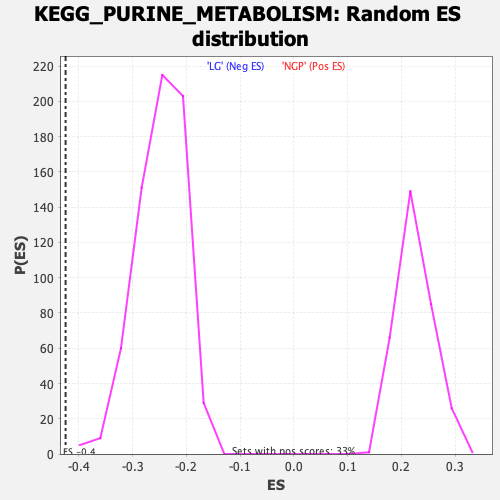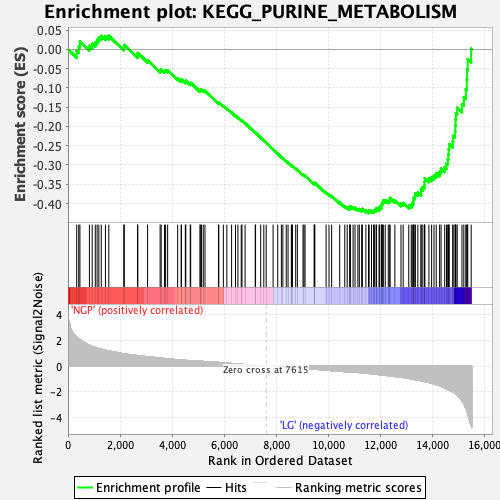
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_LG.phenotype.cls#NGP_versus_LG_repos |
| Phenotype | phenotype.cls#NGP_versus_LG_repos |
| Upregulated in class | LG |
| GeneSet | KEGG_PURINE_METABOLISM |
| Enrichment Score (ES) | -0.424942 |
| Normalized Enrichment Score (NES) | -1.7113243 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.026973516 |
| FWER p-Value | 0.298 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PDE3A | Pde3a | 335 | 2.266 | -0.0043 | No | ||
| 2 | PDE8A | Pde8a | 413 | 2.143 | 0.0072 | No | ||
| 3 | PDE6B | Pde6b | 458 | 2.079 | 0.0204 | No | ||
| 4 | AMPD3 | Ampd3 | 823 | 1.637 | 0.0093 | No | ||
| 5 | PDE5A | Pde5a | 930 | 1.540 | 0.0143 | No | ||
| 6 | GUCY1A2 | Gucy1a2 | 1049 | 1.461 | 0.0179 | No | ||
| 7 | ADCY1 | Adcy1 | 1119 | 1.417 | 0.0243 | No | ||
| 8 | NPR1 | Npr1 | 1186 | 1.375 | 0.0306 | No | ||
| 9 | POLR2A | Polr2a | 1279 | 1.322 | 0.0349 | No | ||
| 10 | PDE9A | Pde9a | 1439 | 1.241 | 0.0341 | No | ||
| 11 | ALLC | Allc | 1569 | 1.187 | 0.0348 | No | ||
| 12 | POLR3A | Polr3a | 2144 | 0.970 | 0.0050 | No | ||
| 13 | ADPRM | Adprm | 2164 | 0.962 | 0.0112 | No | ||
| 14 | POLR1C | Polr1c | 2676 | 0.827 | -0.0156 | No | ||
| 15 | NPR2 | Npr2 | 2682 | 0.825 | -0.0096 | No | ||
| 16 | ENTPD3 | Entpd3 | 3060 | 0.741 | -0.0284 | No | ||
| 17 | NUDT9 | Nudt9 | 3542 | 0.638 | -0.0548 | No | ||
| 18 | ATIC | Atic | 3582 | 0.630 | -0.0524 | No | ||
| 19 | AK4 | Ak4 | 3711 | 0.602 | -0.0561 | No | ||
| 20 | ADCY5 | Adcy5 | 3753 | 0.589 | -0.0542 | No | ||
| 21 | PRUNE1 | Prune1 | 3831 | 0.571 | -0.0548 | No | ||
| 22 | PRPS1L1 | Prps1l1 | 4216 | 0.500 | -0.0759 | No | ||
| 23 | GUCY2C | Gucy2c | 4352 | 0.480 | -0.0810 | No | ||
| 24 | ENTPD8 | Entpd8 | 4359 | 0.479 | -0.0777 | No | ||
| 25 | POLR3F | Polr3f | 4510 | 0.458 | -0.0839 | No | ||
| 26 | NME5 | Nme5 | 4530 | 0.454 | -0.0817 | No | ||
| 27 | PDE4A | Pde4a | 4699 | 0.434 | -0.0893 | No | ||
| 28 | NT5C | Nt5c | 4717 | 0.430 | -0.0870 | No | ||
| 29 | DGUOK | Dguok | 5078 | 0.390 | -0.1074 | No | ||
| 30 | CANT1 | Cant1 | 5083 | 0.390 | -0.1047 | No | ||
