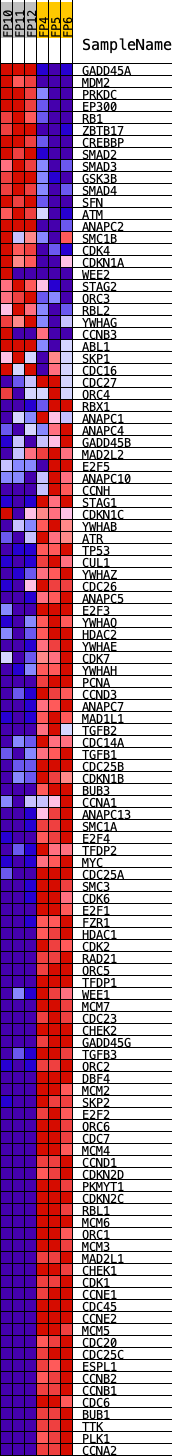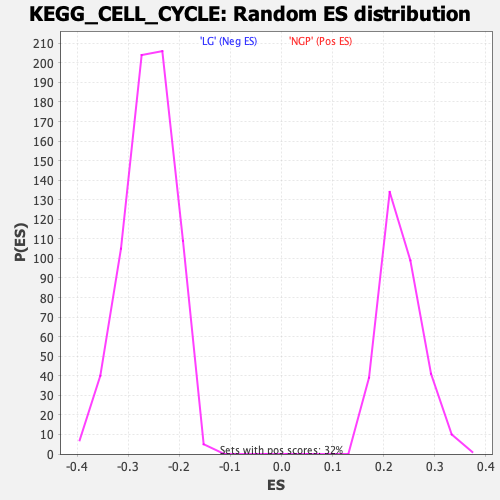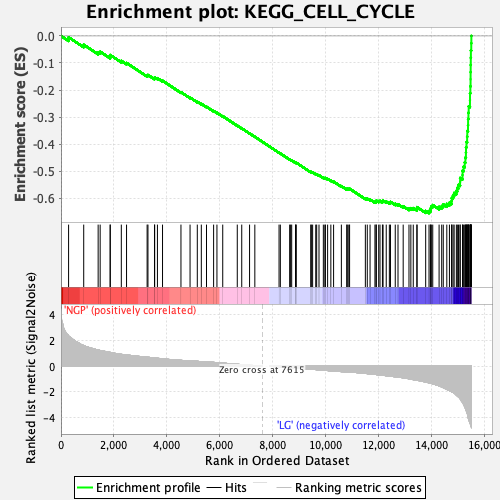
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_LG.phenotype.cls#NGP_versus_LG_repos |
| Phenotype | phenotype.cls#NGP_versus_LG_repos |
| Upregulated in class | LG |
| GeneSet | KEGG_CELL_CYCLE |
| Enrichment Score (ES) | -0.6553098 |
| Normalized Enrichment Score (NES) | -2.522597 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | GADD45A | Gadd45a | 287 | 2.360 | -0.0049 | No | ||
| 2 | MDM2 | Mdm2 | 859 | 1.605 | -0.0326 | No | ||
| 3 | PRKDC | Prkdc | 1405 | 1.257 | -0.0606 | No | ||
| 4 | EP300 | Ep300 | 1481 | 1.221 | -0.0584 | No | ||
| 5 | RB1 | Rb1 | 1859 | 1.074 | -0.0766 | No | ||
| 6 | ZBTB17 | Zbtb17 | 1863 | 1.072 | -0.0706 | No | ||
| 7 | CREBBP | Crebbp | 2281 | 0.921 | -0.0923 | No | ||
| 8 | SMAD2 | Smad2 | 2479 | 0.869 | -0.1000 | No | ||
| 9 | SMAD3 | Smad3 | 3257 | 0.699 | -0.1464 | No | ||
| 10 | GSK3B | Gsk3b | 3289 | 0.693 | -0.1443 | No | ||
| 11 | SMAD4 | Smad4 | 3536 | 0.639 | -0.1566 | No | ||
| 12 | SFN | Sfn | 3543 | 0.638 | -0.1532 | No | ||
| 13 | ATM | Atm | 3647 | 0.615 | -0.1563 | No | ||
| 14 | ANAPC2 | Anapc2 | 3840 | 0.569 | -0.1655 | No | ||
| 15 | SMC1B | Smc1b | 4537 | 0.453 | -0.2080 | No | ||
| 16 | CDK4 | Cdk4 | 4881 | 0.408 | -0.2279 | No | ||
| 17 | CDKN1A | Cdkn1a | 5155 | 0.378 | -0.2434 | No | ||
| 18 | WEE2 | Wee2 | 5307 | 0.356 | -0.2511 | No | ||
| 19 | STAG2 | Stag2 | 5503 | 0.341 | -0.2618 | No | ||
| 20 | ORC3 | Orc3 | 5772 | 0.294 | -0.2775 | No | ||
| 21 | RBL2 | Rbl2 | 5898 | 0.274 | -0.2840 | No | ||
| 22 | YWHAG | Ywhag | 6117 | 0.240 | -0.2968 | No | ||
| 23 | CCNB3 | Ccnb3 | 6667 | 0.148 | -0.3315 | No | ||
| 24 | ABL1 | Abl1 | 6836 | 0.119 | -0.3417 | No | ||
| 25 | SKP1 | Skp1a | 7132 | 0.074 | -0.3605 | No | ||
| 26 | CDC16 | Cdc16 | 7331 | 0.044 | -0.3730 | No | ||
| 27 | CDC27 | Cdc27 | 8247 | -0.057 | -0.4321 | No | ||
| 28 | ORC4 | Orc4 | 8296 | -0.065 | -0.4348 | No | ||
| 29 | RBX1 | Rbx1 | 8648 | -0.115 | -0.4569 | No | ||
| 30 | ANAPC1 | Anapc1 | 8685 | -0.120 | -0.4586 | No | ||
| 31 | ANAPC4 | Anapc4 | 8725 | -0.128 | -0.4604 | No | ||
| 32 | GADD45B | Gadd45b | 8870 | -0.149 | -0.4688 | No | ||
| 33 | MAD2L2 | Mad2l2 | 8884 | -0.151 | -0.4688 | No | ||
| 34 | E2F5 | E2f5 | 8895 | -0.152 | -0.4686 | No | ||
