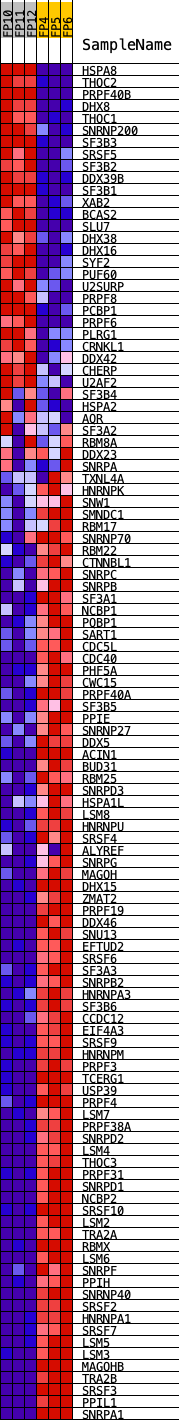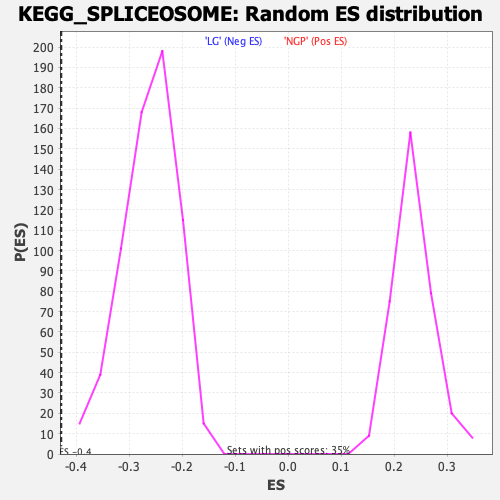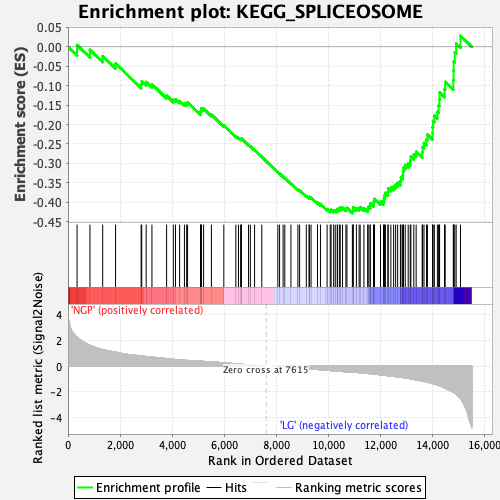
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_LG.phenotype.cls#NGP_versus_LG_repos |
| Phenotype | phenotype.cls#NGP_versus_LG_repos |
| Upregulated in class | LG |
| GeneSet | KEGG_SPLICEOSOME |
| Enrichment Score (ES) | -0.42828196 |
| Normalized Enrichment Score (NES) | -1.6373243 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.04644959 |
| FWER p-Value | 0.541 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | HSPA8 | Hspa8 | 348 | 2.249 | 0.0047 | No | ||
| 2 | THOC2 | Thoc2 | 844 | 1.618 | -0.0078 | No | ||
| 3 | PRPF40B | Prpf40b | 1334 | 1.294 | -0.0239 | No | ||
| 4 | DHX8 | Dhx8 | 1828 | 1.086 | -0.0427 | No | ||
| 5 | THOC1 | Thoc1 | 2811 | 0.791 | -0.0968 | No | ||
| 6 | SNRNP200 | Snrnp200 | 2835 | 0.787 | -0.0888 | No | ||
| 7 | SF3B3 | Sf3b3 | 3005 | 0.751 | -0.0906 | No | ||
| 8 | SRSF5 | Srsf5 | 3227 | 0.706 | -0.0964 | No | ||
| 9 | SF3B2 | Sf3b2 | 3791 | 0.580 | -0.1259 | No | ||
| 10 | DDX39B | Ddx39b | 4049 | 0.532 | -0.1361 | No | ||
| 11 | SF3B1 | Sf3b1 | 4132 | 0.515 | -0.1352 | No | ||
| 12 | XAB2 | Xab2 | 4291 | 0.489 | -0.1395 | No | ||
| 13 | BCAS2 | Bcas2 | 4476 | 0.462 | -0.1459 | No | ||
| 14 | SLU7 | Slu7 | 4562 | 0.449 | -0.1459 | No | ||
| 15 | DHX38 | Dhx38 | 4598 | 0.443 | -0.1428 | No | ||
| 16 | DHX16 | Dhx16 | 5096 | 0.388 | -0.1704 | No | ||
| 17 | SYF2 | Syf2 | 5097 | 0.388 | -0.1657 | No | ||
| 18 | PUF60 | Puf60 | 5122 | 0.384 | -0.1626 | No | ||
| 19 | U2SURP | U2surp | 5124 | 0.383 | -0.1580 | No | ||
| 20 | PRPF8 | Prpf8 | 5213 | 0.368 | -0.1592 | No | ||
| 21 | PCBP1 | Pcbp1 | 5513 | 0.340 | -0.1745 | No | ||
| 22 | PRPF6 | Prpf6 | 5991 | 0.260 | -0.2023 | No | ||
| 23 | PLRG1 | Plrg1 | 6453 | 0.184 | -0.2300 | No | ||
| 24 | CRNKL1 | Crnkl1 | 6554 | 0.167 | -0.2345 | No | ||
| 25 | DDX42 | Ddx42 | 6640 | 0.152 | -0.2382 | No | ||
| 26 | CHERP | Cherp | 6661 | 0.148 | -0.2377 | No | ||
| 27 | U2AF2 | U2af2 | 6663 | 0.148 | -0.2359 | No | ||
| 28 | SF3B4 | Sf3b4 | 6941 | 0.103 | -0.2526 | No | ||
| 29 | HSPA2 | Hspa2 | 7011 | 0.092 | -0.2560 | No | ||
| 30 | AQR | Aqr | 7173 | 0.067 | -0.2656 | No | ||
| 31 | SF3A2 | Sf3a2 | 7450 | 0.025 | -0.2832 | No | ||
| 32 | RBM8A | Rbm8a | 8068 | -0.030 | -0.3229 | No | ||
| 33 | DDX23 | Ddx23 | 8134 | -0.040 | -0.3266 | No | ||
