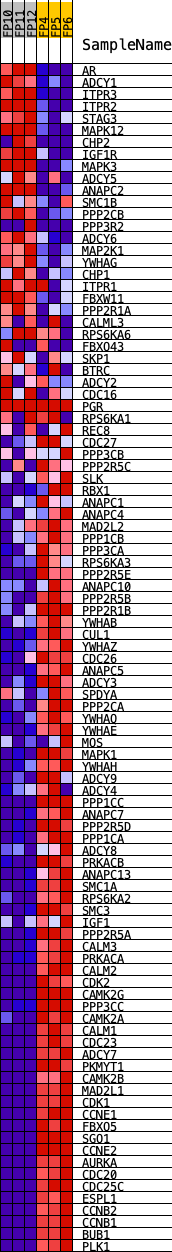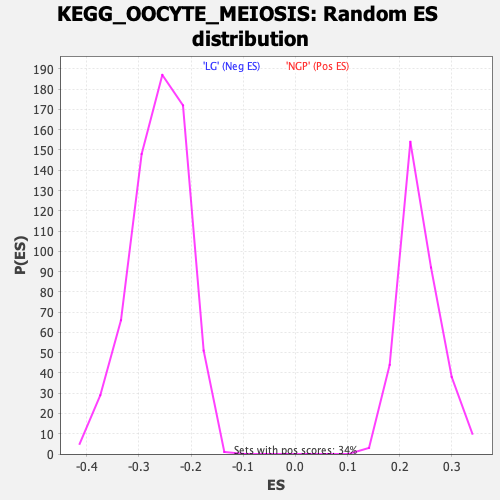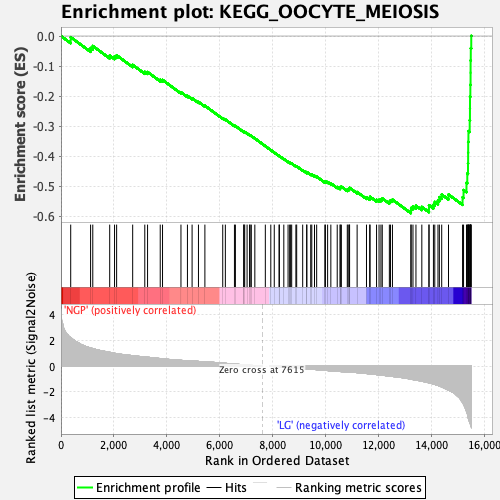
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_LG.phenotype.cls#NGP_versus_LG_repos |
| Phenotype | phenotype.cls#NGP_versus_LG_repos |
| Upregulated in class | LG |
| GeneSet | KEGG_OOCYTE_MEIOSIS |
| Enrichment Score (ES) | -0.5890038 |
| Normalized Enrichment Score (NES) | -2.2450452 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | AR | Ar | 367 | 2.230 | -0.0039 | No | ||
| 2 | ADCY1 | Adcy1 | 1119 | 1.417 | -0.0399 | No | ||
| 3 | ITPR3 | Itpr3 | 1202 | 1.362 | -0.0331 | No | ||
| 4 | ITPR2 | Itpr2 | 1842 | 1.079 | -0.0649 | No | ||
| 5 | STAG3 | Stag3 | 2025 | 1.005 | -0.0677 | No | ||
| 6 | MAPK12 | Mapk12 | 2104 | 0.983 | -0.0640 | No | ||
| 7 | CHP2 | Chp2 | 2711 | 0.819 | -0.0960 | No | ||
| 8 | IGF1R | Igf1r | 3169 | 0.719 | -0.1192 | No | ||
| 9 | MAPK3 | Mapk3 | 3274 | 0.697 | -0.1197 | No | ||
| 10 | ADCY5 | Adcy5 | 3753 | 0.589 | -0.1455 | No | ||
| 11 | ANAPC2 | Anapc2 | 3840 | 0.569 | -0.1460 | No | ||
| 12 | SMC1B | Smc1b | 4537 | 0.453 | -0.1871 | No | ||
| 13 | PPP2CB | Ppp2cb | 4782 | 0.427 | -0.1991 | No | ||
| 14 | PPP3R2 | Ppp3r2 | 4958 | 0.401 | -0.2068 | No | ||
| 15 | ADCY6 | Adcy6 | 5199 | 0.371 | -0.2191 | No | ||
| 16 | MAP2K1 | Map2k1 | 5442 | 0.350 | -0.2316 | No | ||
| 17 | YWHAG | Ywhag | 6117 | 0.240 | -0.2732 | No | ||
| 18 | CHP1 | Chp1 | 6216 | 0.224 | -0.2775 | No | ||
| 19 | ITPR1 | Itpr1 | 6560 | 0.165 | -0.2983 | No | ||
| 20 | FBXW11 | Fbxw11 | 6596 | 0.161 | -0.2991 | No | ||
| 21 | PPP2R1A | Ppp2r1a | 6912 | 0.107 | -0.3186 | No | ||
| 22 | CALML3 | Calml3 | 6917 | 0.107 | -0.3179 | No | ||
| 23 | RPS6KA6 | Rps6ka6 | 6948 | 0.102 | -0.3189 | No | ||
| 24 | FBXO43 | Fbxo43 | 7038 | 0.086 | -0.3239 | No | ||
| 25 | SKP1 | Skp1a | 7132 | 0.074 | -0.3293 | No | ||
| 26 | BTRC | Btrc | 7154 | 0.070 | -0.3301 | No | ||
| 27 | ADCY2 | Adcy2 | 7198 | 0.064 | -0.3323 | No | ||
| 28 | CDC16 | Cdc16 | 7331 | 0.044 | -0.3404 | No | ||
| 29 | PGR | Pgr | 7728 | 0.000 | -0.3661 | No | ||
