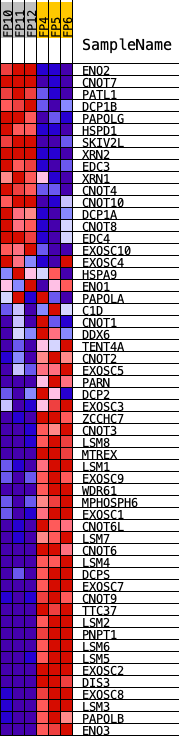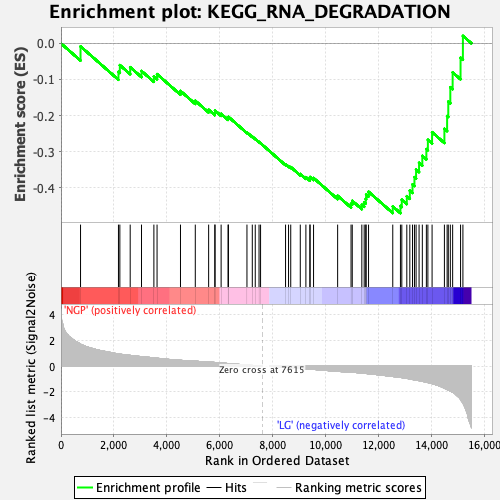
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_LG.phenotype.cls#NGP_versus_LG_repos |
| Phenotype | phenotype.cls#NGP_versus_LG_repos |
| Upregulated in class | LG |
| GeneSet | KEGG_RNA_DEGRADATION |
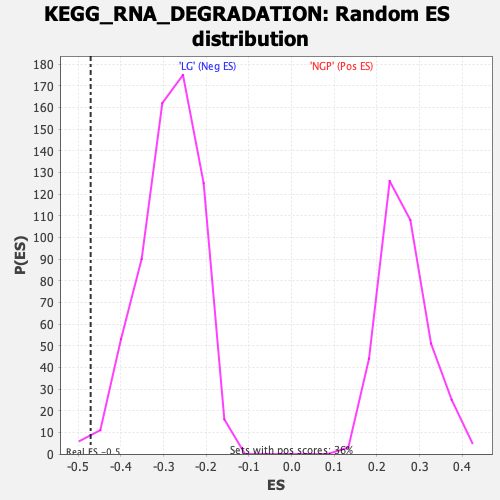
| Enrichment Score (ES) | -0.47082466 |
| Normalized Enrichment Score (NES) | -1.6395197 |
| Nominal p-value | 0.009404388 |
| FDR q-value | 0.051017795 |
| FWER p-Value | 0.533 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ENO2 | Eno2 | 739 | 1.710 | -0.0082 | No | ||
| 2 | CNOT7 | Cnot7 | 2173 | 0.959 | -0.0787 | No | ||
| 3 | PATL1 | Patl1 | 2226 | 0.945 | -0.0602 | No | ||
| 4 | DCP1B | Dcp1b | 2617 | 0.839 | -0.0660 | No | ||
| 5 | PAPOLG | Papolg | 3045 | 0.743 | -0.0764 | No | ||
| 6 | HSPD1 | Hspd1 | 3513 | 0.643 | -0.0918 | No | ||
| 7 | SKIV2L | Skiv2l | 3633 | 0.617 | -0.0852 | No | ||
| 8 | XRN2 | Xrn2 | 4519 | 0.456 | -0.1318 | No | ||
| 9 | EDC3 | Edc3 | 5079 | 0.390 | -0.1590 | No | ||
| 10 | XRN1 | Xrn1 | 5584 | 0.326 | -0.1840 | No | ||
| 11 | CNOT4 | Cnot4 | 5818 | 0.288 | -0.1924 | No | ||
| 12 | CNOT10 | Cnot10 | 5826 | 0.287 | -0.1862 | No | ||
| 13 | DCP1A | Dcp1a | 6056 | 0.249 | -0.1953 | No | ||
| 14 | CNOT8 | Cnot8 | 6318 | 0.204 | -0.2074 | No | ||
| 15 | EDC4 | Edc4 | 6333 | 0.202 | -0.2037 | No | ||
| 16 | EXOSC10 | Exosc10 | 7033 | 0.087 | -0.2468 | No | ||
| 17 | EXOSC4 | Exosc4 | 7235 | 0.058 | -0.2585 | No | ||
| 18 | HSPA9 | Hspa9 | 7347 | 0.042 | -0.2647 | No | ||
| 19 | ENO1 | Eno1 | 7490 | 0.019 | -0.2734 | No | ||
| 20 | PAPOLA | Papola | 7548 | 0.012 | -0.2768 | No | ||
| 21 | C1D | C1d | 8493 | -0.093 | -0.3357 | No | ||
| 22 | CNOT1 | Cnot1 | 8606 | -0.112 | -0.3404 | No | ||
| 23 | DDX6 | Ddx6 | 8689 | -0.122 | -0.3429 | No | ||
| 24 | TENT4A | Tent4a | 9050 | -0.172 | -0.3622 | No | ||