| 35 | ANAPC10 | Anapc10 | 9445 | -0.234 | -0.5028 | No | ||
| 36 | CCNH | Ccnh | 9469 | -0.237 | -0.5029 | No | ||
| 37 | STAG1 | Stag1 | 9517 | -0.245 | -0.5046 | No | ||
| 38 | CDKN1C | Cdkn1c | 9635 | -0.261 | -0.5106 | No | ||
| 39 | YWHAB | Ywhab | 9670 | -0.266 | -0.5113 | No | ||
| 40 | ATR | Atr | 9758 | -0.283 | -0.5153 | No | ||
| 41 | TP53 | Trp53 | 9922 | -0.312 | -0.5241 | No | ||
| 42 | CUL1 | Cul1 | 9979 | -0.320 | -0.5258 | No | ||
| 43 | YWHAZ | Ywhaz | 10001 | -0.323 | -0.5253 | No | ||
| 44 | CDC26 | Cdc26 | 10084 | -0.337 | -0.5287 | No | ||
| 45 | ANAPC5 | Anapc5 | 10206 | -0.364 | -0.5344 | No | ||
| 46 | E2F3 | E2f3 | 10310 | -0.381 | -0.5389 | No | ||
| 47 | YWHAQ | Ywhaq | 10604 | -0.421 | -0.5554 | No | ||
| 48 | HDAC2 | Hdac2 | 10806 | -0.436 | -0.5659 | No | ||
| 49 | YWHAE | Ywhae | 10828 | -0.440 | -0.5647 | No | ||
| 50 | CDK7 | Cdk7 | 10885 | -0.449 | -0.5658 | No | ||
| 51 | YWHAH | Ywhah | 10915 | -0.456 | -0.5650 | No | ||
| 52 | PCNA | Pcna | 11511 | -0.560 | -0.6003 | No | ||
| 53 | CCND3 | Ccnd3 | 11584 | -0.575 | -0.6017 | No | ||
| 54 | ANAPC7 | Anapc7 | 11689 | -0.596 | -0.6050 | No | ||
| 55 | MAD1L1 | Mad1l1 | 11874 | -0.636 | -0.6132 | No | ||
| 56 | TGFB2 | Tgfb2 | 11914 | -0.646 | -0.6120 | No | ||
| 57 | CDC14A | Cdc14a | 11927 | -0.649 | -0.6090 | No | ||
| 58 | TGFB1 | Tgfb1 | 12016 | -0.671 | -0.6108 | No | ||
| 59 | CDC25B | Cdc25b | 12065 | -0.682 | -0.6099 | No | ||
| 60 | CDKN1B | Cdkn1b | 12157 | -0.705 | -0.6117 | No | ||
| 61 | BUB3 | Bub3 | 12181 | -0.710 | -0.6091 | No | ||
| 62 | CCNA1 | Ccna1 | 12301 | -0.740 | -0.6125 | No | ||
| 63 | ANAPC13 | Anapc13 | 12419 | -0.766 | -0.6156 | No | ||
| 64 | SMC1A | Smc1a | 12461 | -0.775 | -0.6138 | No | ||
| 65 | E2F4 | E2f4 | 12642 | -0.823 | -0.6207 | No | ||
| 66 | TFDP2 | Tfdp2 | 12746 | -0.846 | -0.6224 | No | ||
| 67 | MYC | Myc | 12942 | -0.905 | -0.6298 | No | ||
| 68 | CDC25A | Cdc25a | 13161 | -0.974 | -0.6383 | No | ||
| 69 | SMC3 | Smc3 | 13232 | -1.004 | -0.6370 | No | ||
| 70 | CDK6 | Cdk6 | 13323 | -1.045 | -0.6368 | No | ||
| 71 | E2F1 | E2f1 | 13461 | -1.093 | -0.6393 | No | ||
| 72 | FZR1 | Fzr1 | 13466 | -1.093 | -0.6332 | No | ||
| 73 | HDAC1 | Hdac1 | 13793 | -1.225 | -0.6472 | No | ||
| 74 | CDK2 | Cdk2 | 13919 | -1.295 | -0.6478 | Yes | ||
| 75 | RAD21 | Rad21 | 13971 | -1.318 | -0.6434 | Yes | ||
| 76 | ORC5 | Orc5 | 13985 | -1.327 | -0.6365 | Yes | ||
