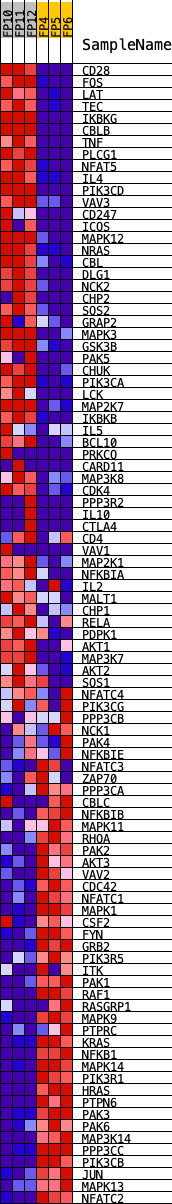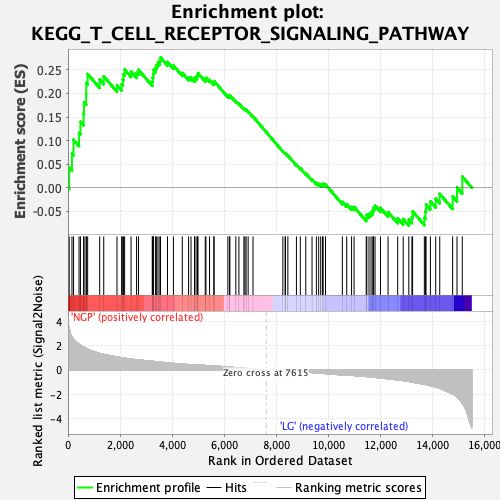
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_LG.phenotype.cls#NGP_versus_LG_repos |
| Phenotype | phenotype.cls#NGP_versus_LG_repos |
| Upregulated in class | NGP |
| GeneSet | KEGG_T_CELL_RECEPTOR_SIGNALING_PATHWAY |
| Enrichment Score (ES) | 0.2765091 |
| Normalized Enrichment Score (NES) | 1.1353918 |
| Nominal p-value | 0.1993865 |
| FDR q-value | 0.6675182 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CD28 | Cd28 | 45 | 3.429 | 0.0427 | Yes | ||
| 2 | FOS | Fos | 152 | 2.726 | 0.0720 | Yes | ||
| 3 | LAT | Lat | 213 | 2.533 | 0.1018 | Yes | ||
| 4 | TEC | Tec | 424 | 2.118 | 0.1164 | Yes | ||
| 5 | IKBKG | Ikbkg | 478 | 2.041 | 0.1401 | Yes | ||
| 6 | CBLB | Cblb | 600 | 1.881 | 0.1572 | Yes | ||
| 7 | TNF | Tnf | 615 | 1.864 | 0.1811 | Yes | ||
| 8 | PLCG1 | Plcg1 | 698 | 1.760 | 0.1992 | Yes | ||
| 9 | NFAT5 | Nfat5 | 701 | 1.754 | 0.2224 | Yes | ||
| 10 | IL4 | Il4 | 756 | 1.694 | 0.2414 | Yes | ||
| 11 | PIK3CD | Pik3cd | 1218 | 1.353 | 0.2295 | Yes | ||
| 12 | VAV3 | Vav3 | 1374 | 1.276 | 0.2364 | Yes | ||
| 13 | CD247 | Cd247 | 1883 | 1.064 | 0.2177 | Yes | ||
| 14 | ICOS | Icos | 2061 | 0.991 | 0.2194 | Yes | ||
| 15 | MAPK12 | Mapk12 | 2104 | 0.983 | 0.2297 | Yes | ||
| 16 | NRAS | Nras | 2132 | 0.975 | 0.2409 | Yes | ||
| 17 | CBL | Cbl | 2177 | 0.958 | 0.2508 | Yes | ||
| 18 | DLG1 | Dlg1 | 2424 | 0.886 | 0.2466 | Yes | ||
| 19 | NCK2 | Nck2 | 2636 | 0.836 | 0.2441 | Yes | ||
| 20 | CHP2 | Chp2 | 2711 | 0.819 | 0.2502 | Yes | ||
| 21 | SOS2 | Sos2 | 3239 | 0.702 | 0.2254 | Yes | ||
| 22 | GRAP2 | Grap2 | 3254 | 0.699 | 0.2337 | Yes | ||
| 23 | MAPK3 | Mapk3 | 3274 | 0.697 | 0.2418 | Yes | ||
| 24 | GSK3B | Gsk3b | 3289 | 0.693 | 0.2501 | Yes | ||
| 25 | PAK5 | Pak7 | 3367 | 0.678 | 0.2541 | Yes | ||
| 26 | CHUK | Chuk | 3408 | 0.666 | 0.2604 | Yes | ||
