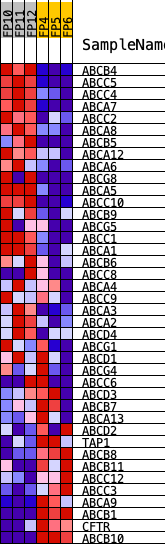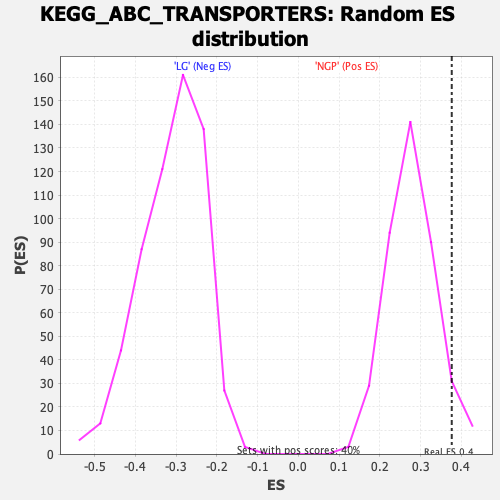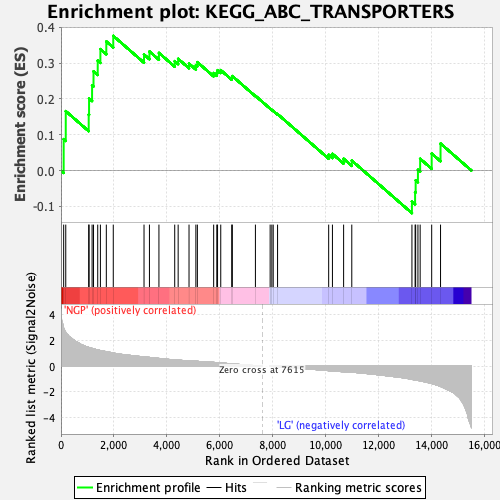
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_LG.phenotype.cls#NGP_versus_LG_repos |
| Phenotype | phenotype.cls#NGP_versus_LG_repos |
| Upregulated in class | NGP |
| GeneSet | KEGG_ABC_TRANSPORTERS |
| Enrichment Score (ES) | 0.3762589 |
| Normalized Enrichment Score (NES) | 1.3488419 |
| Nominal p-value | 0.0675 |
| FDR q-value | 0.43926054 |
| FWER p-Value | 0.991 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ABCB4 | Abcb4 | 101 | 2.979 | 0.0878 | Yes | ||
| 2 | ABCC5 | Abcc5 | 180 | 2.631 | 0.1660 | Yes | ||
| 3 | ABCC4 | Abcc4 | 1048 | 1.462 | 0.1563 | Yes | ||
| 4 | ABCA7 | Abca7 | 1062 | 1.453 | 0.2015 | Yes | ||
| 5 | ABCC2 | Abcc2 | 1176 | 1.385 | 0.2380 | Yes | ||
| 6 | ABCA8 | Abca8b | 1230 | 1.348 | 0.2773 | Yes | ||
| 7 | ABCB5 | Abcb5 | 1389 | 1.267 | 0.3072 | Yes | ||
| 8 | ABCA12 | Abca12 | 1490 | 1.218 | 0.3392 | Yes | ||
| 9 | ABCA6 | Abca6 | 1714 | 1.130 | 0.3606 | Yes | ||
| 10 | ABCG8 | Abcg8 | 1977 | 1.029 | 0.3763 | Yes | ||
| 11 | ABCA5 | Abca5 | 3140 | 0.728 | 0.3243 | No | ||
| 12 | ABCC10 | Abcc10 | 3345 | 0.682 | 0.3327 | No | ||
| 13 | ABCB9 | Abcb9 | 3704 | 0.603 | 0.3287 | No | ||
| 14 | ABCG5 | Abcg5 | 4301 | 0.488 | 0.3056 | No | ||
| 15 | ABCC1 | Abcc1 | 4431 | 0.470 | 0.3122 | No | ||
| 16 | ABCA1 | Abca1 | 4842 | 0.416 | 0.2989 | No | ||
| 17 | ABCB6 | Abcb6 | 5101 | 0.387 | 0.2945 | No | ||
| 18 | ABCC8 | Abcc8 | 5158 | 0.377 | 0.3028 | No | ||
| 19 | ABCA4 | Abca4 | 5774 | 0.294 | 0.2724 | No | ||
| 20 | ABCC9 | Abcc9 | 5893 | 0.275 | 0.2735 | No | ||
| 21 | ABCA3 | Abca3 | 5921 | 0.272 | 0.2803 | No | ||
| 22 | ABCA2 | Abca2 | 6042 | 0.251 | 0.2806 | No | ||
| 23 | ABCD4 | Abcd4 | 6457 | 0.184 | 0.2596 | No | ||
| 24 | ABCG1 | Abcg1 | 6480 | 0.180 | 0.2639 | No | ||
| 25 | ABCD1 | Abcd1 | 7353 | 0.041 | 0.2089 | No | ||
| 26 | ABCG4 | Abcg4 | 7910 | -0.004 | 0.1731 | No | ||
| 27 | ABCC6 | Abcc6 | 7975 | -0.014 | 0.1694 | No | ||
| 28 | ABCD3 | Abcd3 | 8026 | -0.024 | 0.1670 | No | ||
| 29 | ABCB7 | Abcb7 | 8191 | -0.048 | 0.1579 | No | ||
| 30 | ABCA13 | Abca13 | 10124 | -0.346 | 0.0441 | No | ||
| 31 | ABCD2 | Abcd2 | 10266 | -0.374 | 0.0468 | No | ||
| 32 | TAP1 | Tap1 | 10689 | -0.425 | 0.0330 | No | ||
| 33 | ABCB8 | Abcb8 | 10998 | -0.469 | 0.0280 | No | ||
| 34 | ABCB11 | Abcb11 | 13273 | -1.027 | -0.0864 | No | ||
| 35 | ABCC12 | Abcc12 | 13393 | -1.072 | -0.0601 | No | ||
| 36 | ABCC3 | Abcc3 | 13414 | -1.079 | -0.0272 | No | ||
| 37 | ABCA9 | Abca9 | 13500 | -1.104 | 0.0022 | No | ||
| 38 | ABCB1 | Abcb1a | 13584 | -1.142 | 0.0330 | No | ||
| 39 | CFTR | Cftr | 14020 | -1.347 | 0.0476 | No | ||
| 40 | ABCB10 | Abcb10 | 14355 | -1.565 | 0.0756 | No |