| 31 | PDE10A | Pde10a | 5131 | 0.382 | -0.1048 | No | ||
| 32 | ADCY6 | Adcy6 | 5199 | 0.371 | -0.1063 | No | ||
| 33 | PDE6G | Pde6g | 5266 | 0.359 | -0.1078 | No | ||
| 34 | NME4 | Nme4 | 5785 | 0.293 | -0.1392 | No | ||
| 35 | AK1 | Ak1 | 5798 | 0.291 | -0.1378 | No | ||
| 36 | PKM | Pkm | 5973 | 0.264 | -0.1471 | No | ||
| 37 | AMPD2 | Ampd2 | 6107 | 0.241 | -0.1538 | No | ||
| 38 | PDE11A | Pde11a | 6290 | 0.209 | -0.1641 | No | ||
| 39 | POLR3C | Polr3c | 6439 | 0.187 | -0.1722 | No | ||
| 40 | POLR2K | Polr2k | 6533 | 0.170 | -0.1770 | No | ||
| 41 | POLR3B | Polr3b | 6660 | 0.148 | -0.1840 | No | ||
| 42 | AK7 | Ak7 | 6689 | 0.146 | -0.1847 | No | ||
| 43 | ADA | Ada | 6808 | 0.124 | -0.1914 | No | ||
| 44 | ADCY2 | Adcy2 | 7198 | 0.064 | -0.2162 | No | ||
| 45 | POLR1D | Polr1d | 7211 | 0.062 | -0.2165 | No | ||
| 46 | PDE1B | Pde1b | 7408 | 0.032 | -0.2290 | No | ||
| 47 | RRM2B | Rrm2b | 7528 | 0.015 | -0.2366 | No | ||
| 48 | PKLR | Pklr | 7619 | 0.000 | -0.2425 | No | ||
| 49 | POLR2E | Polr2e | 7885 | -0.000 | -0.2597 | No | ||
| 50 | PRPS2 | Prps2 | 8063 | -0.030 | -0.2710 | No | ||
| 51 | PDE2A | Gm45837 | 8203 | -0.048 | -0.2797 | No | ||
| 52 | GUCY1A1 | Gucy1a1 | 8261 | -0.059 | -0.2829 | No | ||
| 53 | NUDT5 | Nudt5 | 8390 | -0.077 | -0.2906 | No | ||
| 54 | PDE7A | Pde7a | 8460 | -0.088 | -0.2944 | No | ||
| 55 | PAICS | Paics | 8586 | -0.106 | -0.3018 | No | ||
| 56 | POLR2B | Polr2b | 8603 | -0.111 | -0.3019 | No | ||
| 57 | AMPD1 | Ampd1 | 8630 | -0.112 | -0.3028 | No | ||
| 58 | POLE4 | Pole4 | 8754 | -0.132 | -0.3097 | No | ||
| 59 | POLR3D | Polr3d | 8818 | -0.144 | -0.3127 | No | ||
| 60 | GUCY2F | Gucy2f | 9036 | -0.171 | -0.3255 | No | ||
| 61 | PRPS1 | Prps1 | 9072 | -0.175 | -0.3264 | No | ||
| 62 | ENTPD5 | Entpd5 | 9114 | -0.181 | -0.3277 | No | ||
| 63 | PDE6D | Pde6d | 9462 | -0.236 | -0.3484 | No | ||
| 64 | ADCY10 | Adcy10 | 9477 | -0.238 | -0.3475 | No | ||
| 65 | GUCY1B1 | Gucy1b1 | 9490 | -0.241 | -0.3464 | No | ||
| 66 | NME7 | Nme7 | 9923 | -0.312 | -0.3721 | No | ||
| 67 | PDE6A | Pde6a | 10035 | -0.328 | -0.3768 | No | ||
| 68 | POLR2H | Polr2h | 10136 | -0.349 | -0.3806 | No | ||