| 27 | PIK3CA | Pik3ca | 3461 | 0.655 | 0.2657 | Yes | ||
| 28 | LCK | Lck | 3532 | 0.640 | 0.2697 | Yes | ||
| 29 | MAP2K7 | Map2k7 | 3558 | 0.635 | 0.2765 | Yes | ||
| 30 | IKBKB | Ikbkb | 3825 | 0.572 | 0.2669 | No | ||
| 31 | IL5 | Il5 | 4053 | 0.531 | 0.2592 | No | ||
| 32 | BCL10 | Bcl10 | 4394 | 0.475 | 0.2435 | No | ||
| 33 | PRKCQ | Prkcq | 4635 | 0.440 | 0.2338 | No | ||
| 34 | CARD11 | Card11 | 4729 | 0.428 | 0.2335 | No | ||
| 35 | MAP3K8 | Map3k8 | 4865 | 0.412 | 0.2302 | No | ||
| 36 | CDK4 | Cdk4 | 4881 | 0.408 | 0.2346 | No | ||
| 37 | PPP3R2 | Ppp3r2 | 4958 | 0.401 | 0.2351 | No | ||
| 38 | IL10 | Il10 | 4976 | 0.401 | 0.2393 | No | ||
| 39 | CTLA4 | Ctla4 | 4998 | 0.401 | 0.2432 | No | ||
| 40 | CD4 | Cd4 | 5276 | 0.357 | 0.2300 | No | ||
| 41 | VAV1 | Vav1 | 5304 | 0.356 | 0.2330 | No | ||
| 42 | MAP2K1 | Map2k1 | 5442 | 0.350 | 0.2288 | No | ||
| 43 | NFKBIA | Nfkbia | 5607 | 0.322 | 0.2225 | No | ||
| 44 | IL2 | Il2 | 5616 | 0.320 | 0.2262 | No | ||
| 45 | MALT1 | Malt1 | 6146 | 0.236 | 0.1951 | No | ||
| 46 | CHP1 | Chp1 | 6216 | 0.224 | 0.1936 | No | ||
| 47 | RELA | Rela | 6219 | 0.223 | 0.1964 | No | ||
| 48 | PDPK1 | Pdpk1 | 6452 | 0.184 | 0.1838 | No | ||
| 49 | AKT1 | Akt1 | 6571 | 0.164 | 0.1783 | No | ||
| 50 | MAP3K7 | Map3k7 | 6762 | 0.130 | 0.1678 | No | ||
| 51 | AKT2 | Akt2 | 6803 | 0.125 | 0.1668 | No | ||
| 52 | SOS1 | Sos1 | 6848 | 0.116 | 0.1655 | No | ||
| 53 | NFATC4 | Nfatc4 | 6926 | 0.106 | 0.1619 | No | ||
| 54 | PIK3CG | Pik3cg | 7113 | 0.076 | 0.1509 | No | ||
| 55 | PPP3CB | Ppp3cb | 8260 | -0.059 | 0.0774 | No | ||
| 56 | NCK1 | Nck1 | 8346 | -0.073 | 0.0729 | No | ||
| 57 | PAK4 | Pak4 | 8362 | -0.075 | 0.0729 | No | ||
| 58 | NFKBIE | Nfkbie | 8453 | -0.087 | 0.0682 | No | ||
| 59 | NFATC3 | Nfatc3 | 8782 | -0.138 | 0.0488 | No | ||
| 60 | ZAP70 | Zap70 | 8932 | -0.159 | 0.0413 | No | ||
| 61 | PPP3CA | Ppp3ca | 9141 | -0.186 | 0.0302 | No | ||
