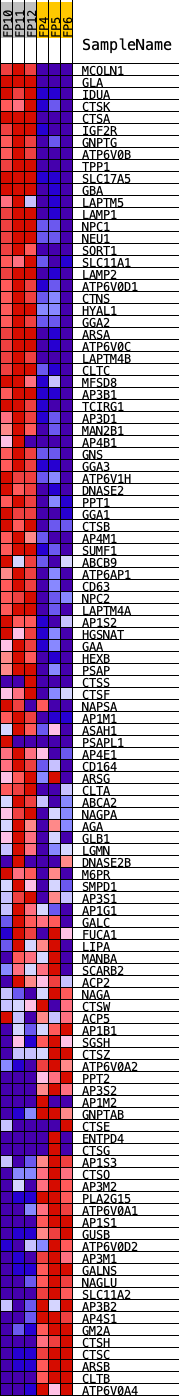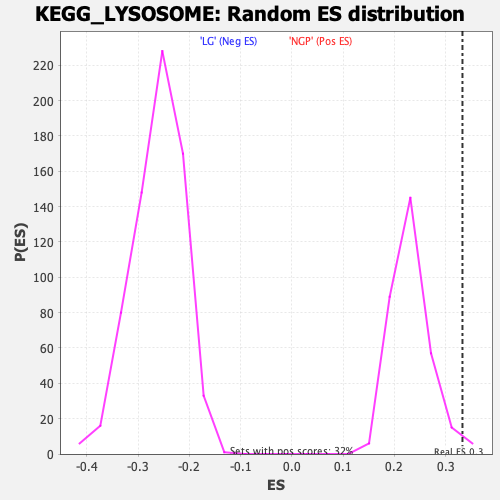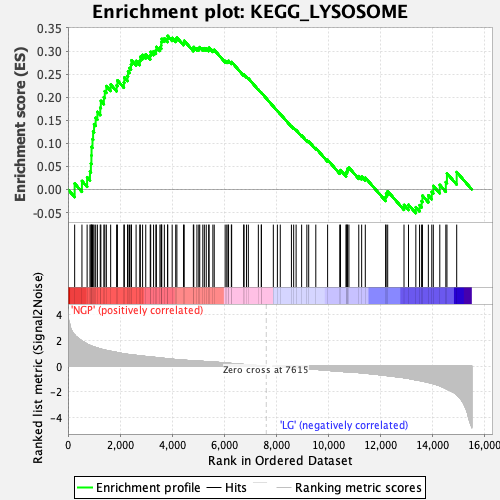
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_LG.phenotype.cls#NGP_versus_LG_repos |
| Phenotype | phenotype.cls#NGP_versus_LG_repos |
| Upregulated in class | NGP |
| GeneSet | KEGG_LYSOSOME |
| Enrichment Score (ES) | 0.3328711 |
| Normalized Enrichment Score (NES) | 1.4377623 |
| Nominal p-value | 0.018867925 |
| FDR q-value | 0.3729831 |
| FWER p-Value | 0.917 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | MCOLN1 | Mcoln1 | 256 | 2.433 | 0.0129 | Yes | ||
| 2 | GLA | Gla | 536 | 1.948 | 0.0185 | Yes | ||
| 3 | IDUA | Idua | 735 | 1.713 | 0.0264 | Yes | ||
| 4 | CTSK | Ctsk | 845 | 1.617 | 0.0389 | Yes | ||
| 5 | CTSA | Ctsa | 881 | 1.582 | 0.0559 | Yes | ||
| 6 | IGF2R | Igf2r | 900 | 1.570 | 0.0738 | Yes | ||
| 7 | GNPTG | Gnptg | 906 | 1.564 | 0.0924 | Yes | ||
| 8 | ATP6V0B | Atp6v0b | 942 | 1.535 | 0.1088 | Yes | ||
| 9 | TPP1 | Tpp1 | 965 | 1.521 | 0.1258 | Yes | ||
| 10 | SLC17A5 | Slc17a5 | 1002 | 1.498 | 0.1416 | Yes | ||
| 11 | GBA | Gba | 1064 | 1.452 | 0.1553 | Yes | ||
| 12 | LAPTM5 | Laptm5 | 1130 | 1.406 | 0.1682 | Yes | ||
| 13 | LAMP1 | Lamp1 | 1240 | 1.341 | 0.1774 | Yes | ||
| 14 | NPC1 | Npc1 | 1263 | 1.331 | 0.1921 | Yes | ||
| 15 | NEU1 | Neu1 | 1379 | 1.272 | 0.2001 | Yes | ||
| 16 | SORT1 | Sort1 | 1415 | 1.252 | 0.2130 | Yes | ||
| 17 | SLC11A1 | Slc11a1 | 1474 | 1.225 | 0.2241 | Yes | ||
| 18 | LAMP2 | Lamp2 | 1640 | 1.159 | 0.2274 | Yes | ||
| 19 | ATP6V0D1 | Atp6v0d1 | 1869 | 1.069 | 0.2256 | Yes | ||
| 20 | CTNS | Ctns | 1900 | 1.055 | 0.2365 | Yes | ||
| 21 | HYAL1 | Hyal1 | 2149 | 0.969 | 0.2321 | Yes | ||
| 22 | GGA2 | Gga2 | 2166 | 0.961 | 0.2428 | Yes | ||
| 23 | ARSA | Arsa | 2288 | 0.919 | 0.2461 | Yes | ||
| 24 | ATP6V0C | Atp6v0c | 2308 | 0.915 | 0.2559 | Yes | ||
| 25 | LAPTM4B | Laptm4b | 2364 | 0.900 | 0.2633 | Yes | ||
| 26 | CLTC | Cltc | 2422 | 0.886 | 0.2703 | Yes | ||
| 27 | MFSD8 | Mfsd8 | 2440 | 0.881 | 0.2799 | Yes | ||
| 28 | AP3B1 | Ap3b1 | 2618 | 0.839 | 0.2786 | Yes | ||
| 29 | TCIRG1 | Tcirg1 | 2760 | 0.808 | 0.2793 | Yes | ||
| 30 | AP3D1 | Ap3d1 | 2781 | 0.801 | 0.2877 | Yes | ||
| 31 | MAN2B1 | Man2b1 | 2868 | 0.780 | 0.2916 | Yes | ||