| 30 | RPS6KA1 | Rps6ka1 | 7935 | -0.008 | -0.3794 | No | ||
| 31 | REC8 | Rec8 | 8066 | -0.030 | -0.3876 | No | ||
| 32 | CDC27 | Cdc27 | 8247 | -0.057 | -0.3987 | No | ||
| 33 | PPP3CB | Ppp3cb | 8260 | -0.059 | -0.3990 | No | ||
| 34 | PPP2R5C | Ppp2r5c | 8427 | -0.084 | -0.4090 | No | ||
| 35 | SLK | Slk | 8588 | -0.106 | -0.4184 | No | ||
| 36 | RBX1 | Rbx1 | 8648 | -0.115 | -0.4212 | No | ||
| 37 | ANAPC1 | Anapc1 | 8685 | -0.120 | -0.4225 | No | ||
| 38 | ANAPC4 | Anapc4 | 8725 | -0.128 | -0.4238 | No | ||
| 39 | MAD2L2 | Mad2l2 | 8884 | -0.151 | -0.4327 | No | ||
| 40 | PPP1CB | Ppp1cb | 8914 | -0.156 | -0.4332 | No | ||
| 41 | PPP3CA | Ppp3ca | 9141 | -0.186 | -0.4462 | No | ||
| 42 | RPS6KA3 | Rps6ka3 | 9294 | -0.208 | -0.4542 | No | ||
| 43 | PPP2R5E | Ppp2r5e | 9296 | -0.208 | -0.4524 | No | ||
| 44 | ANAPC10 | Anapc10 | 9445 | -0.234 | -0.4599 | No | ||
| 45 | PPP2R5B | Ppp2r5b | 9485 | -0.240 | -0.4603 | No | ||
| 46 | PPP2R1B | Ppp2r1b | 9588 | -0.253 | -0.4647 | No | ||
| 47 | YWHAB | Ywhab | 9670 | -0.266 | -0.4676 | No | ||
| 48 | CUL1 | Cul1 | 9979 | -0.320 | -0.4847 | No | ||
| 49 | YWHAZ | Ywhaz | 10001 | -0.323 | -0.4831 | No | ||
| 50 | CDC26 | Cdc26 | 10084 | -0.337 | -0.4855 | No | ||
| 51 | ANAPC5 | Anapc5 | 10206 | -0.364 | -0.4900 | No | ||
| 52 | ADCY3 | Adcy3 | 10447 | -0.403 | -0.5020 | No | ||
| 53 | SPDYA | Spdya | 10556 | -0.410 | -0.5054 | No | ||
| 54 | PPP2CA | Ppp2ca | 10563 | -0.411 | -0.5021 | No | ||
| 55 | YWHAQ | Ywhaq | 10604 | -0.421 | -0.5009 | No | ||
| 56 | YWHAE | Ywhae | 10828 | -0.440 | -0.5114 | No | ||
| 57 | MOS | Mos | 10861 | -0.445 | -0.5096 | No | ||
| 58 | MAPK1 | Mapk1 | 10902 | -0.453 | -0.5081 | No | ||
| 59 | YWHAH | Ywhah | 10915 | -0.456 | -0.5048 | No | ||
| 60 | ADCY9 | Adcy9 | 11200 | -0.498 | -0.5188 | No | ||
| 61 | ADCY4 | Adcy4 | 11554 | -0.569 | -0.5366 | No | ||
| 62 | PPP1CC | Ppp1cc | 11676 | -0.594 | -0.5391 | No | ||
| 63 | ANAPC7 | Anapc7 | 11689 | -0.596 | -0.5346 | No | ||
| 64 | PPP2R5D | Ppp2r5d | 11935 | -0.651 | -0.5447 | No | ||