| 62 | CBLC | Cblc | 9382 | -0.224 | 0.0177 | No | ||
| 63 | NFKBIB | Nfkbib | 9549 | -0.248 | 0.0102 | No | ||
| 64 | MAPK11 | Mapk11 | 9637 | -0.261 | 0.0081 | No | ||
| 65 | RHOA | Rhoa | 9715 | -0.276 | 0.0067 | No | ||
| 66 | PAK2 | Pak2 | 9784 | -0.286 | 0.0061 | No | ||
| 67 | AKT3 | Akt3 | 9809 | -0.291 | 0.0084 | No | ||
| 68 | VAV2 | Vav2 | 9900 | -0.308 | 0.0067 | No | ||
| 69 | CDC42 | Cdc42 | 10548 | -0.409 | -0.0298 | No | ||
| 70 | NFATC1 | Nfatc1 | 10715 | -0.429 | -0.0348 | No | ||
| 71 | MAPK1 | Mapk1 | 10902 | -0.453 | -0.0409 | No | ||
| 72 | CSF2 | Csf2 | 10999 | -0.469 | -0.0409 | No | ||
| 73 | FYN | Fyn | 11465 | -0.549 | -0.0637 | No | ||
| 74 | GRB2 | Grb2 | 11488 | -0.555 | -0.0577 | No | ||
| 75 | PIK3R5 | Pik3r5 | 11573 | -0.572 | -0.0556 | No | ||
| 76 | ITK | Itk | 11651 | -0.589 | -0.0527 | No | ||
| 77 | PAK1 | Pak1 | 11717 | -0.600 | -0.0490 | No | ||
| 78 | RAF1 | Raf1 | 11747 | -0.605 | -0.0428 | No | ||
| 79 | RASGRP1 | Rasgrp1 | 11804 | -0.619 | -0.0382 | No | ||
| 80 | MAPK9 | Mapk9 | 12013 | -0.670 | -0.0428 | No | ||
| 81 | PTPRC | Ptprc | 12305 | -0.740 | -0.0518 | No | ||
| 82 | KRAS | Kras | 12675 | -0.828 | -0.0647 | No | ||
| 83 | NFKB1 | Nfkb1 | 12888 | -0.885 | -0.0667 | No | ||
| 84 | MAPK14 | Mapk14 | 13103 | -0.956 | -0.0678 | No | ||
| 85 | PIK3R1 | Pik3r1 | 13222 | -1.001 | -0.0622 | No | ||
| 86 | HRAS | Hras | 13248 | -1.014 | -0.0503 | No | ||
| 87 | PTPN6 | Ptpn6 | 13700 | -1.188 | -0.0637 | No | ||
| 88 | PAK3 | Pak3 | 13741 | -1.206 | -0.0503 | No | ||
| 89 | PAK6 | Pak6 | 13766 | -1.215 | -0.0357 | No | ||
| 90 | MAP3K14 | Map3k14 | 13933 | -1.303 | -0.0291 | No | ||
| 91 | PPP3CC | Ppp3cc | 14137 | -1.416 | -0.0234 | No | ||
| 92 | PIK3CB | Pik3cb | 14292 | -1.520 | -0.0132 | No | ||
| 93 | JUN | Jun | 14788 | -2.002 | -0.0187 | No | ||
| 94 | MAPK13 | Mapk13 | 14956 | -2.243 | 0.0003 | No | ||
| 95 | NFATC2 | Nfatc2 | 15159 | -2.746 | 0.0237 | No |