| 32 | AP4B1 | Ap4b1 | 2989 | 0.754 | 0.2930 | Yes | ||
| 33 | GNS | Gns | 3162 | 0.722 | 0.2906 | Yes | ||
| 34 | GGA3 | Gga3 | 3179 | 0.716 | 0.2982 | Yes | ||
| 35 | ATP6V1H | Atp6v1h | 3290 | 0.693 | 0.2995 | Yes | ||
| 36 | DNASE2 | Dnase2a | 3380 | 0.674 | 0.3019 | Yes | ||
| 37 | PPT1 | Ppt1 | 3397 | 0.670 | 0.3090 | Yes | ||
| 38 | GGA1 | Gga1 | 3535 | 0.640 | 0.3078 | Yes | ||
| 39 | CTSB | Ctsb | 3584 | 0.630 | 0.3124 | Yes | ||
| 40 | AP4M1 | Ap4m1 | 3587 | 0.629 | 0.3199 | Yes | ||
| 41 | SUMF1 | Sumf1 | 3605 | 0.625 | 0.3264 | Yes | ||
| 42 | ABCB9 | Abcb9 | 3704 | 0.603 | 0.3273 | Yes | ||
| 43 | ATP6AP1 | Atp6ap1 | 3827 | 0.572 | 0.3263 | Yes | ||
| 44 | CD63 | Cd63 | 3834 | 0.570 | 0.3329 | Yes | ||
| 45 | NPC2 | Npc2 | 4005 | 0.540 | 0.3284 | No | ||
| 46 | LAPTM4A | Laptm4a | 4141 | 0.514 | 0.3259 | No | ||
| 47 | AP1S2 | Ap1s2 | 4186 | 0.506 | 0.3292 | No | ||
| 48 | HGSNAT | Hgsnat | 4440 | 0.470 | 0.3184 | No | ||
| 49 | GAA | Gaa | 4468 | 0.465 | 0.3223 | No | ||
| 50 | HEXB | Hexb | 4819 | 0.419 | 0.3047 | No | ||
| 51 | PSAP | Psap | 4835 | 0.416 | 0.3088 | No | ||
| 52 | CTSS | Ctss | 4957 | 0.401 | 0.3058 | No | ||
| 53 | CTSF | Ctsf | 5029 | 0.399 | 0.3060 | No | ||
| 54 | NAPSA | Napsa | 5065 | 0.393 | 0.3085 | No | ||
| 55 | AP1M1 | Ap1m1 | 5183 | 0.372 | 0.3055 | No | ||
| 56 | ASAH1 | Asah1 | 5242 | 0.364 | 0.3061 | No | ||
| 57 | PSAPL1 | Psapl1 | 5315 | 0.356 | 0.3058 | No | ||
| 58 | AP4E1 | Ap4e1 | 5409 | 0.356 | 0.3040 | No | ||
| 59 | CD164 | Cd164 | 5424 | 0.354 | 0.3074 | No | ||
| 60 | ARSG | Arsg | 5570 | 0.331 | 0.3020 | No | ||
| 61 | CLTA | Clta | 5622 | 0.319 | 0.3026 | No | ||
| 62 | ABCA2 | Abca2 | 6042 | 0.251 | 0.2785 | No | ||
| 63 | NAGPA | Nagpa | 6105 | 0.242 | 0.2774 | No | ||
| 64 | AGA | Aga | 6153 | 0.235 | 0.2772 | No | ||
| 65 | GLB1 | Glb1 | 6173 | 0.231 | 0.2788 | No | ||
| 66 | LGMN | Lgmn | 6280 | 0.211 | 0.2744 | No | ||
| 67 | DNASE2B | Dnase2b | 6292 | 0.209 | 0.2763 | No | ||
| 68 | M6PR | M6pr | 6753 | 0.133 | 0.2480 | No | ||
| 69 | SMPD1 | Smpd1 | 6768 | 0.130 | 0.2487 | No | ||
| 70 | AP3S1 | Ap3s1 | 6862 | 0.114 | 0.2441 | No | ||
| 71 | AP1G1 | Ap1g1 | 6933 | 0.104 | 0.2408 | No | ||
| 72 | GALC | Galc | 7318 | 0.046 | 0.2164 | No | ||
