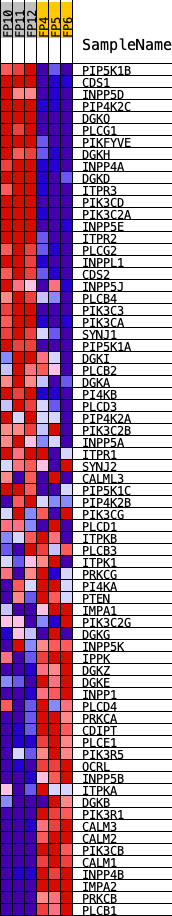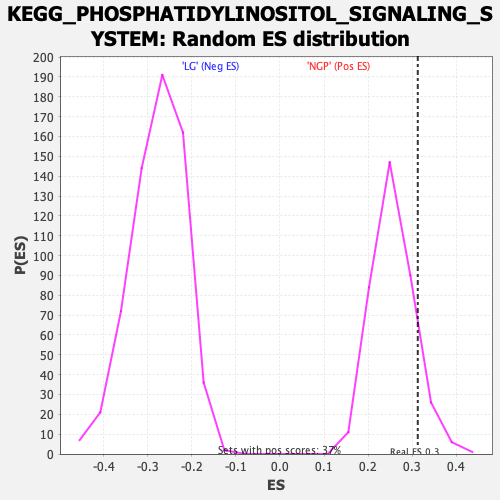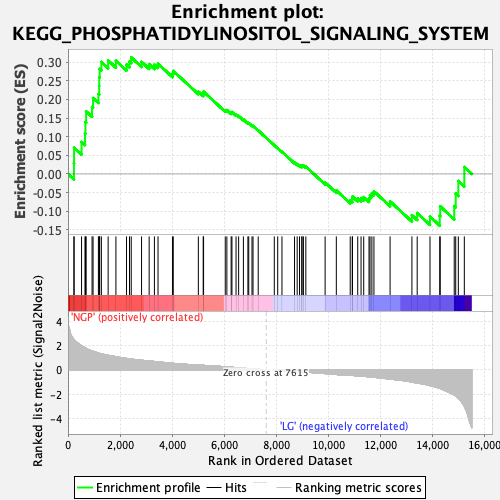
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_LG.phenotype.cls#NGP_versus_LG_repos |
| Phenotype | phenotype.cls#NGP_versus_LG_repos |
| Upregulated in class | NGP |
| GeneSet | KEGG_PHOSPHATIDYLINOSITOL_SIGNALING_SYSTEM |
| Enrichment Score (ES) | 0.31263858 |
| Normalized Enrichment Score (NES) | 1.2217058 |
| Nominal p-value | 0.12328767 |
| FDR q-value | 0.5966352 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PIP5K1B | Pip5k1b | 228 | 2.492 | 0.0282 | Yes | ||
| 2 | CDS1 | Cds1 | 234 | 2.470 | 0.0705 | Yes | ||
| 3 | INPP5D | Inpp5d | 521 | 1.970 | 0.0860 | Yes | ||
| 4 | PIP4K2C | Pip4k2c | 654 | 1.826 | 0.1090 | Yes | ||
| 5 | DGKQ | Dgkq | 671 | 1.808 | 0.1392 | Yes | ||
| 6 | PLCG1 | Plcg1 | 698 | 1.760 | 0.1678 | Yes | ||
| 7 | PIKFYVE | Pikfyve | 925 | 1.543 | 0.1799 | Yes | ||
| 8 | DGKH | Dgkh | 963 | 1.522 | 0.2037 | Yes | ||
| 9 | INPP4A | Inpp4a | 1169 | 1.391 | 0.2145 | Yes | ||
| 10 | DGKD | Dgkd | 1200 | 1.362 | 0.2360 | Yes | ||
| 11 | ITPR3 | Itpr3 | 1202 | 1.362 | 0.2594 | Yes | ||
| 12 | PIK3CD | Pik3cd | 1218 | 1.353 | 0.2818 | Yes | ||
| 13 | PIK3C2A | Pik3c2a | 1283 | 1.321 | 0.3005 | Yes | ||
| 14 | INPP5E | Inpp5e | 1543 | 1.197 | 0.3043 | Yes | ||
| 15 | ITPR2 | Itpr2 | 1842 | 1.079 | 0.3037 | Yes | ||
| 16 | PLCG2 | Plcg2 | 2250 | 0.934 | 0.2935 | Yes | ||
| 17 | INPPL1 | Inppl1 | 2365 | 0.899 | 0.3016 | Yes | ||
| 18 | CDS2 | Cds2 | 2431 | 0.883 | 0.3126 | Yes | ||
| 19 | INPP5J | Inpp5j | 2828 | 0.788 | 0.3006 | No | ||
| 20 | PLCB4 | Plcb4 | 3120 | 0.733 | 0.2944 | No | ||
| 21 | PIK3C3 | Pik3c3 | 3322 | 0.686 | 0.2933 | No | ||
| 22 | PIK3CA | Pik3ca | 3461 | 0.655 | 0.2956 | No | ||
| 23 | SYNJ1 | Synj1 | 4016 | 0.538 | 0.2691 | No | ||
| 24 | PIP5K1A | Pip5k1a | 4047 | 0.532 | 0.2763 | No | ||
| 25 | DGKI | Dgki | 5011 | 0.401 | 0.2209 | No | ||
| 26 | PLCB2 | Plcb2 | 5198 | 0.371 | 0.2153 | No | ||
| 27 | DGKA | Dgka | 5212 | 0.369 | 0.2208 | No | ||
| 28 | PI4KB | Pi4kb | 6046 | 0.251 | 0.1712 | No | ||
| 29 | PLCD3 | Plcd3 | 6108 | 0.241 | 0.1714 | No | ||
| 30 | PIP4K2A | Pip4k2a | 6272 | 0.213 | 0.1646 | No | ||
| 31 | PIK3C2B | Pik3c2b | 6303 | 0.207 | 0.1662 | No | ||
