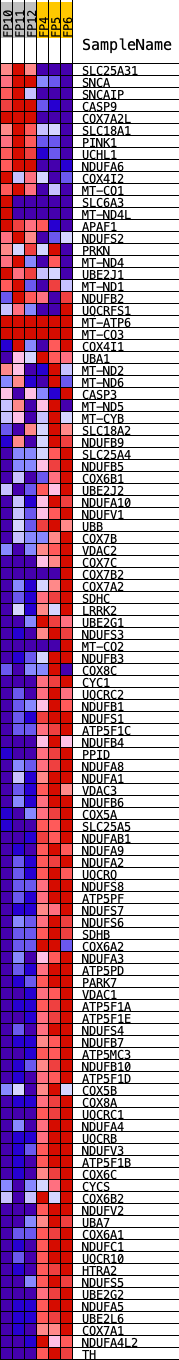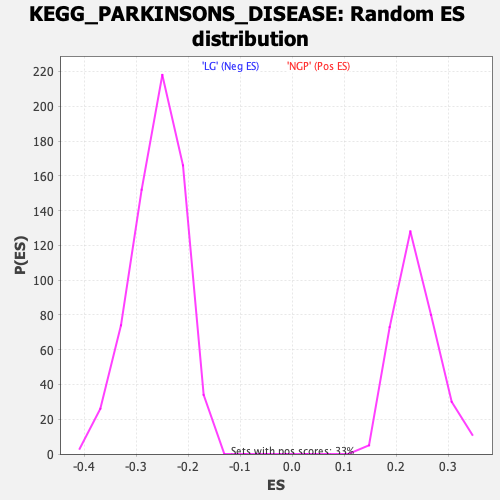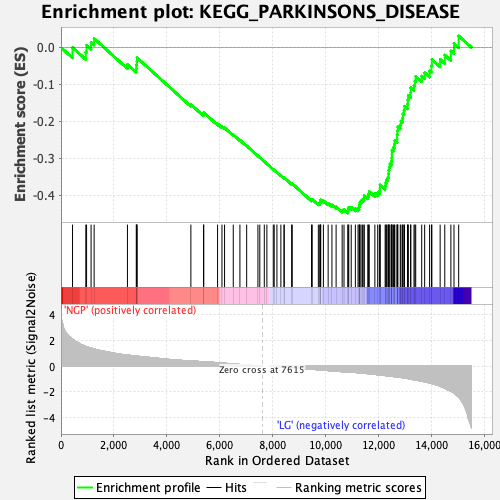
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_LG.phenotype.cls#NGP_versus_LG_repos |
| Phenotype | phenotype.cls#NGP_versus_LG_repos |
| Upregulated in class | LG |
| GeneSet | KEGG_PARKINSONS_DISEASE |
| Enrichment Score (ES) | -0.44820452 |
| Normalized Enrichment Score (NES) | -1.7317423 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.02326611 |
| FWER p-Value | 0.241 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SLC25A31 | Slc25a31 | 436 | 2.108 | -0.0002 | No | ||
| 2 | SNCA | Snca | 939 | 1.538 | -0.0123 | No | ||
| 3 | SNCAIP | Sncaip | 967 | 1.521 | 0.0062 | No | ||
| 4 | CASP9 | Casp9 | 1138 | 1.403 | 0.0139 | No | ||
| 5 | COX7A2L | Cox7a2l | 1253 | 1.335 | 0.0243 | No | ||
| 6 | SLC18A1 | Slc18a1 | 2513 | 0.864 | -0.0459 | No | ||
| 7 | PINK1 | Pink1 | 2848 | 0.785 | -0.0571 | No | ||
| 8 | UCHL1 | Uchl1 | 2854 | 0.783 | -0.0470 | No | ||
| 9 | NDUFA6 | Ndufa6 | 2870 | 0.780 | -0.0376 | No | ||
| 10 | COX4I2 | Cox4i2 | 2876 | 0.778 | -0.0275 | No | ||
| 11 | MT-CO1 | mt-Co1 | 4912 | 0.401 | -0.1542 | No | ||
| 12 | SLC6A3 | Slc6a3 | 5390 | 0.356 | -0.1804 | No | ||
| 13 | MT-ND4L | mt-Nd4l | 5400 | 0.356 | -0.1762 | No | ||
| 14 | APAF1 | Apaf1 | 5922 | 0.272 | -0.2064 | No | ||
| 15 | NDUFS2 | Ndufs2 | 6088 | 0.245 | -0.2139 | No | ||
| 16 | PRKN | Prkn | 6186 | 0.229 | -0.2171 | No | ||
| 17 | MT-ND4 | mt-Nd4 | 6515 | 0.173 | -0.2361 | No | ||
| 18 | UBE2J1 | Ube2j1 | 6767 | 0.130 | -0.2506 | No | ||
| 19 | MT-ND1 | mt-Nd1 | 7022 | 0.089 | -0.2659 | No | ||
| 20 | NDUFB2 | Ndufb2 | 7448 | 0.026 | -0.2931 | No | ||
| 21 | UQCRFS1 | Uqcrfs1 | 7519 | 0.017 | -0.2974 | No | ||
| 22 | MT-ATP6 | mt-Atp6 | 7689 | 0.000 | -0.3084 | No | ||
| 23 | MT-CO3 | mt-Co3 | 7787 | 0.000 | -0.3147 | No | ||
| 24 | COX4I1 | Cox4i1 | 8028 | -0.025 | -0.3299 | No | ||
| 25 | UBA1 | Uba1 | 8064 | -0.030 | -0.3318 | No | ||
| 26 | MT-ND2 | mt-Nd2 | 8167 | -0.045 | -0.3378 | No | ||
| 27 | MT-ND6 | mt-Nd6 | 8316 | -0.068 | -0.3465 | No | ||
| 28 | CASP3 | Casp3 | 8435 | -0.085 | -0.3530 | No | ||
| 29 | MT-ND5 | mt-Nd5 | 8443 | -0.086 | -0.3523 | No | ||
| 30 | MT-CYB | mt-Cytb | 8723 | -0.128 | -0.3687 | No | ||
| 31 | SLC18A2 | Slc18a2 | 8745 | -0.131 | -0.3684 | No | ||
