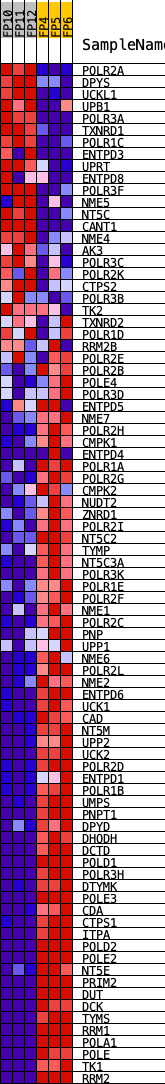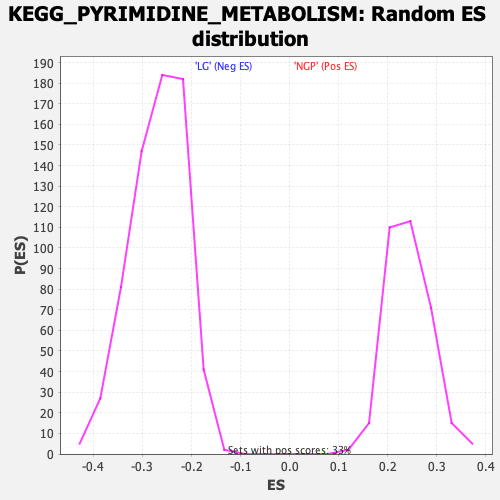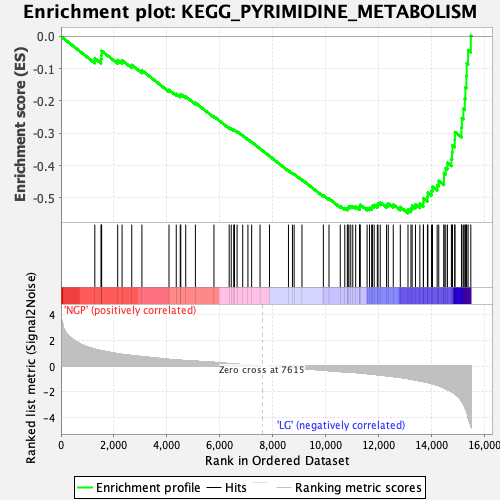
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_LG.phenotype.cls#NGP_versus_LG_repos |
| Phenotype | phenotype.cls#NGP_versus_LG_repos |
| Upregulated in class | LG |
| GeneSet | KEGG_PYRIMIDINE_METABOLISM |
| Enrichment Score (ES) | -0.547432 |
| Normalized Enrichment Score (NES) | -2.0411935 |
| Nominal p-value | 0.0 |
| FDR q-value | 3.0200157E-4 |
| FWER p-Value | 0.002 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | POLR2A | Polr2a | 1279 | 1.322 | -0.0683 | No | ||
| 2 | DPYS | Dpys | 1514 | 1.209 | -0.0701 | No | ||
| 3 | UCKL1 | Uckl1 | 1530 | 1.202 | -0.0578 | No | ||
| 4 | UPB1 | Upb1 | 1537 | 1.198 | -0.0450 | No | ||
| 5 | POLR3A | Polr3a | 2144 | 0.970 | -0.0735 | No | ||
| 6 | TXNRD1 | Txnrd1 | 2310 | 0.914 | -0.0741 | No | ||
| 7 | POLR1C | Polr1c | 2676 | 0.827 | -0.0887 | No | ||
| 8 | ENTPD3 | Entpd3 | 3060 | 0.741 | -0.1053 | No | ||
| 9 | UPRT | Uprt | 4085 | 0.525 | -0.1658 | No | ||
| 10 | ENTPD8 | Entpd8 | 4359 | 0.479 | -0.1782 | No | ||
| 11 | POLR3F | Polr3f | 4510 | 0.458 | -0.1829 | No | ||
| 12 | NME5 | Nme5 | 4530 | 0.454 | -0.1791 | No | ||
| 13 | NT5C | Nt5c | 4717 | 0.430 | -0.1864 | No | ||
| 14 | CANT1 | Cant1 | 5083 | 0.390 | -0.2058 | No | ||
| 15 | NME4 | Nme4 | 5785 | 0.293 | -0.2479 | No | ||
| 16 | AK3 | Ak3 | 6361 | 0.197 | -0.2830 | No | ||
| 17 | POLR3C | Polr3c | 6439 | 0.187 | -0.2859 | No | ||
| 18 | POLR2K | Polr2k | 6533 | 0.170 | -0.2901 | No | ||
| 19 | CTPS2 | Ctps2 | 6562 | 0.165 | -0.2901 | No | ||
| 20 | POLR3B | Polr3b | 6660 | 0.148 | -0.2947 | No | ||
| 21 | TK2 | Tk2 | 6873 | 0.113 | -0.3072 | No | ||
| 22 | TXNRD2 | Txnrd2 | 7070 | 0.082 | -0.3190 | No | ||
| 23 | POLR1D | Polr1d | 7211 | 0.062 | -0.3274 | No | ||
| 24 | RRM2B | Rrm2b | 7528 | 0.015 | -0.3477 | No | ||
| 25 | POLR2E | Polr2e | 7885 | -0.000 | -0.3707 | No | ||
| 26 | POLR2B | Polr2b | 8603 | -0.111 | -0.4159 | No | ||
| 27 | POLE4 | Pole4 | 8754 | -0.132 | -0.4242 | No | ||
| 28 | POLR3D | Polr3d | 8818 | -0.144 | -0.4267 | No | ||
| 29 | ENTPD5 | Entpd5 | 9114 | -0.181 | -0.4438 | No | ||
| 30 | NME7 | Nme7 | 9923 | -0.312 | -0.4927 | No | ||
| 31 | POLR2H | Polr2h | 10136 | -0.349 | -0.5026 | No | ||
| 32 | CMPK1 | Cmpk1 | 10562 | -0.411 | -0.5256 | No | ||
| 33 | ENTPD4 | Entpd4b | 10733 | -0.432 | -0.5318 | No | ||
| 34 | POLR1A | Polr1a | 10830 | -0.440 | -0.5332 | No | ||
| 35 | POLR2G | Polr2g | 10854 | -0.443 | -0.5298 | No | ||
| 36 | CMPK2 | Cmpk2 | 10890 | -0.450 | -0.5271 | No | ||
| 37 | NUDT2 | Nudt2 | 10960 | -0.463 | -0.5264 | No | ||
| 38 | ZNRD1 | Znrd1 | 11035 | -0.476 | -0.5260 | No | ||
