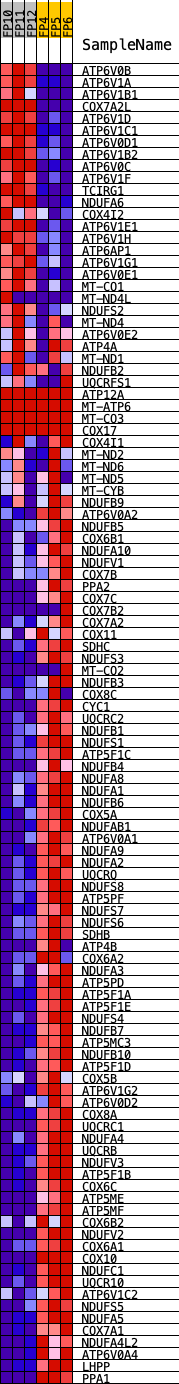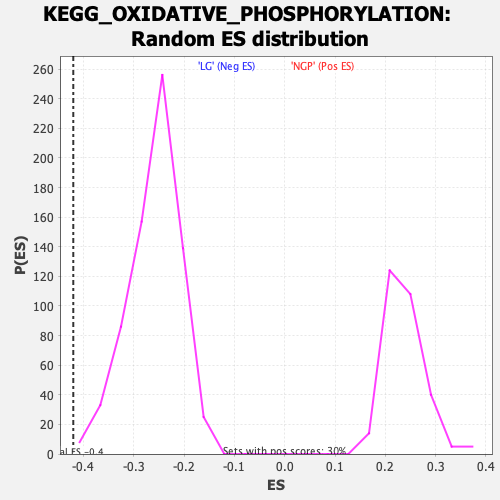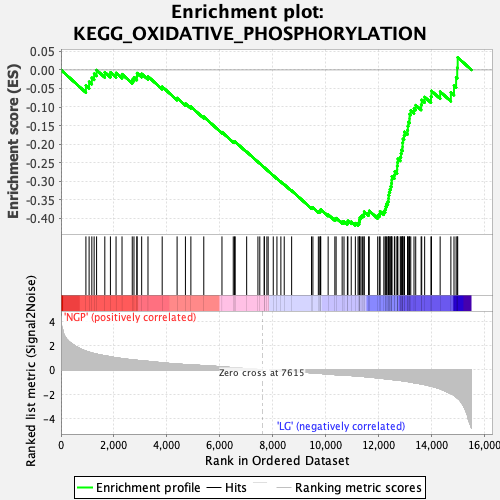
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_LG.phenotype.cls#NGP_versus_LG_repos |
| Phenotype | phenotype.cls#NGP_versus_LG_repos |
| Upregulated in class | LG |
| GeneSet | KEGG_OXIDATIVE_PHOSPHORYLATION |
| Enrichment Score (ES) | -0.4199656 |
| Normalized Enrichment Score (NES) | -1.6175982 |
| Nominal p-value | 0.0028409092 |
| FDR q-value | 0.054505203 |
| FWER p-Value | 0.622 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ATP6V0B | Atp6v0b | 942 | 1.535 | -0.0421 | No | ||
| 2 | ATP6V1A | Atp6v1a | 1063 | 1.452 | -0.0319 | No | ||
| 3 | ATP6V1B1 | Atp6v1b1 | 1166 | 1.391 | -0.0213 | No | ||
| 4 | COX7A2L | Cox7a2l | 1253 | 1.335 | -0.0103 | No | ||
| 5 | ATP6V1D | Atp6v1d | 1346 | 1.288 | -0.0003 | No | ||
| 6 | ATP6V1C1 | Atp6v1c1 | 1657 | 1.152 | -0.0061 | No | ||
| 7 | ATP6V0D1 | Atp6v0d1 | 1869 | 1.069 | -0.0066 | No | ||
| 8 | ATP6V1B2 | Atp6v1b2 | 2082 | 0.986 | -0.0081 | No | ||
| 9 | ATP6V0C | Atp6v0c | 2308 | 0.915 | -0.0114 | No | ||
| 10 | ATP6V1F | Atp6v1f | 2695 | 0.823 | -0.0262 | No | ||
| 11 | TCIRG1 | Tcirg1 | 2760 | 0.808 | -0.0204 | No | ||
| 12 | NDUFA6 | Ndufa6 | 2870 | 0.780 | -0.0178 | No | ||
| 13 | COX4I2 | Cox4i2 | 2876 | 0.778 | -0.0085 | No | ||
| 14 | ATP6V1E1 | Atp6v1e1 | 3051 | 0.742 | -0.0106 | No | ||
| 15 | ATP6V1H | Atp6v1h | 3290 | 0.693 | -0.0174 | No | ||
| 16 | ATP6AP1 | Atp6ap1 | 3827 | 0.572 | -0.0451 | No | ||
| 17 | ATP6V1G1 | Atp6v1g1 | 4391 | 0.476 | -0.0757 | No | ||
| 18 | ATP6V0E1 | Atp6v0e | 4706 | 0.433 | -0.0907 | No | ||
| 19 | MT-CO1 | mt-Co1 | 4912 | 0.401 | -0.0990 | No | ||
| 20 | MT-ND4L | mt-Nd4l | 5400 | 0.356 | -0.1262 | No | ||
| 21 | NDUFS2 | Ndufs2 | 6088 | 0.245 | -0.1678 | No | ||
| 22 | MT-ND4 | mt-Nd4 | 6515 | 0.173 | -0.1932 | No | ||
| 23 | ATP6V0E2 | Atp6v0e2 | 6563 | 0.165 | -0.1943 | No | ||
| 24 | ATP4A | Atp4a | 6588 | 0.162 | -0.1938 | No | ||
| 25 | MT-ND1 | mt-Nd1 | 7022 | 0.089 | -0.2208 | No | ||
| 26 | NDUFB2 | Ndufb2 | 7448 | 0.026 | -0.2480 | No | ||
| 27 | UQCRFS1 | Uqcrfs1 | 7519 | 0.017 | -0.2524 | No | ||
| 28 | ATP12A | Atp12a | 7688 | 0.000 | -0.2633 | No | ||
| 29 | MT-ATP6 | mt-Atp6 | 7689 | 0.000 | -0.2633 | No | ||
| 30 | MT-CO3 | mt-Co3 | 7787 | 0.000 | -0.2696 | No | ||
| 31 | COX17 | Cox17 | 7835 | 0.000 | -0.2726 | No | ||