| 73 | FUCA1 | Fuca1 | 7432 | 0.029 | 0.2094 | No | ||
| 74 | LIPA | Lipa | 7436 | 0.028 | 0.2096 | No | ||
| 75 | MANBA | Manba | 7894 | -0.001 | 0.1800 | No | ||
| 76 | SCARB2 | Scarb2 | 8050 | -0.028 | 0.1702 | No | ||
| 77 | ACP2 | Acp2 | 8161 | -0.045 | 0.1636 | No | ||
| 78 | NAGA | Naga | 8590 | -0.107 | 0.1372 | No | ||
| 79 | CTSW | Ctsw | 8676 | -0.119 | 0.1331 | No | ||
| 80 | ACP5 | Acp5 | 8770 | -0.137 | 0.1287 | No | ||
| 81 | AP1B1 | Ap1b1 | 8982 | -0.166 | 0.1171 | No | ||
| 82 | SGSH | Sgsh | 9183 | -0.191 | 0.1064 | No | ||
| 83 | CTSZ | Ctsz | 9254 | -0.202 | 0.1043 | No | ||
| 84 | ATP6V0A2 | Atp6v0a2 | 9527 | -0.247 | 0.0897 | No | ||
| 85 | PPT2 | Ppt2 | 9981 | -0.320 | 0.0642 | No | ||
| 86 | AP3S2 | Ap3s2 | 10448 | -0.404 | 0.0388 | No | ||
| 87 | AP1M2 | Ap1m2 | 10477 | -0.407 | 0.0419 | No | ||
| 88 | GNPTAB | Gnptab | 10691 | -0.425 | 0.0333 | No | ||
| 89 | CTSE | Ctse | 10700 | -0.427 | 0.0379 | No | ||
| 90 | ENTPD4 | Entpd4b | 10733 | -0.432 | 0.0411 | No | ||
| 91 | CTSG | Ctsg | 10749 | -0.432 | 0.0454 | No | ||
| 92 | AP1S3 | Ap1s3 | 10800 | -0.435 | 0.0474 | No | ||
| 93 | CTSO | Ctso | 11179 | -0.494 | 0.0289 | No | ||
| 94 | AP3M2 | Ap3m2 | 11290 | -0.515 | 0.0280 | No | ||
| 95 | PLA2G15 | Pla2g15 | 11430 | -0.541 | 0.0255 | No | ||
| 96 | ATP6V0A1 | Atp6v0a1 | 12211 | -0.715 | -0.0164 | No | ||
| 97 | AP1S1 | Ap1s1 | 12230 | -0.719 | -0.0088 | No | ||
| 98 | GUSB | Gusb | 12291 | -0.736 | -0.0038 | No | ||
| 99 | ATP6V0D2 | Atp6v0d2 | 12918 | -0.895 | -0.0335 | No | ||
| 100 | AP3M1 | Ap3m1 | 13091 | -0.950 | -0.0332 | No | ||
| 101 | GALNS | Galns | 13373 | -1.065 | -0.0385 | No | ||
| 102 | NAGLU | Naglu | 13516 | -1.112 | -0.0342 | No | ||
| 103 | SLC11A2 | Slc11a2 | 13595 | -1.145 | -0.0254 | No | ||
| 104 | AP3B2 | Ap3b2 | 13627 | -1.157 | -0.0133 | No | ||
| 105 | AP4S1 | Ap4s1 | 13854 | -1.253 | -0.0128 | No | ||
| 106 | GM2A | Gm2a | 13986 | -1.327 | -0.0052 | No | ||
| 107 | CTSH | Ctsh | 14045 | -1.360 | 0.0076 | No | ||
| 108 | CTSC | Ctsc | 14294 | -1.520 | 0.0099 | No | ||
| 109 | ARSB | Arsb | 14526 | -1.735 | 0.0160 | No | ||
| 110 | CLTB | Cltb | 14569 | -1.784 | 0.0349 | No | ||
| 111 | ATP6V0A4 | Atp6v0a4 | 14944 | -2.227 | 0.0377 | No |