| 65 | PPP1CA | Ppp1ca | 12029 | -0.673 | -0.5447 | No | ||
| 66 | ADCY8 | Adcy8 | 12101 | -0.691 | -0.5431 | No | ||
| 67 | PRKACB | Prkacb | 12151 | -0.704 | -0.5400 | No | ||
| 68 | ANAPC13 | Anapc13 | 12419 | -0.766 | -0.5505 | No | ||
| 69 | SMC1A | Smc1a | 12461 | -0.775 | -0.5463 | No | ||
| 70 | RPS6KA2 | Rps6ka2 | 12536 | -0.796 | -0.5440 | No | ||
| 71 | SMC3 | Smc3 | 13232 | -1.004 | -0.5801 | Yes | ||
| 72 | IGF1 | Igf1 | 13244 | -1.013 | -0.5717 | Yes | ||
| 73 | PPP2R5A | Ppp2r5a | 13312 | -1.041 | -0.5668 | Yes | ||
| 74 | CALM3 | Calm3 | 13425 | -1.084 | -0.5644 | Yes | ||
| 75 | PRKACA | Prkaca | 13647 | -1.163 | -0.5683 | Yes | ||
| 76 | CALM2 | Calm2 | 13917 | -1.295 | -0.5742 | Yes | ||
| 77 | CDK2 | Cdk2 | 13919 | -1.295 | -0.5627 | Yes | ||
| 78 | CAMK2G | Camk2g | 14086 | -1.388 | -0.5611 | Yes | ||
| 79 | PPP3CC | Ppp3cc | 14137 | -1.416 | -0.5517 | Yes | ||
| 80 | CAMK2A | Camk2a | 14258 | -1.496 | -0.5462 | Yes | ||
| 81 | CALM1 | Calm1 | 14310 | -1.533 | -0.5358 | Yes | ||
| 82 | CDC23 | Cdc23 | 14399 | -1.616 | -0.5271 | Yes | ||
| 83 | ADCY7 | Adcy7 | 14656 | -1.864 | -0.5271 | Yes | ||
| 84 | PKMYT1 | Pkmyt1 | 15188 | -2.827 | -0.5363 | Yes | ||
| 85 | CAMK2B | Camk2b | 15225 | -2.956 | -0.5122 | Yes | ||
| 86 | MAD2L1 | Mad2l1 | 15336 | -3.510 | -0.4881 | Yes | ||
| 87 | CDK1 | Cdk1 | 15367 | -3.695 | -0.4570 | Yes | ||
| 88 | CCNE1 | Ccne1 | 15394 | -3.949 | -0.4235 | Yes | ||
| 89 | FBXO5 | Fbxo5 | 15399 | -4.006 | -0.3880 | Yes | ||
| 90 | SGO1 | Sgo1 | 15401 | -4.029 | -0.3522 | Yes | ||
| 91 | CCNE2 | Ccne2 | 15410 | -4.096 | -0.3161 | Yes | ||
| 92 | AURKA | Aurka | 15458 | -4.378 | -0.2801 | Yes | ||
| 93 | CDC20 | Cdc20 | 15468 | -4.434 | -0.2412 | Yes | ||
| 94 | CDC25C | Cdc25c | 15470 | -4.435 | -0.2017 | Yes | ||
| 95 | ESPL1 | Espl1 | 15483 | -4.511 | -0.1622 | Yes | ||
| 96 | CCNB2 | Ccnb2 | 15491 | -4.557 | -0.1220 | Yes | ||
| 97 | CCNB1 | Ccnb1 | 15492 | -4.568 | -0.0813 | Yes | ||
| 98 | BUB1 | Bub1 | 15499 | -4.609 | -0.0406 | Yes | ||
| 99 | PLK1 | Plk1 | 15517 | -4.730 | 0.0005 | Yes |