| 32 | COX4I1 | Cox4i1 | 8028 | -0.025 | -0.2848 | No | ||
| 33 | MT-ND2 | mt-Nd2 | 8167 | -0.045 | -0.2931 | No | ||
| 34 | MT-ND6 | mt-Nd6 | 8316 | -0.068 | -0.3019 | No | ||
| 35 | MT-ND5 | mt-Nd5 | 8443 | -0.086 | -0.3090 | No | ||
| 36 | MT-CYB | mt-Cytb | 8723 | -0.128 | -0.3255 | No | ||
| 37 | NDUFB9 | Ndufb9 | 9475 | -0.238 | -0.3713 | No | ||
| 38 | ATP6V0A2 | Atp6v0a2 | 9527 | -0.247 | -0.3715 | No | ||
| 39 | NDUFB5 | Ndufb5 | 9733 | -0.280 | -0.3814 | No | ||
| 40 | COX6B1 | Cox6b1 | 9788 | -0.286 | -0.3813 | No | ||
| 41 | NDUFA10 | Ndufa10 | 9814 | -0.293 | -0.3793 | No | ||
| 42 | NDUFV1 | Ndufv1 | 9824 | -0.293 | -0.3763 | No | ||
| 43 | COX7B | Cox7b | 10104 | -0.341 | -0.3902 | No | ||
| 44 | PPA2 | Ppa2 | 10358 | -0.388 | -0.4018 | No | ||
| 45 | COX7C | Cox7c | 10405 | -0.397 | -0.3998 | No | ||
| 46 | COX7B2 | Cox7b2 | 10634 | -0.421 | -0.4094 | No | ||
| 47 | COX7A2 | Cox7a2 | 10704 | -0.427 | -0.4086 | No | ||
| 48 | COX11 | Cox11 | 10832 | -0.440 | -0.4114 | No | ||
| 49 | SDHC | Sdhc | 10851 | -0.443 | -0.4070 | No | ||
| 50 | NDUFS3 | Ndufs3 | 10977 | -0.465 | -0.4094 | No | ||
| 51 | MT-CO2 | mt-Co2 | 11133 | -0.487 | -0.4134 | No | ||
| 52 | NDUFB3 | Ndufb3 | 11235 | -0.505 | -0.4137 | Yes | ||
| 53 | COX8C | Cox8b | 11277 | -0.512 | -0.4100 | Yes | ||
| 54 | CYC1 | Cyc1 | 11289 | -0.515 | -0.4044 | Yes | ||
| 55 | UQCRC2 | Uqcrc2 | 11303 | -0.518 | -0.3988 | Yes | ||
| 56 | NDUFB1 | Ndufb1-ps | 11347 | -0.526 | -0.3951 | Yes | ||
| 57 | NDUFS1 | Ndufs1 | 11400 | -0.535 | -0.3918 | Yes | ||
| 58 | ATP5F1C | Atp5c1 | 11459 | -0.548 | -0.3888 | Yes | ||
| 59 | NDUFB4 | Ndufb4 | 11466 | -0.550 | -0.3824 | Yes | ||
| 60 | NDUFA8 | Ndufa8 | 11633 | -0.584 | -0.3859 | Yes | ||
| 61 | NDUFA1 | Ndufa1 | 11658 | -0.590 | -0.3801 | Yes | ||
| 62 | NDUFB6 | Ndufb6 | 11978 | -0.661 | -0.3926 | Yes | ||
| 63 | COX5A | Cox5a | 12040 | -0.676 | -0.3882 | Yes | ||
| 64 | NDUFAB1 | Ndufab1 | 12069 | -0.683 | -0.3816 | Yes | ||
| 65 | ATP6V0A1 | Atp6v0a1 | 12211 | -0.715 | -0.3819 | Yes | ||
| 66 | NDUFA9 | Ndufa9 | 12260 | -0.729 | -0.3759 | Yes | ||
| 67 | NDUFA2 | Ndufa2 | 12284 | -0.734 | -0.3683 | Yes | ||
| 68 | UQCRQ | Uqcrq | 12315 | -0.742 | -0.3611 | Yes | ||
| 69 | NDUFS8 | Ndufs8 | 12364 | -0.755 | -0.3549 | Yes | ||
| 70 | ATP5PF | Atp5j | 12388 | -0.760 | -0.3469 | Yes | ||
| 71 | NDUFS7 | Ndufs7 | 12392 | -0.763 | -0.3377 | Yes | ||
