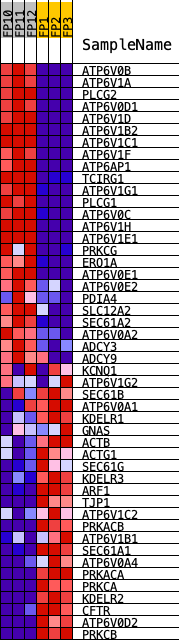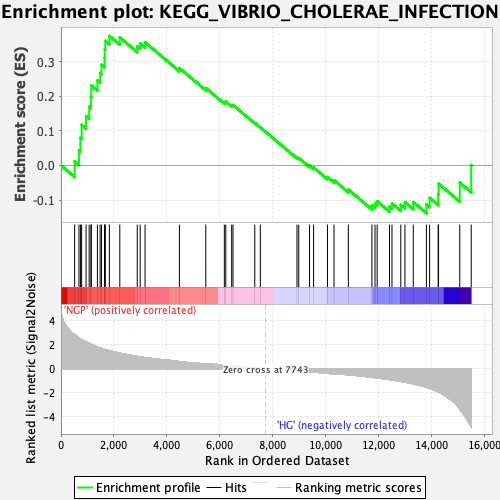
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_HG.phenotype.cls#NGP_versus_HG_repos |
| Phenotype | phenotype.cls#NGP_versus_HG_repos |
| Upregulated in class | NGP |
| GeneSet | KEGG_VIBRIO_CHOLERAE_INFECTION |
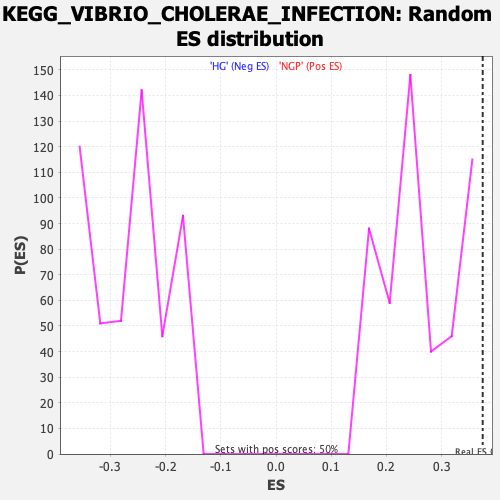
| Enrichment Score (ES) | 0.374299 |
| Normalized Enrichment Score (NES) | 1.4277242 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.31774378 |
| FWER p-Value | 0.856 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ATP6V0B | Atp6v0b | 516 | 2.828 | 0.0124 | Yes | ||
| 2 | ATP6V1A | Atp6v1a | 679 | 2.574 | 0.0435 | Yes | ||
| 3 | PLCG2 | Plcg2 | 735 | 2.492 | 0.0802 | Yes | ||
| 4 | ATP6V0D1 | Atp6v0d1 | 781 | 2.439 | 0.1167 | Yes | ||
| 5 | ATP6V1D | Atp6v1d | 948 | 2.260 | 0.1425 | Yes | ||
| 6 | ATP6V1B2 | Atp6v1b2 | 1065 | 2.129 | 0.1695 | Yes | ||
| 7 | ATP6V1C1 | Atp6v1c1 | 1132 | 2.064 | 0.1986 | Yes | ||
| 8 | ATP6V1F | Atp6v1f | 1147 | 2.051 | 0.2308 | Yes | ||
| 9 | ATP6AP1 | Atp6ap1 | 1377 | 1.821 | 0.2455 | Yes | ||
| 10 | TCIRG1 | Tcirg1 | 1479 | 1.737 | 0.2670 | Yes | ||
| 11 | ATP6V1G1 | Atp6v1g1 | 1533 | 1.696 | 0.2910 | Yes | ||
| 12 | PLCG1 | Plcg1 | 1648 | 1.613 | 0.3097 | Yes | ||
| 13 | ATP6V0C | Atp6v0c | 1653 | 1.608 | 0.3354 | Yes | ||
| 14 | ATP6V1H | Atp6v1h | 1675 | 1.596 | 0.3599 | Yes | ||
| 15 | ATP6V1E1 | Atp6v1e1 | 1829 | 1.504 | 0.3743 | Yes | ||
| 16 | PRKCG | Prkcg | 2224 | 1.299 | 0.3698 | No | ||
| 17 | ERO1A | Ero1l | 2884 | 1.023 | 0.3438 | No | ||
| 18 | ATP6V0E1 | Atp6v0e | 2994 | 0.981 | 0.3526 | No | ||
| 19 | ATP6V0E2 | Atp6v0e2 | 3180 | 0.924 | 0.3556 | No | ||