| 34 | SNRPA | Snrpa | 8270 | -0.060 | -0.3347 | No | ||
| 35 | TXNL4A | Txnl4a | 8335 | -0.071 | -0.3380 | No | ||
| 36 | HNRNPK | Hnrnpk | 8571 | -0.104 | -0.3520 | No | ||
| 37 | SNW1 | Snw1 | 8839 | -0.146 | -0.3675 | No | ||
| 38 | SMNDC1 | Smndc1 | 8905 | -0.154 | -0.3699 | No | ||
| 39 | RBM17 | Rbm17 | 9162 | -0.188 | -0.3842 | No | ||
| 40 | SNRNP70 | Snrnp70 | 9272 | -0.205 | -0.3888 | No | ||
| 41 | RBM22 | Rbm22 | 9274 | -0.206 | -0.3863 | No | ||
| 42 | CTNNBL1 | Ctnnbl1 | 9344 | -0.218 | -0.3882 | No | ||
| 43 | SNRPC | Snrpc | 9592 | -0.254 | -0.4011 | No | ||
| 44 | SNRPB | Snrpb | 9706 | -0.274 | -0.4051 | No | ||
| 45 | SF3A1 | Sf3a1 | 9961 | -0.317 | -0.4178 | No | ||
| 46 | NCBP1 | Ncbp1 | 10081 | -0.336 | -0.4214 | No | ||
| 47 | PQBP1 | Pqbp1 | 10115 | -0.344 | -0.4194 | No | ||
| 48 | SART1 | Sart1 | 10213 | -0.365 | -0.4213 | No | ||
| 49 | CDC5L | Cdc5l | 10275 | -0.375 | -0.4207 | No | ||
| 50 | CDC40 | Cdc40 | 10351 | -0.387 | -0.4208 | No | ||
| 51 | PHF5A | Phf5a | 10367 | -0.389 | -0.4171 | No | ||
| 52 | CWC15 | Cwc15 | 10445 | -0.403 | -0.4172 | No | ||
| 53 | PRPF40A | Prpf40a | 10464 | -0.407 | -0.4135 | No | ||
| 54 | SF3B5 | Sf3b5 | 10555 | -0.410 | -0.4143 | No | ||
| 55 | PPIE | Ppie | 10685 | -0.424 | -0.4175 | No | ||
| 56 | SNRNP27 | Snrnp27 | 10724 | -0.431 | -0.4148 | No | ||
| 57 | DDX5 | Ddx5 | 10933 | -0.459 | -0.4227 | Yes | ||
| 58 | ACIN1 | Acin1 | 10962 | -0.463 | -0.4189 | Yes | ||
| 59 | BUD31 | Bud31 | 10963 | -0.463 | -0.4133 | Yes | ||
| 60 | RBM25 | Rbm25 | 11088 | -0.481 | -0.4155 | Yes | ||
| 61 | SNRPD3 | Snrpd3 | 11192 | -0.497 | -0.4162 | Yes | ||
| 62 | HSPA1L | Hspa1l | 11233 | -0.503 | -0.4127 | Yes | ||
| 63 | LSM8 | Lsm8 | 11378 | -0.532 | -0.4156 | Yes | ||
| 64 | HNRNPU | Hnrnpu | 11522 | -0.562 | -0.4180 | Yes | ||
| 65 | SRSF4 | Srsf4 | 11540 | -0.567 | -0.4123 | Yes | ||
| 66 | ALYREF | 4931428L18Rik | 11615 | -0.581 | -0.4100 | Yes | ||
| 67 | SNRPG | Snrpg | 11620 | -0.582 | -0.4032 | Yes | ||
| 68 | MAGOH | Magoh | 11745 | -0.605 | -0.4039 | Yes | ||
| 69 | DHX15 | Dhx15 | 11758 | -0.608 | -0.3973 | Yes | ||
| 70 | ZMAT2 | Zmat2 | 11785 | -0.615 | -0.3916 | Yes | ||
| 71 | PRPF19 | Prpf19 | 12014 | -0.670 | -0.3983 | Yes | ||
| 72 | DDX46 | Ddx46 | 12131 | -0.699 | -0.3973 | Yes | ||
| 73 | SNU13 | Snu13 | 12149 | -0.703 | -0.3899 | Yes | ||
