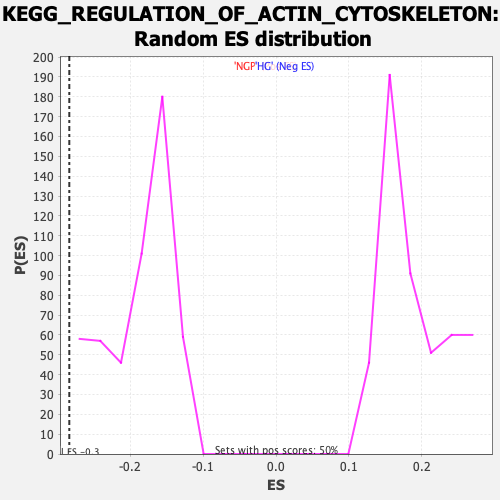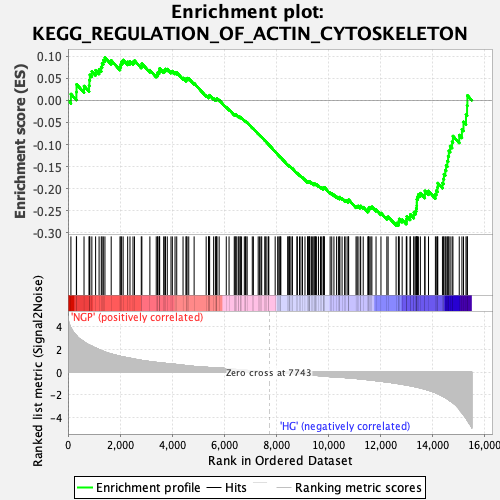
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_HG.phenotype.cls#NGP_versus_HG_repos |
| Phenotype | phenotype.cls#NGP_versus_HG_repos |
| Upregulated in class | HG |
| GeneSet | KEGG_REGULATION_OF_ACTIN_CYTOSKELETON |
| Enrichment Score (ES) | -0.28337482 |
| Normalized Enrichment Score (NES) | -1.5047853 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.15386821 |
| FWER p-Value | 0.765 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | FGF17 | Fgf17 | 112 | 3.899 | 0.0143 | No | ||
| 2 | PIP4K2C | Pip4k2c | 322 | 3.238 | 0.0187 | No | ||
| 3 | CYFIP2 | Cyfip2 | 329 | 3.220 | 0.0361 | No | ||
| 4 | EZR | Ezr | 616 | 2.692 | 0.0324 | No | ||
| 5 | ITGB7 | Itgb7 | 810 | 2.407 | 0.0332 | No | ||
| 6 | ITGA9 | Itga9 | 822 | 2.396 | 0.0458 | No | ||
| 7 | PIP5K1B | Pip5k1b | 841 | 2.373 | 0.0578 | No | ||
| 8 | GIT1 | Git1 | 917 | 2.296 | 0.0656 | No | ||
| 9 | SSH3 | Ssh3 | 1061 | 2.131 | 0.0681 | No | ||
| 10 | FGF22 | Fgf22 | 1195 | 1.992 | 0.0705 | No | ||
| 11 | CHRM1 | Chrm1 | 1279 | 1.910 | 0.0757 | No | ||
| 12 | ITGB5 | Itgb5 | 1310 | 1.874 | 0.0841 | No | ||
| 13 | FGFR3 | Fgfr3 | 1358 | 1.838 | 0.0912 | No | ||
| 14 | PIKFYVE | Pikfyve | 1420 | 1.778 | 0.0971 | No | ||
| 15 | PIK3CD | Pik3cd | 1661 | 1.603 | 0.0904 | No | ||
| 16 | NRAS | Nras | 1999 | 1.405 | 0.0762 | No | ||
| 17 | FGFR2 | Fgfr2 | 2031 | 1.389 | 0.0819 | No | ||
| 18 | ARHGEF6 | Arhgef6 | 2061 | 1.378 | 0.0876 | No | ||
| 19 | ENAH | Enah | 2120 | 1.350 | 0.0913 | No | ||
| 20 | PDGFB | Pdgfb | 2294 | 1.269 | 0.0871 | No | ||
| 21 | GNA13 | Gna13 | 2383 | 1.233 | 0.0882 | No | ||
| 22 | ITGB6 | Itgb6 | 2499 | 1.178 | 0.0872 | No | ||
| 23 | DIAPH2 | Diaph2 | 2556 | 1.151 | 0.0900 | No | ||
| 24 | FGF5 | Fgf5 | 2817 | 1.046 | 0.0788 | No | ||
| 25 | FGFR4 | Fgfr4 | 2840 | 1.037 | 0.0831 | No | ||
| 26 | APC2 | Apc2 | 3148 | 0.936 | 0.0683 | No | ||
| 27 | BCAR1 | Bcar1 | 3392 | 0.865 | 0.0572 | No | ||
| 28 | MYL12B | Myl12b | 3439 | 0.856 | 0.0590 | No | ||
| 29 | ITGA10 | Gm42957 | 3450 | 0.853 | 0.0631 | No | ||
| 30 | FGF11 | Fgf11 | 3502 | 0.839 | 0.0644 | No | ||
| 31 | FGF18 | Fgf18 | 3517 | 0.834 | 0.0681 | No | ||
| 32 | PIK3CG | Pik3cg | 3524 | 0.831 | 0.0723 | No | ||
| 33 | FN1 | Fn1 | 3678 | 0.794 | 0.0668 | No | ||