| 20 | PDIA4 | Pdia4 | 4479 | 0.596 | 0.2814 | No | ||
| 21 | SLC12A2 | Slc12a2 | 5476 | 0.402 | 0.2235 | No | ||
| 22 | SEC61A2 | Sec61a2 | 6178 | 0.302 | 0.1831 | No | ||
| 23 | ATP6V0A2 | Atp6v0a2 | 6233 | 0.291 | 0.1843 | No | ||
| 24 | ADCY3 | Adcy3 | 6455 | 0.245 | 0.1740 | No | ||
| 25 | ADCY9 | Adcy9 | 6506 | 0.234 | 0.1746 | No | ||
| 26 | KCNQ1 | Kcnq1 | 7329 | 0.073 | 0.1226 | No | ||
| 27 | ATP6V1G2 | Atp6v1g2 | 7538 | 0.039 | 0.1098 | No | ||
| 28 | SEC61B | Gm10320 | 8925 | -0.165 | 0.0230 | No | ||
| 29 | ATP6V0A1 | Atp6v0a1 | 8992 | -0.175 | 0.0215 | No | ||
| 30 | KDELR1 | Kdelr1 | 9400 | -0.253 | -0.0007 | No | ||
| 31 | GNAS | Gnas | 9554 | -0.280 | -0.0060 | No | ||
| 32 | ACTB | Actb | 10078 | -0.384 | -0.0336 | No | ||
| 33 | ACTG1 | Actg1 | 10326 | -0.421 | -0.0428 | No | ||
| 34 | SEC61G | Sec61g | 10868 | -0.506 | -0.0696 | No | ||
| 35 | KDELR3 | Kdelr3 | 11760 | -0.715 | -0.1156 | No | ||
| 36 | ARF1 | Arf1 | 11876 | -0.751 | -0.1109 | No | ||
| 37 | TJP1 | Tjp1 | 11958 | -0.773 | -0.1036 | No | ||
| 38 | ATP6V1C2 | Atp6v1c2 | 12423 | -0.908 | -0.1189 | No | ||
| 39 | PRKACB | Prkacb | 12518 | -0.946 | -0.1097 | No | ||
| 40 | ATP6V1B1 | Atp6v1b1 | 12852 | -1.064 | -0.1140 | No | ||
| 41 | SEC61A1 | Sec61a1 | 13008 | -1.123 | -0.1059 | No | ||
| 42 | ATP6V0A4 | Atp6v0a4 | 13323 | -1.264 | -0.1057 | No | ||
| 43 | PRKACA | Prkaca | 13821 | -1.556 | -0.1127 | No | ||
| 44 | PRKCA | Prkca | 13942 | -1.648 | -0.0938 | No | ||
| 45 | KDELR2 | Kdelr2 | 14265 | -1.934 | -0.0833 | No | ||
| 46 | CFTR | Cftr | 14279 | -1.953 | -0.0526 | No | ||
| 47 | ATP6V0D2 | Atp6v0d2 | 15082 | -3.390 | -0.0496 | No | ||
| 48 | PRKCB | Prkcb | 15515 | -4.838 | 0.0006 | No |