| 72 | NDUFS6 | Ndufs6 | 12410 | -0.765 | -0.3293 | Yes | ||
| 73 | SDHB | Sdhb | 12441 | -0.770 | -0.3217 | Yes | ||
| 74 | ATP4B | Atp4b | 12472 | -0.778 | -0.3140 | Yes | ||
| 75 | COX6A2 | Cox6a2 | 12497 | -0.785 | -0.3058 | Yes | ||
| 76 | NDUFA3 | Ndufa3 | 12510 | -0.788 | -0.2968 | Yes | ||
| 77 | ATP5PD | Atp5h | 12515 | -0.789 | -0.2873 | Yes | ||
| 78 | ATP5F1A | Atp5a1 | 12609 | -0.813 | -0.2833 | Yes | ||
| 79 | ATP5F1E | Atp5e | 12626 | -0.818 | -0.2742 | Yes | ||
| 80 | NDUFS4 | Ndufs4 | 12711 | -0.837 | -0.2693 | Yes | ||
| 81 | NDUFB7 | Ndufb7 | 12713 | -0.838 | -0.2589 | Yes | ||
| 82 | ATP5MC3 | Atp5g3 | 12728 | -0.842 | -0.2494 | Yes | ||
| 83 | NDUFB10 | Ndufb10 | 12741 | -0.845 | -0.2397 | Yes | ||
| 84 | ATP5F1D | Atp5d | 12832 | -0.865 | -0.2349 | Yes | ||
| 85 | COX5B | Cox5b | 12858 | -0.875 | -0.2257 | Yes | ||
| 86 | ATP6V1G2 | Atp6v1g2 | 12874 | -0.880 | -0.2157 | Yes | ||
| 87 | ATP6V0D2 | Atp6v0d2 | 12918 | -0.895 | -0.2074 | Yes | ||
| 88 | COX8A | Cox8a | 12920 | -0.897 | -0.1964 | Yes | ||
| 89 | UQCRC1 | Uqcrc1 | 12932 | -0.902 | -0.1859 | Yes | ||
| 90 | NDUFA4 | Ndufa4 | 12975 | -0.914 | -0.1773 | Yes | ||
| 91 | UQCRB | Uqcrb | 12991 | -0.917 | -0.1669 | Yes | ||
| 92 | NDUFV3 | Ndufv3 | 13106 | -0.957 | -0.1625 | Yes | ||
| 93 | ATP5F1B | Atp5b | 13119 | -0.961 | -0.1514 | Yes | ||
| 94 | COX6C | Cox6c | 13135 | -0.967 | -0.1403 | Yes | ||
| 95 | ATP5ME | Atp5k | 13183 | -0.982 | -0.1312 | Yes | ||
| 96 | ATP5MF | Atp5j2 | 13185 | -0.984 | -0.1191 | Yes | ||
| 97 | COX6B2 | Cox6b2 | 13229 | -1.003 | -0.1095 | Yes | ||
| 98 | NDUFV2 | Ndufv2 | 13351 | -1.057 | -0.1042 | Yes | ||
| 99 | COX6A1 | Cox6a1 | 13416 | -1.080 | -0.0950 | Yes | ||
| 100 | COX10 | Cox10 | 13621 | -1.155 | -0.0939 | Yes | ||
| 101 | NDUFC1 | Ndufc1 | 13643 | -1.161 | -0.0809 | Yes | ||
| 102 | UQCR10 | Uqcr10 | 13754 | -1.211 | -0.0730 | Yes | ||
| 103 | ATP6V1C2 | Atp6v1c2 | 13993 | -1.331 | -0.0720 | Yes | ||
| 104 | NDUFS5 | Ndufs5 | 14012 | -1.341 | -0.0565 | Yes | ||
| 105 | NDUFA5 | Ndufa5 | 14340 | -1.554 | -0.0585 | Yes | ||
| 106 | COX7A1 | Cox7a1 | 14746 | -1.951 | -0.0606 | Yes | ||
| 107 | NDUFA4L2 | Ndufa4l2 | 14864 | -2.080 | -0.0424 | Yes | ||
| 108 | ATP6V0A4 | Atp6v0a4 | 14944 | -2.227 | -0.0199 | Yes | ||
| 109 | LHPP | Lhpp | 14986 | -2.306 | 0.0060 | Yes | ||
| 110 | PPA1 | Ppa1 | 15004 | -2.332 | 0.0338 | Yes |