| 74 | EFTUD2 | Eftud2 | 12171 | -0.708 | -0.3827 | Yes | ||
| 75 | SRSF6 | Srsf6 | 12203 | -0.713 | -0.3760 | Yes | ||
| 76 | SF3A3 | Sf3a3 | 12302 | -0.740 | -0.3734 | Yes | ||
| 77 | SNRPB2 | Snrpb2 | 12309 | -0.741 | -0.3648 | Yes | ||
| 78 | HNRNPA3 | Gm17190 | 12413 | -0.765 | -0.3622 | Yes | ||
| 79 | SF3B6 | Sf3b6 | 12520 | -0.791 | -0.3595 | Yes | ||
| 80 | CCDC12 | Ccdc12 | 12596 | -0.811 | -0.3546 | Yes | ||
| 81 | EIF4A3 | Eif4a3 | 12674 | -0.828 | -0.3495 | Yes | ||
| 82 | SRSF9 | Srsf9 | 12780 | -0.852 | -0.3460 | Yes | ||
| 83 | HNRNPM | Hnrnpm | 12800 | -0.858 | -0.3368 | Yes | ||
| 84 | PRPF3 | Prpf3 | 12869 | -0.879 | -0.3306 | Yes | ||
| 85 | TCERG1 | Tcerg1 | 12880 | -0.883 | -0.3205 | Yes | ||
| 86 | USP39 | Usp39 | 12898 | -0.889 | -0.3109 | Yes | ||
| 87 | PRPF4 | Prpf4 | 12966 | -0.910 | -0.3042 | Yes | ||
| 88 | LSM7 | Lsm7 | 13079 | -0.947 | -0.3000 | Yes | ||
| 89 | PRPF38A | Prpf38a | 13157 | -0.973 | -0.2932 | Yes | ||
| 90 | SNRPD2 | Snrpd2 | 13182 | -0.982 | -0.2828 | Yes | ||
| 91 | LSM4 | Lsm4 | 13295 | -1.035 | -0.2775 | Yes | ||
| 92 | THOC3 | Thoc3 | 13385 | -1.070 | -0.2703 | Yes | ||
| 93 | PRPF31 | Prpf31 | 13617 | -1.154 | -0.2713 | Yes | ||
| 94 | SNRPD1 | Snrpd1 | 13634 | -1.159 | -0.2583 | Yes | ||
| 95 | NCBP2 | Ncbp2 | 13687 | -1.181 | -0.2474 | Yes | ||
| 96 | SRSF10 | Srsf10 | 13784 | -1.220 | -0.2388 | Yes | ||
| 97 | LSM2 | Lsm2 | 13819 | -1.237 | -0.2260 | Yes | ||
| 98 | TRA2A | Tra2a | 14015 | -1.342 | -0.2224 | Yes | ||
| 99 | RBMX | Rbmx | 14019 | -1.346 | -0.2063 | Yes | ||
| 100 | LSM6 | Lsm6 | 14035 | -1.356 | -0.1908 | Yes | ||
| 101 | SNRPF | Snrpf | 14084 | -1.387 | -0.1771 | Yes | ||
| 102 | PPIH | Gm7879 | 14201 | -1.461 | -0.1669 | Yes | ||
| 103 | SNRNP40 | Snrnp40 | 14249 | -1.492 | -0.1519 | Yes | ||
| 104 | SRSF2 | Srsf2 | 14277 | -1.511 | -0.1353 | Yes | ||
| 105 | HNRNPA1 | Hnrnpa1 | 14286 | -1.513 | -0.1175 | Yes | ||
| 106 | SRSF7 | Srsf7 | 14476 | -1.685 | -0.1094 | Yes | ||
| 107 | LSM5 | Lsm5 | 14501 | -1.708 | -0.0902 | Yes | ||
| 108 | LSM3 | Lsm3 | 14813 | -2.023 | -0.0859 | Yes | ||
| 109 | MAGOHB | Magohb | 14823 | -2.031 | -0.0618 | Yes | ||
| 110 | TRA2B | Tra2b | 14828 | -2.039 | -0.0374 | Yes | ||
| 111 | SRSF3 | Srsf3 | 14871 | -2.098 | -0.0147 | Yes | ||
| 112 | PPIL1 | Ppil1 | 14923 | -2.180 | 0.0084 | Yes | ||
| 113 | SNRPA1 | Snrpa1 | 15089 | -2.520 | 0.0283 | Yes |