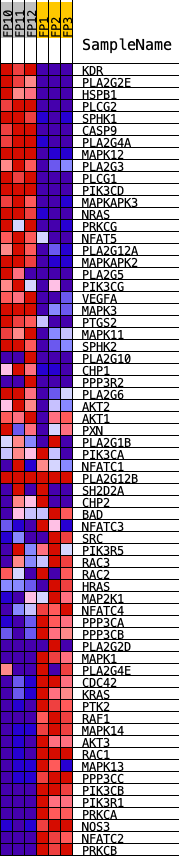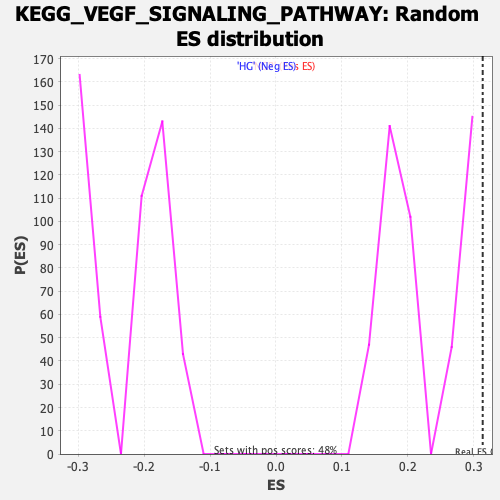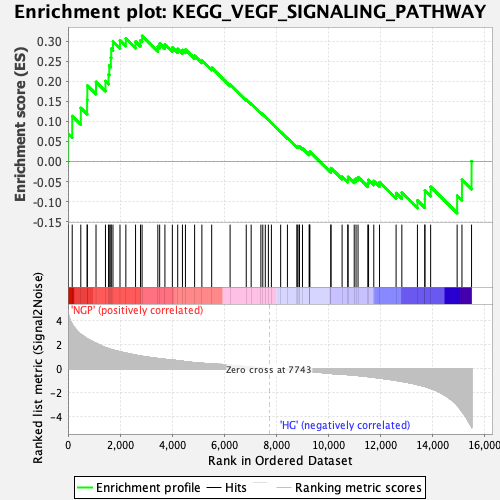
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_HG.phenotype.cls#NGP_versus_HG_repos |
| Phenotype | phenotype.cls#NGP_versus_HG_repos |
| Upregulated in class | NGP |
| GeneSet | KEGG_VEGF_SIGNALING_PATHWAY |
| Enrichment Score (ES) | 0.31334507 |
| Normalized Enrichment Score (NES) | 1.3852144 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.27192703 |
| FWER p-Value | 0.856 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | KDR | Kdr | 6 | 4.826 | 0.0694 | Yes | ||
| 2 | PLA2G2E | Pla2g2e | 162 | 3.725 | 0.1133 | Yes | ||
| 3 | HSPB1 | Hspb1 | 492 | 2.867 | 0.1335 | Yes | ||
| 4 | PLCG2 | Plcg2 | 735 | 2.492 | 0.1539 | Yes | ||
| 5 | SPHK1 | Sphk1 | 744 | 2.485 | 0.1894 | Yes | ||
| 6 | CASP9 | Casp9 | 1077 | 2.120 | 0.1986 | Yes | ||
| 7 | PLA2G4A | Pla2g4a | 1440 | 1.760 | 0.2006 | Yes | ||
| 8 | MAPK12 | Mapk12 | 1560 | 1.674 | 0.2171 | Yes | ||
| 9 | PLA2G3 | Pla2g3 | 1586 | 1.650 | 0.2394 | Yes | ||
| 10 | PLCG1 | Plcg1 | 1648 | 1.613 | 0.2588 | Yes | ||
| 11 | PIK3CD | Pik3cd | 1661 | 1.603 | 0.2812 | Yes | ||
| 12 | MAPKAPK3 | Mapkapk3 | 1729 | 1.562 | 0.2995 | Yes | ||
| 13 | NRAS | Nras | 1999 | 1.405 | 0.3024 | Yes | ||
| 14 | PRKCG | Prkcg | 2224 | 1.299 | 0.3067 | Yes | ||
| 15 | NFAT5 | Nfat5 | 2597 | 1.137 | 0.2991 | Yes | ||
| 16 | PLA2G12A | Pla2g12a | 2787 | 1.061 | 0.3023 | Yes | ||
| 17 | MAPKAPK2 | Mapkapk2 | 2848 | 1.035 | 0.3133 | Yes | ||
| 18 | PLA2G5 | Pla2g5 | 3454 | 0.853 | 0.2866 | No | ||
| 19 | PIK3CG | Pik3cg | 3524 | 0.831 | 0.2941 | No | ||
| 20 | VEGFA | Vegfa | 3726 | 0.782 | 0.2924 | No | ||
| 21 | MAPK3 | Mapk3 | 4012 | 0.720 | 0.2844 | No | ||
| 22 | PTGS2 | Ptgs2 | 4218 | 0.666 | 0.2808 | No | ||
| 23 | MAPK11 | Mapk11 | 4400 | 0.621 | 0.2781 | No | ||
| 24 | SPHK2 | Sphk2 | 4515 | 0.587 | 0.2792 | No | ||
| 25 | PLA2G10 | Pla2g10 | 4868 | 0.503 | 0.2637 | No | ||
| 26 | CHP1 | Chp1 | 5147 | 0.454 | 0.2523 | No | ||
| 27 | PPP3R2 | Ppp3r2 | 5524 | 0.401 | 0.2338 | No | ||
| 28 | PLA2G6 | Pla2g6 | 6232 | 0.291 | 0.1923 | No | ||