| 32 | NDUFB9 | Ndufb9 | 9475 | -0.238 | -0.4125 | No | ||
| 33 | SLC25A4 | Slc25a4 | 9501 | -0.243 | -0.4109 | No | ||
| 34 | NDUFB5 | Ndufb5 | 9733 | -0.280 | -0.4221 | No | ||
| 35 | COX6B1 | Cox6b1 | 9788 | -0.286 | -0.4218 | No | ||
| 36 | UBE2J2 | Ube2j2 | 9805 | -0.290 | -0.4190 | No | ||
| 37 | NDUFA10 | Ndufa10 | 9814 | -0.293 | -0.4156 | No | ||
| 38 | NDUFV1 | Ndufv1 | 9824 | -0.293 | -0.4123 | No | ||
| 39 | UBB | Ubb | 9924 | -0.312 | -0.4145 | No | ||
| 40 | COX7B | Cox7b | 10104 | -0.341 | -0.4216 | No | ||
| 41 | VDAC2 | Vdac2 | 10247 | -0.371 | -0.4259 | No | ||
| 42 | COX7C | Cox7c | 10405 | -0.397 | -0.4308 | No | ||
| 43 | COX7B2 | Cox7b2 | 10634 | -0.421 | -0.4399 | No | ||
| 44 | COX7A2 | Cox7a2 | 10704 | -0.427 | -0.4387 | No | ||
| 45 | SDHC | Sdhc | 10851 | -0.443 | -0.4423 | Yes | ||
| 46 | LRRK2 | Lrrk2 | 10858 | -0.444 | -0.4368 | Yes | ||
| 47 | UBE2G1 | Ube2g1 | 10884 | -0.449 | -0.4324 | Yes | ||
| 48 | NDUFS3 | Ndufs3 | 10977 | -0.465 | -0.4322 | Yes | ||
| 49 | MT-CO2 | mt-Co2 | 11133 | -0.487 | -0.4358 | Yes | ||
| 50 | NDUFB3 | Ndufb3 | 11235 | -0.505 | -0.4356 | Yes | ||
| 51 | COX8C | Cox8b | 11277 | -0.512 | -0.4314 | Yes | ||
| 52 | CYC1 | Cyc1 | 11289 | -0.515 | -0.4253 | Yes | ||
| 53 | UQCRC2 | Uqcrc2 | 11303 | -0.518 | -0.4192 | Yes | ||
| 54 | NDUFB1 | Ndufb1-ps | 11347 | -0.526 | -0.4150 | Yes | ||
| 55 | NDUFS1 | Ndufs1 | 11400 | -0.535 | -0.4113 | Yes | ||
| 56 | ATP5F1C | Atp5c1 | 11459 | -0.548 | -0.4077 | Yes | ||
| 57 | NDUFB4 | Ndufb4 | 11466 | -0.550 | -0.4008 | Yes | ||
| 58 | PPID | Ppid | 11603 | -0.578 | -0.4019 | Yes | ||
| 59 | NDUFA8 | Ndufa8 | 11633 | -0.584 | -0.3960 | Yes | ||
| 60 | NDUFA1 | Ndufa1 | 11658 | -0.590 | -0.3897 | Yes | ||
| 61 | VDAC3 | Vdac3 | 11871 | -0.635 | -0.3950 | Yes | ||
| 62 | NDUFB6 | Ndufb6 | 11978 | -0.661 | -0.3931 | Yes | ||
| 63 | COX5A | Cox5a | 12040 | -0.676 | -0.3880 | Yes | ||
| 64 | SLC25A5 | Slc25a5 | 12066 | -0.682 | -0.3805 | Yes | ||
| 65 | NDUFAB1 | Ndufab1 | 12069 | -0.683 | -0.3716 | Yes | ||
| 66 | NDUFA9 | Ndufa9 | 12260 | -0.729 | -0.3742 | Yes | ||
| 67 | NDUFA2 | Ndufa2 | 12284 | -0.734 | -0.3659 | Yes | ||
| 68 | UQCRQ | Uqcrq | 12315 | -0.742 | -0.3580 | Yes | ||
| 69 | NDUFS8 | Ndufs8 | 12364 | -0.755 | -0.3510 | Yes | ||
| 70 | ATP5PF | Atp5j | 12388 | -0.760 | -0.3424 | Yes | ||