| 34 | WASL | Wasl | 3722 | 0.784 | 0.0683 | No | ||
| 35 | PIP5K1A | Pip5k1a | 3748 | 0.776 | 0.0710 | No | ||
| 36 | GNA12 | Gna12 | 3817 | 0.756 | 0.0707 | No | ||
| 37 | FGF23 | Fgf23 | 3960 | 0.732 | 0.0655 | No | ||
| 38 | MAPK3 | Mapk3 | 4012 | 0.720 | 0.0662 | No | ||
| 39 | PIP4K2A | Pip4k2a | 4115 | 0.695 | 0.0634 | No | ||
| 40 | PAK5 | Pak7 | 4180 | 0.678 | 0.0630 | No | ||
| 41 | MYH9 | Myh9 | 4423 | 0.614 | 0.0506 | No | ||
| 42 | ARHGAP35 | Arhgap35 | 4541 | 0.582 | 0.0462 | No | ||
| 43 | SOS2 | Sos2 | 4544 | 0.582 | 0.0493 | No | ||
| 44 | CHRM4 | Chrm4 | 4578 | 0.575 | 0.0503 | No | ||
| 45 | ARAF | Araf | 4642 | 0.557 | 0.0493 | No | ||
| 46 | CFL2 | Cfl2 | 4849 | 0.507 | 0.0387 | No | ||
| 47 | CHRM5 | Chrm5 | 5312 | 0.428 | 0.0109 | No | ||
| 48 | ITGB8 | Itgb8 | 5402 | 0.417 | 0.0075 | No | ||
| 49 | MYL10 | Myl10 | 5411 | 0.415 | 0.0092 | No | ||
| 50 | DIAPH1 | Diaph1 | 5425 | 0.412 | 0.0107 | No | ||
| 51 | BRAF | Braf | 5445 | 0.408 | 0.0117 | No | ||
| 52 | ITGA2B | Itga2b | 5599 | 0.396 | 0.0039 | No | ||
| 53 | ITGA3 | Itga3 | 5667 | 0.382 | 0.0017 | No | ||
| 54 | IQGAP1 | Iqgap1 | 5698 | 0.375 | 0.0018 | No | ||
| 55 | ARPC1A | Arpc1a | 5719 | 0.369 | 0.0025 | No | ||
| 56 | ARHGEF12 | Arhgef12 | 5723 | 0.368 | 0.0044 | No | ||
| 57 | VAV1 | Vav1 | 5811 | 0.356 | 0.0007 | No | ||
| 58 | VAV2 | Vav2 | 6085 | 0.320 | -0.0154 | No | ||
| 59 | APC | Apc | 6192 | 0.299 | -0.0206 | No | ||
| 60 | FGF2 | Fgf2 | 6392 | 0.258 | -0.0322 | No | ||
| 61 | FGFR1 | Fgfr1 | 6421 | 0.253 | -0.0326 | No | ||
| 62 | INSRR | Insrr | 6454 | 0.246 | -0.0333 | No | ||
| 63 | FGD3 | Fgd3 | 6466 | 0.241 | -0.0327 | No | ||
| 64 | CSK | Csk | 6540 | 0.226 | -0.0362 | No | ||
| 65 | MRAS | Mras | 6567 | 0.221 | -0.0367 | No | ||
| 66 | GNG12 | Gng12 | 6617 | 0.212 | -0.0387 | No | ||
| 67 | ARPC4 | Arpc4 | 6622 | 0.210 | -0.0378 | No | ||
| 68 | ARPC5 | Arpc5 | 6673 | 0.201 | -0.0399 | No | ||
| 69 | LIMK1 | Limk1 | 6779 | 0.181 | -0.0458 | No | ||
| 70 | F2 | F2 | 6823 | 0.170 | -0.0476 | No | ||
| 71 | CHRM3 | Chrm3 | 6842 | 0.166 | -0.0479 | No | ||
| 72 | NCKAP1 | Nckap1 | 6888 | 0.157 | -0.0499 | No | ||
| 73 | ROCK1 | Rock1 | 7085 | 0.119 | -0.0621 | No | ||
| 74 | CRK | Crk | 7127 | 0.113 | -0.0641 | No | ||
| 75 | MSN | Msn | 7312 | 0.075 | -0.0757 | No | ||
