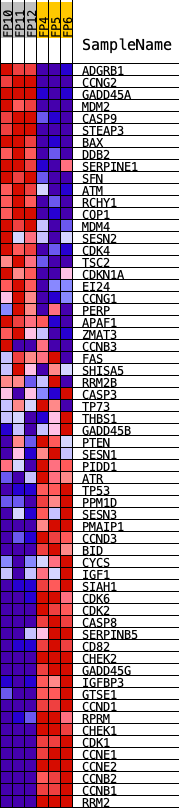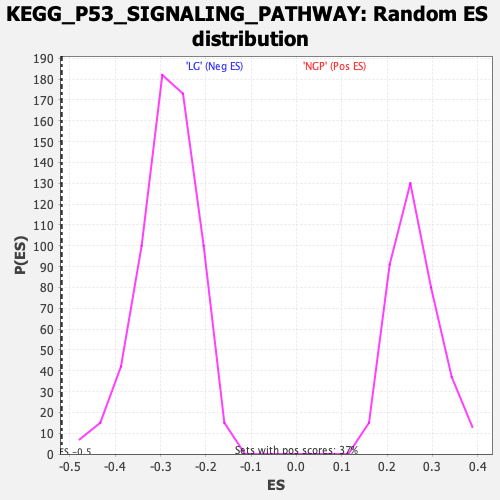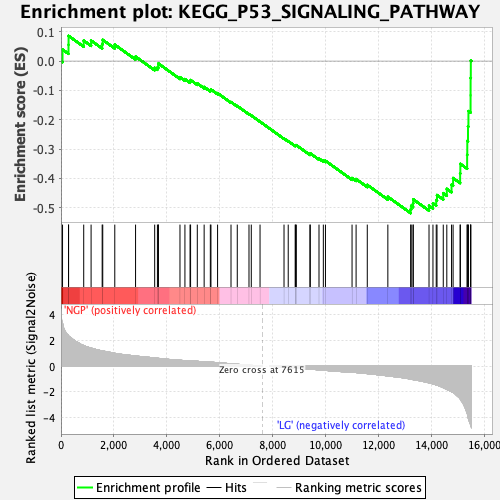
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_LG.phenotype.cls#NGP_versus_LG_repos |
| Phenotype | phenotype.cls#NGP_versus_LG_repos |
| Upregulated in class | LG |
| GeneSet | KEGG_P53_SIGNALING_PATHWAY |
| Enrichment Score (ES) | -0.5181166 |
| Normalized Enrichment Score (NES) | -1.8115513 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.009750378 |
| FWER p-Value | 0.092 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ADGRB1 | Adgrb1 | 58 | 3.326 | 0.0393 | No | ||
| 2 | CCNG2 | Ccng2 | 284 | 2.369 | 0.0554 | No | ||
| 3 | GADD45A | Gadd45a | 287 | 2.360 | 0.0858 | No | ||
| 4 | MDM2 | Mdm2 | 859 | 1.605 | 0.0697 | No | ||
| 5 | CASP9 | Casp9 | 1138 | 1.403 | 0.0699 | No | ||
| 6 | STEAP3 | Steap3 | 1557 | 1.192 | 0.0583 | No | ||
| 7 | BAX | Bax | 1579 | 1.182 | 0.0722 | No | ||
| 8 | DDB2 | Ddb2 | 2034 | 1.001 | 0.0558 | No | ||
| 9 | SERPINE1 | Serpine1 | 2820 | 0.790 | 0.0153 | No | ||
| 10 | SFN | Sfn | 3543 | 0.638 | -0.0231 | No | ||
| 11 | ATM | Atm | 3647 | 0.615 | -0.0218 | No | ||
| 12 | RCHY1 | Rchy1 | 3681 | 0.607 | -0.0161 | No | ||
| 13 | COP1 | Cop1 | 3682 | 0.607 | -0.0082 | No | ||
| 14 | MDM4 | Mdm4 | 4500 | 0.459 | -0.0551 | No | ||
| 15 | SESN2 | Sesn2 | 4687 | 0.435 | -0.0615 | No | ||
| 16 | CDK4 | Cdk4 | 4881 | 0.408 | -0.0687 | No | ||
| 17 | TSC2 | Tsc2 | 4901 | 0.402 | -0.0647 | No | ||
| 18 | CDKN1A | Cdkn1a | 5155 | 0.378 | -0.0762 | No | ||
| 19 | EI24 | Ei24 | 5414 | 0.355 | -0.0883 | No | ||
| 20 | CCNG1 | Ccng1 | 5648 | 0.314 | -0.0993 | No | ||
| 21 | PERP | Perp | 5671 | 0.311 | -0.0967 | No | ||
| 22 | APAF1 | Apaf1 | 5922 | 0.272 | -0.1093 | No | ||
| 23 | ZMAT3 | Zmat3 | 6430 | 0.188 | -0.1397 | No | ||
| 24 | CCNB3 | Ccnb3 | 6667 | 0.148 | -0.1531 | No | ||
| 25 | FAS | Fas | 7110 | 0.077 | -0.1806 | No | ||
| 26 | SHISA5 | Shisa5 | 7197 | 0.064 | -0.1854 | No | ||
| 27 | RRM2B | Rrm2b | 7528 | 0.015 | -0.2065 | No | ||