| 25 | CNOT2 | Cnot2 | 9262 | -0.205 | -0.3711 | No | ||
| 26 | EXOSC5 | Exosc5 | 9414 | -0.229 | -0.3756 | No | ||
| 27 | PARN | Parn | 9420 | -0.230 | -0.3706 | No | ||
| 28 | DCP2 | Dcp2 | 9552 | -0.249 | -0.3733 | No | ||
| 29 | EXOSC3 | Exosc3 | 10463 | -0.406 | -0.4227 | No | ||
| 30 | ZCCHC7 | Zcchc7 | 10970 | -0.465 | -0.4447 | No | ||
| 31 | CNOT3 | Cnot3 | 11016 | -0.474 | -0.4366 | No | ||
| 32 | LSM8 | Lsm8 | 11378 | -0.532 | -0.4477 | No | ||
| 33 | MTREX | Mtrex | 11467 | -0.550 | -0.4406 | No | ||
| 34 | LSM1 | Lsm1 | 11525 | -0.563 | -0.4313 | No | ||
| 35 | EXOSC9 | Exosc9 | 11542 | -0.568 | -0.4192 | No | ||
| 36 | WDR61 | Wdr61 | 11632 | -0.584 | -0.4115 | No | ||
| 37 | MPHOSPH6 | Mphosph6 | 12548 | -0.799 | -0.4521 | Yes | ||
| 38 | EXOSC1 | Exosc1 | 12838 | -0.867 | -0.4508 | Yes | ||
| 39 | CNOT6L | Cnot6l | 12890 | -0.886 | -0.4336 | Yes | ||
| 40 | LSM7 | Lsm7 | 13079 | -0.947 | -0.4238 | Yes | ||
| 41 | CNOT6 | Cnot6 | 13193 | -0.987 | -0.4083 | Yes | ||
| 42 | LSM4 | Lsm4 | 13295 | -1.035 | -0.3909 | Yes | ||
| 43 | DCPS | Dcps | 13368 | -1.063 | -0.3710 | Yes | ||
| 44 | EXOSC7 | Exosc7 | 13430 | -1.085 | -0.3499 | Yes | ||
| 45 | CNOT9 | Cnot9 | 13543 | -1.124 | -0.3311 | Yes | ||
| 46 | TTC37 | Ttc37 | 13666 | -1.174 | -0.3119 | Yes | ||
| 47 | LSM2 | Lsm2 | 13819 | -1.237 | -0.2931 | Yes | ||
| 48 | PNPT1 | Pnpt1 | 13876 | -1.270 | -0.2674 | Yes | ||
| 49 | LSM6 | Lsm6 | 14035 | -1.356 | -0.2462 | Yes | ||
| 50 | LSM5 | Lsm5 | 14501 | -1.708 | -0.2368 | Yes | ||
| 51 | EXOSC2 | Exosc2 | 14604 | -1.818 | -0.2013 | Yes | ||
| 52 | DIS3 | Dis3 | 14648 | -1.853 | -0.1613 | Yes | ||
| 53 | EXOSC8 | Exosc8 | 14725 | -1.934 | -0.1215 | Yes | ||
| 54 | LSM3 | Lsm3 | 14813 | -2.023 | -0.0803 | Yes | ||
| 55 | PAPOLB | Papolb | 15112 | -2.587 | -0.0398 | Yes | ||
| 56 | ENO3 | Eno3 | 15201 | -2.871 | 0.0209 | Yes |