| 76 | SSH1 | Ssh1 | 7359 | 0.067 | -0.0783 | No | ||
| 77 | PXN | Pxn | 7416 | 0.059 | -0.0816 | No | ||
| 78 | ABI2 | Abi2 | 7447 | 0.056 | -0.0833 | No | ||
| 79 | PIP5K1C | Pip5k1c | 7567 | 0.035 | -0.0909 | No | ||
| 80 | PIK3CA | Pik3ca | 7585 | 0.030 | -0.0918 | No | ||
| 81 | WAS | Was | 7646 | 0.019 | -0.0956 | No | ||
| 82 | BDKRB2 | Bdkrb2 | 7707 | 0.008 | -0.0995 | No | ||
| 83 | FGF9 | Fgf9 | 7721 | 0.004 | -0.1003 | No | ||
| 84 | FGF6 | Fgf6 | 7965 | 0.000 | -0.1162 | No | ||
| 85 | FGF19 | Fgf15 | 8061 | -0.009 | -0.1223 | No | ||
| 86 | MYLK3 | Mylk3 | 8106 | -0.017 | -0.1251 | No | ||
| 87 | LIMK2 | Limk2 | 8144 | -0.026 | -0.1273 | No | ||
| 88 | SOS1 | Sos1 | 8166 | -0.030 | -0.1286 | No | ||
| 89 | F2R | F2r | 8183 | -0.033 | -0.1294 | No | ||
| 90 | MYLK2 | Mylk2 | 8447 | -0.083 | -0.1461 | No | ||
| 91 | VAV3 | Vav3 | 8467 | -0.087 | -0.1469 | No | ||
| 92 | MYH14 | Myh14 | 8506 | -0.095 | -0.1488 | No | ||
| 93 | ARPC3 | Arpc3 | 8527 | -0.098 | -0.1496 | No | ||
| 94 | MYL2 | Myl2 | 8535 | -0.099 | -0.1495 | No | ||
| 95 | PAK6 | Pak6 | 8612 | -0.113 | -0.1538 | No | ||
| 96 | FGD1 | Fgd1 | 8628 | -0.115 | -0.1542 | No | ||
| 97 | ITGA5 | Itga5 | 8800 | -0.145 | -0.1645 | No | ||
| 98 | TIAM1 | Tiam1 | 8817 | -0.149 | -0.1647 | No | ||
| 99 | PIK3R5 | Pik3r5 | 8900 | -0.160 | -0.1692 | No | ||
| 100 | ITGAL | Itgal | 8955 | -0.169 | -0.1718 | No | ||
| 101 | RAC3 | Rac3 | 9015 | -0.180 | -0.1746 | No | ||
| 102 | PDGFA | Pdgfa | 9111 | -0.196 | -0.1797 | No | ||
| 103 | ACTN2 | Actn2 | 9209 | -0.215 | -0.1849 | No | ||
| 104 | RHOA | Rhoa | 9229 | -0.220 | -0.1849 | No | ||
| 105 | FGF12 | Fgf12 | 9237 | -0.221 | -0.1841 | No | ||
| 106 | RAC2 | Rac2 | 9271 | -0.227 | -0.1850 | No | ||
| 107 | HRAS | Hras | 9287 | -0.231 | -0.1847 | No | ||
| 108 | MAP2K1 | Map2k1 | 9298 | -0.232 | -0.1841 | No | ||
| 109 | WASF2 | Wasf2 | 9358 | -0.245 | -0.1866 | No | ||
| 110 | PFN1 | Pfn1 | 9394 | -0.252 | -0.1874 | No | ||
| 111 | ACTN1 | Actn1 | 9458 | -0.264 | -0.1901 | No | ||
| 112 | ARHGEF7 | Arhgef7 | 9461 | -0.264 | -0.1887 | No | ||
| 113 | ITGAM | Itgam | 9504 | -0.270 | -0.1900 | No | ||
| 114 | PAK4 | Pak4 | 9515 | -0.272 | -0.1891 | No | ||
| 115 | FGF10 | Fgf10 | 9558 | -0.280 | -0.1903 | No | ||
| 116 | ARPC2 | Arpc2 | 9632 | -0.295 | -0.1934 | No | ||