| 28 | CASP3 | Casp3 | 8435 | -0.085 | -0.2640 | No | ||
| 29 | TP73 | Trp73 | 8594 | -0.108 | -0.2728 | No | ||
| 30 | THBS1 | Thbs1 | 8857 | -0.147 | -0.2879 | No | ||
| 31 | GADD45B | Gadd45b | 8870 | -0.149 | -0.2867 | No | ||
| 32 | PTEN | Pten | 8901 | -0.154 | -0.2867 | No | ||
| 33 | SESN1 | Sesn1 | 9415 | -0.229 | -0.3169 | No | ||
| 34 | PIDD1 | Pidd1 | 9428 | -0.230 | -0.3147 | No | ||
| 35 | ATR | Atr | 9758 | -0.283 | -0.3323 | No | ||
| 36 | TP53 | Trp53 | 9922 | -0.312 | -0.3388 | No | ||
| 37 | PPM1D | Ppm1d | 10002 | -0.323 | -0.3397 | No | ||
| 38 | SESN3 | Sesn3 | 11008 | -0.471 | -0.3986 | No | ||
| 39 | PMAIP1 | Pmaip1 | 11161 | -0.493 | -0.4020 | No | ||
| 40 | CCND3 | Ccnd3 | 11584 | -0.575 | -0.4219 | No | ||
| 41 | BID | Bid | 12362 | -0.754 | -0.4624 | No | ||
| 42 | CYCS | Cycs | 13225 | -1.002 | -0.5051 | Yes | ||
| 43 | IGF1 | Igf1 | 13244 | -1.013 | -0.4932 | Yes | ||
| 44 | SIAH1 | Siah1a | 13316 | -1.042 | -0.4843 | Yes | ||
| 45 | CDK6 | Cdk6 | 13323 | -1.045 | -0.4711 | Yes | ||
| 46 | CDK2 | Cdk2 | 13919 | -1.295 | -0.4929 | Yes | ||
| 47 | CASP8 | Casp8 | 14068 | -1.374 | -0.4846 | Yes | ||
| 48 | SERPINB5 | Serpinb5 | 14192 | -1.456 | -0.4737 | Yes | ||
| 49 | CD82 | Cd82 | 14221 | -1.472 | -0.4565 | Yes | ||
| 50 | CHEK2 | Chek2 | 14458 | -1.668 | -0.4502 | Yes | ||
| 51 | GADD45G | Gadd45g | 14592 | -1.803 | -0.4354 | Yes | ||
| 52 | IGFBP3 | Igfbp3 | 14771 | -1.987 | -0.4212 | Yes | ||
| 53 | GTSE1 | Gtse1 | 14830 | -2.039 | -0.3986 | Yes | ||
| 54 | CCND1 | Ccnd1 | 15096 | -2.535 | -0.3829 | Yes | ||
| 55 | RPRM | Rprm | 15104 | -2.555 | -0.3503 | Yes | ||
| 56 | CHEK1 | Chek1 | 15361 | -3.652 | -0.3196 | Yes | ||
| 57 | CDK1 | Cdk1 | 15367 | -3.695 | -0.2721 | Yes | ||
| 58 | CCNE1 | Ccne1 | 15394 | -3.949 | -0.2226 | Yes | ||
| 59 | CCNE2 | Ccne2 | 15410 | -4.096 | -0.1706 | Yes | ||
| 60 | CCNB2 | Ccnb2 | 15491 | -4.557 | -0.1167 | Yes | ||
| 61 | CCNB1 | Ccnb1 | 15492 | -4.568 | -0.0576 | Yes | ||
| 62 | RRM2 | Rrm2 | 15500 | -4.610 | 0.0016 | Yes |