| 32 | INPP5A | Inpp5a | 6461 | 0.183 | 0.1592 | No | ||
| 33 | ITPR1 | Itpr1 | 6560 | 0.165 | 0.1557 | No | ||
| 34 | SYNJ2 | Synj2 | 6743 | 0.135 | 0.1463 | No | ||
| 35 | CALML3 | Calml3 | 6917 | 0.107 | 0.1369 | No | ||
| 36 | PIP5K1C | Pip5k1c | 6949 | 0.102 | 0.1367 | No | ||
| 37 | PIP4K2B | Pip4k2b | 7074 | 0.082 | 0.1301 | No | ||
| 38 | PIK3CG | Pik3cg | 7113 | 0.076 | 0.1289 | No | ||
| 39 | PLCD1 | Plcd1 | 7313 | 0.047 | 0.1168 | No | ||
| 40 | ITPKB | Itpkb | 7931 | -0.007 | 0.0770 | No | ||
| 41 | PLCB3 | Plcb3 | 8062 | -0.030 | 0.0691 | No | ||
| 42 | ITPK1 | Itpk1 | 8226 | -0.053 | 0.0595 | No | ||
| 43 | PRKCG | Prkcg | 8709 | -0.126 | 0.0305 | No | ||
| 44 | PI4KA | Pi4ka | 8811 | -0.143 | 0.0264 | No | ||
| 45 | PTEN | Pten | 8901 | -0.154 | 0.0233 | No | ||
| 46 | IMPA1 | Impa1 | 8989 | -0.167 | 0.0206 | No | ||
| 47 | PIK3C2G | Pik3c2g | 8996 | -0.167 | 0.0231 | No | ||
| 48 | DGKG | Dgkg | 9051 | -0.172 | 0.0225 | No | ||
| 49 | INPP5K | Inpp5k | 9144 | -0.186 | 0.0198 | No | ||
| 50 | IPPK | Ippk | 9885 | -0.305 | -0.0228 | No | ||
| 51 | DGKZ | Dgkz | 10316 | -0.382 | -0.0441 | No | ||
| 52 | DGKE | Dgke | 10850 | -0.443 | -0.0709 | No | ||
| 53 | INPP1 | Inpp1 | 10928 | -0.458 | -0.0680 | No | ||
| 54 | PLCD4 | Plcd4 | 10937 | -0.460 | -0.0606 | No | ||
| 55 | PRKCA | Prkca | 11139 | -0.487 | -0.0652 | No | ||
| 56 | CDIPT | Cdipt | 11267 | -0.510 | -0.0646 | No | ||
| 57 | PLCE1 | Plce1 | 11369 | -0.530 | -0.0620 | No | ||
| 58 | PIK3R5 | Pik3r5 | 11573 | -0.572 | -0.0652 | No | ||
| 59 | OCRL | Ocrl | 11611 | -0.581 | -0.0576 | No | ||
| 60 | INPP5B | Inpp5b | 11683 | -0.595 | -0.0519 | No | ||
| 61 | ITPKA | Itpka | 11762 | -0.611 | -0.0465 | No | ||
| 62 | DGKB | Dgkb | 12383 | -0.758 | -0.0735 | No | ||
| 63 | PIK3R1 | Pik3r1 | 13222 | -1.001 | -0.1104 | No | ||
| 64 | CALM3 | Calm3 | 13425 | -1.084 | -0.1048 | No | ||
| 65 | CALM2 | Calm2 | 13917 | -1.295 | -0.1142 | No | ||
| 66 | PIK3CB | Pik3cb | 14292 | -1.520 | -0.1122 | No | ||
| 67 | CALM1 | Calm1 | 14310 | -1.533 | -0.0869 | No | ||
| 68 | INPP4B | Inpp4b | 14849 | -2.062 | -0.0861 | No | ||
| 69 | IMPA2 | Impa2 | 14906 | -2.149 | -0.0527 | No | ||
| 70 | PRKCB | Prkcb | 15007 | -2.336 | -0.0188 | No | ||
| 71 | PLCB1 | Plcb1 | 15237 | -3.030 | 0.0186 | No |