| 117 | CRKL | Crkl | 9708 | -0.310 | -0.1966 | No | ||
| 118 | PPP1R12A | Ppp1r12a | 9753 | -0.318 | -0.1977 | No | ||
| 119 | CFL1 | Cfl1 | 9812 | -0.332 | -0.1997 | No | ||
| 120 | ARPC1B | Arpc1b | 9829 | -0.334 | -0.1989 | No | ||
| 121 | PDGFC | Pdgfc | 9842 | -0.336 | -0.1978 | No | ||
| 122 | PAK3 | Pak3 | 9856 | -0.340 | -0.1967 | No | ||
| 123 | ACTB | Actb | 10078 | -0.384 | -0.2090 | No | ||
| 124 | SLC9A1 | Slc9a1 | 10140 | -0.396 | -0.2108 | No | ||
| 125 | ITGB1 | Itgb1 | 10224 | -0.414 | -0.2139 | No | ||
| 126 | ACTG1 | Actg1 | 10326 | -0.421 | -0.2182 | No | ||
| 127 | ITGA11 | Itga11 | 10396 | -0.426 | -0.2203 | No | ||
| 128 | PPP1CB | Ppp1cb | 10446 | -0.435 | -0.2211 | No | ||
| 129 | BAIAP2 | Baiap2 | 10449 | -0.435 | -0.2188 | No | ||
| 130 | ROCK2 | Rock2 | 10537 | -0.442 | -0.2220 | No | ||
| 131 | PAK2 | Pak2 | 10634 | -0.463 | -0.2257 | No | ||
| 132 | PFN2 | Pfn2 | 10684 | -0.475 | -0.2263 | No | ||
| 133 | MAPK1 | Mapk1 | 10768 | -0.489 | -0.2290 | No | ||
| 134 | CYFIP1 | Cyfip1 | 10770 | -0.491 | -0.2263 | No | ||
| 135 | MYLPF | Mylpf | 10790 | -0.494 | -0.2248 | No | ||
| 136 | CDC42 | Cdc42 | 11074 | -0.550 | -0.2402 | No | ||
| 137 | VCL | Vcl | 11113 | -0.558 | -0.2396 | No | ||
| 138 | KRAS | Kras | 11151 | -0.567 | -0.2389 | No | ||
| 139 | TIAM2 | Tiam2 | 11232 | -0.586 | -0.2408 | No | ||
| 140 | ARPC5L | Arpc5l | 11253 | -0.591 | -0.2389 | No | ||
| 141 | PIP4K2B | Pip4k2b | 11350 | -0.615 | -0.2417 | No | ||
| 142 | PTK2 | Ptk2 | 11527 | -0.655 | -0.2496 | No | ||
| 143 | RAF1 | Raf1 | 11555 | -0.662 | -0.2477 | No | ||
| 144 | FGF21 | Fgf21 | 11557 | -0.662 | -0.2440 | No | ||
| 145 | BRK1 | Brk1 | 11580 | -0.666 | -0.2418 | No | ||
| 146 | MYL12A | Myl12a | 11642 | -0.682 | -0.2420 | No | ||
| 147 | ITGAD | Itgad | 11683 | -0.694 | -0.2407 | No | ||
| 148 | ITGA7 | Itga7 | 11843 | -0.740 | -0.2470 | No | ||
| 149 | ITGAV | Itgav | 12032 | -0.796 | -0.2549 | No | ||
| 150 | PPP1CA | Ppp1ca | 12259 | -0.857 | -0.2648 | No | ||
| 151 | ARHGEF1 | Arhgef1 | 12307 | -0.869 | -0.2631 | No | ||
| 152 | RAC1 | Rac1 | 12614 | -0.976 | -0.2776 | No | ||
| 153 | TMSB4X | Tmsb4x | 12703 | -1.008 | -0.2778 | Yes | ||
| 154 | PPP1CC | Ppp1cc | 12721 | -1.015 | -0.2733 | Yes | ||
| 155 | SSH2 | Ssh2 | 12732 | -1.019 | -0.2683 | Yes | ||
| 156 | MYL9 | Myl9 | 12847 | -1.060 | -0.2698 | Yes | ||