| 77 | TFDP1 | Tfdp1 | 14006 | -1.336 | -0.6300 | Yes | ||
| 78 | WEE1 | Wee1 | 14060 | -1.370 | -0.6255 | Yes | ||
| 79 | MCM7 | Mcm7 | 14300 | -1.525 | -0.6321 | Yes | ||
| 80 | CDC23 | Cdc23 | 14399 | -1.616 | -0.6291 | Yes | ||
| 81 | CHEK2 | Chek2 | 14458 | -1.668 | -0.6232 | Yes | ||
| 82 | GADD45G | Gadd45g | 14592 | -1.803 | -0.6213 | Yes | ||
| 83 | TGFB3 | Tgfb3 | 14691 | -1.893 | -0.6166 | Yes | ||
| 84 | ORC2 | Orc2 | 14773 | -1.989 | -0.6103 | Yes | ||
| 85 | DBF4 | Dbf4 | 14780 | -1.994 | -0.5991 | Yes | ||
| 86 | MCM2 | Mcm2 | 14827 | -2.038 | -0.5902 | Yes | ||
| 87 | SKP2 | Skp2 | 14869 | -2.091 | -0.5807 | Yes | ||
| 88 | E2F2 | E2f2 | 14953 | -2.242 | -0.5730 | Yes | ||
| 89 | ORC6 | Orc6 | 14990 | -2.313 | -0.5619 | Yes | ||
| 90 | CDC7 | Cdc7 | 15033 | -2.387 | -0.5508 | Yes | ||
| 91 | MCM4 | Mcm4 | 15094 | -2.529 | -0.5399 | Yes | ||
| 92 | CCND1 | Ccnd1 | 15096 | -2.535 | -0.5252 | Yes | ||
| 93 | CDKN2D | Cdkn2d | 15186 | -2.823 | -0.5146 | Yes | ||
| 94 | PKMYT1 | Pkmyt1 | 15188 | -2.827 | -0.4982 | Yes | ||
| 95 | CDKN2C | Cdkn2c | 15228 | -2.983 | -0.4834 | Yes | ||
| 96 | RBL1 | Rbl1 | 15270 | -3.172 | -0.4676 | Yes | ||
| 97 | MCM6 | Mcm6 | 15295 | -3.297 | -0.4499 | Yes | ||
| 98 | ORC1 | Orc1 | 15315 | -3.391 | -0.4314 | Yes | ||
| 99 | MCM3 | Mcm3 | 15319 | -3.408 | -0.4118 | Yes | ||
| 100 | MAD2L1 | Mad2l1 | 15336 | -3.510 | -0.3924 | Yes | ||
| 101 | CHEK1 | Chek1 | 15361 | -3.652 | -0.3727 | Yes | ||
| 102 | CDK1 | Cdk1 | 15367 | -3.695 | -0.3515 | Yes | ||
| 103 | CCNE1 | Ccne1 | 15394 | -3.949 | -0.3302 | Yes | ||
| 104 | CDC45 | Cdc45 | 15398 | -4.001 | -0.3071 | Yes | ||
| 105 | CCNE2 | Ccne2 | 15410 | -4.096 | -0.2840 | Yes | ||
| 106 | MCM5 | Mcm5 | 15420 | -4.154 | -0.2604 | Yes | ||
| 107 | CDC20 | Cdc20 | 15468 | -4.434 | -0.2376 | Yes | ||
| 108 | CDC25C | Cdc25c | 15470 | -4.435 | -0.2119 | Yes | ||
| 109 | ESPL1 | Espl1 | 15483 | -4.511 | -0.1864 | Yes | ||
| 110 | CCNB2 | Ccnb2 | 15491 | -4.557 | -0.1603 | Yes | ||
| 111 | CCNB1 | Ccnb1 | 15492 | -4.568 | -0.1337 | Yes | ||
| 112 | CDC6 | Cdc6 | 15495 | -4.576 | -0.1072 | Yes | ||
| 113 | BUB1 | Bub1 | 15499 | -4.609 | -0.0806 | Yes | ||
| 114 | TTK | Ttk | 15503 | -4.625 | -0.0539 | Yes | ||
| 115 | PLK1 | Plk1 | 15517 | -4.730 | -0.0272 | Yes | ||
| 116 | CCNA2 | Ccna2 | 15521 | -4.748 | 0.0003 | Yes |