| 29 | AKT2 | Akt2 | 6853 | 0.164 | 0.1546 | No | ||
| 30 | AKT1 | Akt1 | 7042 | 0.129 | 0.1443 | No | ||
| 31 | PXN | Pxn | 7416 | 0.059 | 0.1210 | No | ||
| 32 | PLA2G1B | Pla2g1b | 7489 | 0.048 | 0.1170 | No | ||
| 33 | PIK3CA | Pik3ca | 7585 | 0.030 | 0.1113 | No | ||
| 34 | NFATC1 | Nfatc1 | 7704 | 0.008 | 0.1038 | No | ||
| 35 | PLA2G12B | Pla2g12b | 7820 | 0.000 | 0.0964 | No | ||
| 36 | SH2D2A | Sh2d2a | 8176 | -0.033 | 0.0739 | No | ||
| 37 | CHP2 | Chp2 | 8436 | -0.082 | 0.0583 | No | ||
| 38 | BAD | Bad | 8802 | -0.146 | 0.0368 | No | ||
| 39 | NFATC3 | Nfatc3 | 8813 | -0.148 | 0.0383 | No | ||
| 40 | SRC | Src | 8880 | -0.156 | 0.0363 | No | ||
| 41 | PIK3R5 | Pik3r5 | 8900 | -0.160 | 0.0374 | No | ||
| 42 | RAC3 | Rac3 | 9015 | -0.180 | 0.0326 | No | ||
| 43 | RAC2 | Rac2 | 9271 | -0.227 | 0.0194 | No | ||
| 44 | HRAS | Hras | 9287 | -0.231 | 0.0218 | No | ||
| 45 | MAP2K1 | Map2k1 | 9298 | -0.232 | 0.0245 | No | ||
| 46 | NFATC4 | Nfatc4 | 10107 | -0.391 | -0.0221 | No | ||
| 47 | PPP3CA | Ppp3ca | 10109 | -0.391 | -0.0165 | No | ||
| 48 | PPP3CB | Ppp3cb | 10538 | -0.443 | -0.0378 | No | ||
| 49 | PLA2G2D | Pla2g2d | 10756 | -0.489 | -0.0448 | No | ||
| 50 | MAPK1 | Mapk1 | 10768 | -0.489 | -0.0384 | No | ||
| 51 | PLA2G4E | Pla2g4e | 11004 | -0.534 | -0.0459 | No | ||
| 52 | CDC42 | Cdc42 | 11074 | -0.550 | -0.0424 | No | ||
| 53 | KRAS | Kras | 11151 | -0.567 | -0.0391 | No | ||
| 54 | PTK2 | Ptk2 | 11527 | -0.655 | -0.0539 | No | ||
| 55 | RAF1 | Raf1 | 11555 | -0.662 | -0.0460 | No | ||
| 56 | MAPK14 | Mapk14 | 11756 | -0.714 | -0.0486 | No | ||
| 57 | AKT3 | Akt3 | 11977 | -0.777 | -0.0516 | No | ||
| 58 | RAC1 | Rac1 | 12614 | -0.976 | -0.0787 | No | ||
| 59 | MAPK13 | Mapk13 | 12835 | -1.057 | -0.0776 | No | ||
| 60 | PPP3CC | Ppp3cc | 13434 | -1.328 | -0.0971 | No | ||
| 61 | PIK3CB | Pik3cb | 13719 | -1.479 | -0.0940 | No | ||
| 62 | PIK3R1 | Pik3r1 | 13721 | -1.480 | -0.0727 | No | ||
| 63 | PRKCA | Prkca | 13942 | -1.648 | -0.0631 | No | ||
| 64 | NOS3 | Nos3 | 14961 | -3.019 | -0.0852 | No | ||
| 65 | NFATC2 | Nfatc2 | 15146 | -3.562 | -0.0456 | No | ||
| 66 | PRKCB | Prkcb | 15515 | -4.838 | 0.0006 | No |