| 157 | SCIN | Scin | 13005 | -1.122 | -0.2738 | Yes | ||
| 158 | MYH10 | Myh10 | 13018 | -1.127 | -0.2684 | Yes | ||
| 159 | DOCK1 | Dock1 | 13026 | -1.132 | -0.2625 | Yes | ||
| 160 | PAK1 | Pak1 | 13152 | -1.182 | -0.2641 | Yes | ||
| 161 | EGF | Egf | 13161 | -1.188 | -0.2581 | Yes | ||
| 162 | MYL7 | Myl7 | 13295 | -1.248 | -0.2598 | Yes | ||
| 163 | RRAS | Rras | 13300 | -1.249 | -0.2531 | Yes | ||
| 164 | RDX | Rdx | 13370 | -1.294 | -0.2505 | Yes | ||
| 165 | ACTN4 | Actn4 | 13388 | -1.302 | -0.2443 | Yes | ||
| 166 | FGF7 | Fgf7 | 13408 | -1.311 | -0.2383 | Yes | ||
| 167 | WASF1 | Wasf1 | 13411 | -1.313 | -0.2312 | Yes | ||
| 168 | FGF14 | Fgf14 | 13412 | -1.313 | -0.2239 | Yes | ||
| 169 | ITGA8 | Itga8 | 13437 | -1.329 | -0.2181 | Yes | ||
| 170 | GSN | Gsn | 13471 | -1.343 | -0.2128 | Yes | ||
| 171 | MOS | Mos | 13547 | -1.385 | -0.2100 | Yes | ||
| 172 | PIK3CB | Pik3cb | 13719 | -1.479 | -0.2129 | Yes | ||
| 173 | PIK3R1 | Pik3r1 | 13721 | -1.480 | -0.2048 | Yes | ||
| 174 | RRAS2 | Rras2 | 13860 | -1.582 | -0.2050 | Yes | ||
| 175 | ITGB4 | Itgb4 | 14125 | -1.795 | -0.2123 | Yes | ||
| 176 | ITGA6 | Itga6 | 14158 | -1.823 | -0.2042 | Yes | ||
| 177 | BDKRB1 | Bdkrb1 | 14206 | -1.872 | -0.1969 | Yes | ||
| 178 | PFN4 | Pfn4 | 14217 | -1.884 | -0.1871 | Yes | ||
| 179 | ITGA4 | Itga4 | 14396 | -2.091 | -0.1871 | Yes | ||
| 180 | PDGFRA | Pdgfra | 14432 | -2.130 | -0.1776 | Yes | ||
| 181 | PDGFD | Pdgfd | 14454 | -2.166 | -0.1669 | Yes | ||
| 182 | PDGFRB | Pdgfrb | 14505 | -2.226 | -0.1578 | Yes | ||
| 183 | EGFR | Egfr | 14536 | -2.270 | -0.1472 | Yes | ||
| 184 | CD14 | Cd14 | 14584 | -2.340 | -0.1373 | Yes | ||
| 185 | ITGA1 | Itga1 | 14614 | -2.383 | -0.1259 | Yes | ||
| 186 | IQGAP2 | Iqgap2 | 14642 | -2.438 | -0.1142 | Yes | ||
| 187 | ITGA2 | Itga2 | 14691 | -2.524 | -0.1033 | Yes | ||
| 188 | ACTN3 | Actn3 | 14766 | -2.627 | -0.0935 | Yes | ||
| 189 | MYLK | Mylk | 14799 | -2.685 | -0.0807 | Yes | ||
| 190 | ITGAE | Itgae | 15042 | -3.244 | -0.0785 | Yes | ||
| 191 | NCKAP1L | Nckap1l | 15145 | -3.560 | -0.0654 | Yes | ||
| 192 | FGF1 | Fgf1 | 15206 | -3.722 | -0.0487 | Yes | ||
| 193 | ITGAX | Itgax | 15301 | -4.050 | -0.0323 | Yes | ||
| 194 | DIAPH3 | Diaph3 | 15343 | -4.194 | -0.0117 | Yes | ||
| 195 | IQGAP3 | Iqgap3 | 15355 | -4.245 | 0.0111 | Yes |