| 71 | NDUFS7 | Ndufs7 | 12392 | -0.763 | -0.3324 | Yes | ||
| 72 | NDUFS6 | Ndufs6 | 12410 | -0.765 | -0.3233 | Yes | ||
| 73 | SDHB | Sdhb | 12441 | -0.770 | -0.3150 | Yes | ||
| 74 | COX6A2 | Cox6a2 | 12497 | -0.785 | -0.3081 | Yes | ||
| 75 | NDUFA3 | Ndufa3 | 12510 | -0.788 | -0.2984 | Yes | ||
| 76 | ATP5PD | Atp5h | 12515 | -0.789 | -0.2882 | Yes | ||
| 77 | PARK7 | Park7 | 12517 | -0.790 | -0.2777 | Yes | ||
| 78 | VDAC1 | Vdac1 | 12573 | -0.806 | -0.2706 | Yes | ||
| 79 | ATP5F1A | Atp5a1 | 12609 | -0.813 | -0.2620 | Yes | ||
| 80 | ATP5F1E | Atp5e | 12626 | -0.818 | -0.2521 | Yes | ||
| 81 | NDUFS4 | Ndufs4 | 12711 | -0.837 | -0.2464 | Yes | ||
| 82 | NDUFB7 | Ndufb7 | 12713 | -0.838 | -0.2353 | Yes | ||
| 83 | ATP5MC3 | Atp5g3 | 12728 | -0.842 | -0.2250 | Yes | ||
| 84 | NDUFB10 | Ndufb10 | 12741 | -0.845 | -0.2146 | Yes | ||
| 85 | ATP5F1D | Atp5d | 12832 | -0.865 | -0.2089 | Yes | ||
| 86 | COX5B | Cox5b | 12858 | -0.875 | -0.1989 | Yes | ||
| 87 | COX8A | Cox8a | 12920 | -0.897 | -0.1909 | Yes | ||
| 88 | UQCRC1 | Uqcrc1 | 12932 | -0.902 | -0.1796 | Yes | ||
| 89 | NDUFA4 | Ndufa4 | 12975 | -0.914 | -0.1701 | Yes | ||
| 90 | UQCRB | Uqcrb | 12991 | -0.917 | -0.1589 | Yes | ||
| 91 | NDUFV3 | Ndufv3 | 13106 | -0.957 | -0.1535 | Yes | ||
| 92 | ATP5F1B | Atp5b | 13119 | -0.961 | -0.1415 | Yes | ||
| 93 | COX6C | Cox6c | 13135 | -0.967 | -0.1296 | Yes | ||
| 94 | CYCS | Cycs | 13225 | -1.002 | -0.1220 | Yes | ||
| 95 | COX6B2 | Cox6b2 | 13229 | -1.003 | -0.1089 | Yes | ||
| 96 | NDUFV2 | Ndufv2 | 13351 | -1.057 | -0.1026 | Yes | ||
| 97 | UBA7 | Uba7 | 13386 | -1.070 | -0.0906 | Yes | ||
| 98 | COX6A1 | Cox6a1 | 13416 | -1.080 | -0.0781 | Yes | ||
| 99 | NDUFC1 | Ndufc1 | 13643 | -1.161 | -0.0773 | Yes | ||
| 100 | UQCR10 | Uqcr10 | 13754 | -1.211 | -0.0683 | Yes | ||
| 101 | HTRA2 | Htra2 | 13941 | -1.305 | -0.0630 | Yes | ||
| 102 | NDUFS5 | Ndufs5 | 14012 | -1.341 | -0.0497 | Yes | ||
| 103 | UBE2G2 | Ube2g2 | 14027 | -1.350 | -0.0326 | Yes | ||
| 104 | NDUFA5 | Ndufa5 | 14340 | -1.554 | -0.0321 | Yes | ||
| 105 | UBE2L6 | Ube2l6 | 14515 | -1.727 | -0.0204 | Yes | ||
| 106 | COX7A1 | Cox7a1 | 14746 | -1.951 | -0.0093 | Yes | ||
| 107 | NDUFA4L2 | Ndufa4l2 | 14864 | -2.080 | 0.0108 | Yes | ||
| 108 | TH | Th | 15041 | -2.405 | 0.0314 | Yes |