| 69 | ADCY3 | Adcy3 | 10447 | -0.403 | -0.3977 | No | ||
| 70 | GUCY2D | Gucy2e | 10641 | -0.421 | -0.4070 | No | ||
| 71 | ENTPD4 | Entpd4b | 10733 | -0.432 | -0.4096 | No | ||
| 72 | POLR1A | Polr1a | 10830 | -0.440 | -0.4124 | No | ||
| 73 | ENTPD2 | Entpd2 | 10852 | -0.443 | -0.4103 | No | ||
| 74 | POLR2G | Polr2g | 10854 | -0.443 | -0.4070 | No | ||
| 75 | NUDT2 | Nudt2 | 10960 | -0.463 | -0.4102 | No | ||
| 76 | ZNRD1 | Znrd1 | 11035 | -0.476 | -0.4114 | No | ||
| 77 | POLR2I | Polr2i | 11145 | -0.489 | -0.4147 | No | ||
| 78 | ADCY9 | Adcy9 | 11200 | -0.498 | -0.4144 | No | ||
| 79 | NT5C2 | Nt5c2 | 11291 | -0.515 | -0.4163 | No | ||
| 80 | NT5C3A | Nt5c3 | 11318 | -0.519 | -0.4139 | No | ||
| 81 | APRT | Aprt | 11453 | -0.546 | -0.4184 | No | ||
| 82 | ADCY4 | Adcy4 | 11554 | -0.569 | -0.4206 | Yes | ||
| 83 | POLR3K | Polr3k | 11577 | -0.573 | -0.4176 | Yes | ||
| 84 | POLR1E | Polr1e | 11668 | -0.593 | -0.4189 | Yes | ||
| 85 | POLR2F | Polr2f | 11750 | -0.606 | -0.4194 | Yes | ||
| 86 | NME1 | Nme1 | 11779 | -0.614 | -0.4165 | Yes | ||
| 87 | POLR2C | Polr2c | 11845 | -0.629 | -0.4159 | Yes | ||
| 88 | IMPDH1 | Impdh1 | 11858 | -0.632 | -0.4118 | Yes | ||
| 89 | PNP | Pnp | 11964 | -0.658 | -0.4136 | Yes | ||
| 90 | PDE6H | Pde6h | 11966 | -0.658 | -0.4086 | Yes | ||
| 91 | AK5 | Ak5 | 12046 | -0.677 | -0.4085 | Yes | ||
| 92 | PDE7B | Pde7b | 12049 | -0.678 | -0.4034 | Yes | ||
| 93 | NME6 | Nme6 | 12074 | -0.684 | -0.3997 | Yes | ||
| 94 | ADCY8 | Adcy8 | 12101 | -0.691 | -0.3960 | Yes | ||
| 95 | FHIT | Fhit | 12121 | -0.698 | -0.3919 | Yes | ||
| 96 | IMPDH2 | Impdh2 | 12196 | -0.712 | -0.3912 | Yes | ||
| 97 | POLR2L | Polr2l | 12313 | -0.742 | -0.3930 | Yes | ||
| 98 | PAPSS2 | Papss2 | 12374 | -0.756 | -0.3911 | Yes | ||
| 99 | NME2 | Nme2 | 12378 | -0.757 | -0.3855 | Yes | ||
| 100 | ENTPD6 | Entpd6 | 12567 | -0.804 | -0.3915 | Yes | ||
| 101 | PFAS | Pfas | 12802 | -0.861 | -0.4001 | Yes | ||
| 102 | GUK1 | Guk1 | 12886 | -0.885 | -0.3986 | Yes | ||
| 103 | PDE3B | Pde3b | 13101 | -0.955 | -0.4052 | Yes | ||
| 104 | HPRT1 | Hprt | 13196 | -0.988 | -0.4037 | Yes | ||
| 105 | NT5M | Nt5m | 13236 | -1.007 | -0.3985 | Yes | ||