| 39 | POLR2I | Polr2i | 11145 | -0.489 | -0.5277 | No | ||
| 40 | NT5C2 | Nt5c2 | 11291 | -0.515 | -0.5314 | No | ||
| 41 | TYMP | Tymp | 11295 | -0.516 | -0.5259 | No | ||
| 42 | NT5C3A | Nt5c3 | 11318 | -0.519 | -0.5216 | No | ||
| 43 | POLR3K | Polr3k | 11577 | -0.573 | -0.5320 | No | ||
| 44 | POLR1E | Polr1e | 11668 | -0.593 | -0.5313 | No | ||
| 45 | POLR2F | Polr2f | 11750 | -0.606 | -0.5298 | No | ||
| 46 | NME1 | Nme1 | 11779 | -0.614 | -0.5249 | No | ||
| 47 | POLR2C | Polr2c | 11845 | -0.629 | -0.5221 | No | ||
| 48 | PNP | Pnp | 11964 | -0.658 | -0.5225 | No | ||
| 49 | UPP1 | Upp1 | 11990 | -0.663 | -0.5168 | No | ||
| 50 | NME6 | Nme6 | 12074 | -0.684 | -0.5147 | No | ||
| 51 | POLR2L | Polr2l | 12313 | -0.742 | -0.5219 | No | ||
| 52 | NME2 | Nme2 | 12378 | -0.757 | -0.5177 | No | ||
| 53 | ENTPD6 | Entpd6 | 12567 | -0.804 | -0.5210 | No | ||
| 54 | UCK1 | Uck1 | 12833 | -0.865 | -0.5287 | No | ||
| 55 | CAD | Cad | 13124 | -0.964 | -0.5368 | Yes | ||
| 56 | NT5M | Nt5m | 13236 | -1.007 | -0.5329 | Yes | ||
| 57 | UPP2 | Upp2 | 13281 | -1.029 | -0.5244 | Yes | ||
| 58 | UCK2 | Uck2 | 13407 | -1.077 | -0.5206 | Yes | ||
| 59 | POLR2D | Polr2d | 13573 | -1.136 | -0.5188 | Yes | ||
| 60 | ENTPD1 | Entpd1 | 13706 | -1.190 | -0.5142 | Yes | ||
| 61 | POLR1B | Polr1b | 13708 | -1.191 | -0.5012 | Yes | ||
| 62 | UMPS | Umps | 13859 | -1.260 | -0.4970 | Yes | ||
| 63 | PNPT1 | Pnpt1 | 13876 | -1.270 | -0.4840 | Yes | ||
| 64 | DPYD | Dpyd | 14014 | -1.341 | -0.4781 | Yes | ||
| 65 | DHODH | Dhodh | 14051 | -1.364 | -0.4654 | Yes | ||
| 66 | DCTD | Dctd | 14227 | -1.477 | -0.4604 | Yes | ||
| 67 | POLD1 | Pold1 | 14287 | -1.514 | -0.4476 | Yes | ||
| 68 | POLR3H | Polr3h | 14483 | -1.689 | -0.4416 | Yes | ||
| 69 | DTYMK | Dtymk | 14485 | -1.692 | -0.4230 | Yes | ||
| 70 | POLE3 | Pole3 | 14552 | -1.771 | -0.4077 | Yes | ||
| 71 | CDA | Cda | 14616 | -1.831 | -0.3916 | Yes | ||
| 72 | CTPS1 | Ctps | 14766 | -1.979 | -0.3795 | Yes | ||
| 73 | ITPA | Itpa | 14791 | -2.005 | -0.3589 | Yes | ||
| 74 | POLD2 | Pold2 | 14805 | -2.019 | -0.3375 | Yes | ||
| 75 | POLE2 | Pole2 | 14892 | -2.125 | -0.3196 | Yes | ||
| 76 | NT5E | Nt5e | 14899 | -2.139 | -0.2964 | Yes | ||
| 77 | PRIM2 | Prim2 | 15147 | -2.715 | -0.2825 | Yes | ||
| 78 | DUT | Dut | 15162 | -2.753 | -0.2531 | Yes | ||
| 79 | DCK | Dck | 15218 | -2.933 | -0.2243 | Yes | ||
| 80 | TYMS | Tyms | 15273 | -3.185 | -0.1927 | Yes | ||
| 81 | RRM1 | Rrm1 | 15294 | -3.282 | -0.1578 | Yes | ||
| 82 | POLA1 | Pola1 | 15332 | -3.491 | -0.1217 | Yes | ||
| 83 | POLE | Pole | 15345 | -3.558 | -0.0832 | Yes | ||
| 84 | TK1 | Tk1 | 15397 | -3.985 | -0.0426 | Yes | ||
| 85 | RRM2 | Rrm2 | 15500 | -4.610 | 0.0016 | Yes |