| 106 | ADSS2 | Adss | 13280 | -1.029 | -0.3933 | Yes | ||
| 107 | GMPR2 | Gmpr2 | 13287 | -1.031 | -0.3858 | Yes | ||
| 108 | PDE1C | Pde1c | 13335 | -1.053 | -0.3807 | Yes | ||
| 109 | XDH | Xdh | 13354 | -1.058 | -0.3737 | Yes | ||
| 110 | GART | Gart | 13450 | -1.091 | -0.3715 | Yes | ||
| 111 | POLR2D | Polr2d | 13573 | -1.136 | -0.3707 | Yes | ||
| 112 | ADSL | Adsl | 13574 | -1.137 | -0.3619 | Yes | ||
| 113 | GMPS | Gmps | 13642 | -1.161 | -0.3573 | Yes | ||
| 114 | ENTPD1 | Entpd1 | 13706 | -1.190 | -0.3522 | Yes | ||
| 115 | POLR1B | Polr1b | 13708 | -1.191 | -0.3431 | Yes | ||
| 116 | ADK | Adk | 13719 | -1.200 | -0.3345 | Yes | ||
| 117 | PNPT1 | Pnpt1 | 13876 | -1.270 | -0.3349 | Yes | ||
| 118 | ADSS1 | Adssl1 | 13980 | -1.325 | -0.3314 | Yes | ||
| 119 | PDE4D | Pde4d | 14071 | -1.378 | -0.3266 | Yes | ||
| 120 | GMPR | Gmpr | 14157 | -1.433 | -0.3211 | Yes | ||
| 121 | POLD1 | Pold1 | 14287 | -1.514 | -0.3178 | Yes | ||
| 122 | GDA | Gda | 14343 | -1.556 | -0.3094 | Yes | ||
| 123 | POLR3H | Polr3h | 14483 | -1.689 | -0.3054 | Yes | ||
| 124 | POLE3 | Pole3 | 14552 | -1.771 | -0.2961 | Yes | ||
| 125 | ENPP1 | Enpp1 | 14600 | -1.813 | -0.2852 | Yes | ||
| 126 | PAPSS1 | Papss1 | 14623 | -1.838 | -0.2725 | Yes | ||
| 127 | PDE1A | Pde1a | 14637 | -1.847 | -0.2591 | Yes | ||
| 128 | ADCY7 | Adcy7 | 14656 | -1.864 | -0.2459 | Yes | ||
| 129 | ITPA | Itpa | 14791 | -2.005 | -0.2391 | Yes | ||
| 130 | POLD2 | Pold2 | 14805 | -2.019 | -0.2244 | Yes | ||
| 131 | AK2 | Ak2 | 14878 | -2.105 | -0.2129 | Yes | ||
| 132 | POLE2 | Pole2 | 14892 | -2.125 | -0.1973 | Yes | ||
| 133 | NT5E | Nt5e | 14899 | -2.139 | -0.1813 | Yes | ||
| 134 | PPAT | Ppat | 14915 | -2.162 | -0.1656 | Yes | ||
| 135 | PDE4B | Pde4b | 14963 | -2.263 | -0.1512 | Yes | ||
| 136 | PRIM2 | Prim2 | 15147 | -2.715 | -0.1421 | Yes | ||
| 137 | DCK | Dck | 15218 | -2.933 | -0.1241 | Yes | ||
| 138 | RRM1 | Rrm1 | 15294 | -3.282 | -0.1037 | Yes | ||
| 139 | POLA1 | Pola1 | 15332 | -3.491 | -0.0792 | Yes | ||
| 140 | POLE | Pole | 15345 | -3.558 | -0.0525 | Yes | ||
| 141 | ENPP3 | Enpp3 | 15369 | -3.703 | -0.0255 | Yes | ||
| 142 | RRM2 | Rrm2 | 15500 | -4.610 | 0.0016 | Yes |