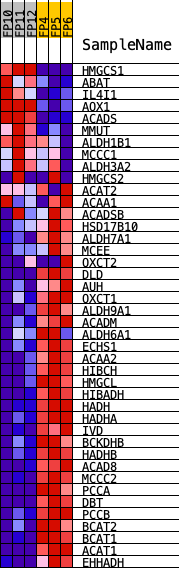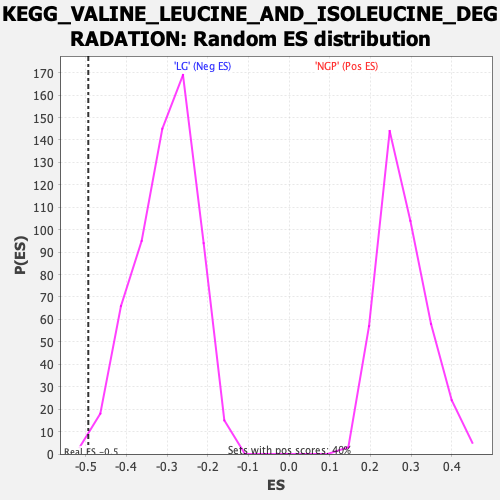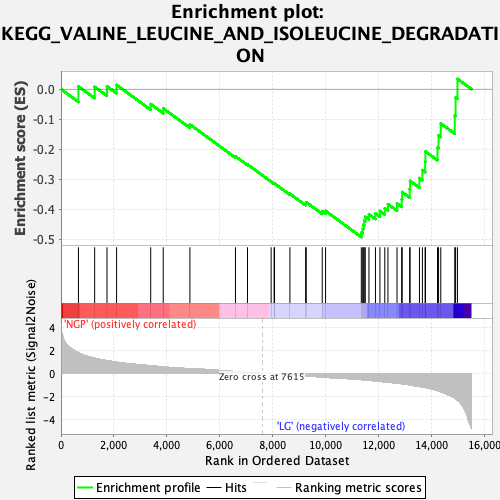
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_LG.phenotype.cls#NGP_versus_LG_repos |
| Phenotype | phenotype.cls#NGP_versus_LG_repos |
| Upregulated in class | LG |
| GeneSet | KEGG_VALINE_LEUCINE_AND_ISOLEUCINE_DEGRADATION |
| Enrichment Score (ES) | -0.49353573 |
| Normalized Enrichment Score (NES) | -1.6387469 |
| Nominal p-value | 0.0033057851 |
| FDR q-value | 0.048460312 |
| FWER p-Value | 0.535 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | HMGCS1 | Hmgcs1 | 660 | 1.817 | 0.0093 | No | ||
| 2 | ABAT | Abat | 1273 | 1.327 | 0.0077 | No | ||
| 3 | IL4I1 | Il4i1 | 1739 | 1.120 | 0.0097 | No | ||
| 4 | AOX1 | Aox1 | 2102 | 0.983 | 0.0144 | No | ||
| 5 | ACADS | Acads | 3393 | 0.671 | -0.0498 | No | ||
| 6 | MMUT | Mmut | 3866 | 0.564 | -0.0641 | No | ||
| 7 | ALDH1B1 | Aldh1b1 | 4875 | 0.410 | -0.1175 | No | ||
| 8 | MCCC1 | Mccc1 | 6598 | 0.161 | -0.2241 | No | ||
| 9 | ALDH3A2 | Aldh3a2 | 7053 | 0.084 | -0.2510 | No | ||
| 10 | HMGCS2 | Hmgcs2 | 7948 | -0.009 | -0.3085 | No | ||
| 11 | ACAT2 | Acat2 | 8060 | -0.030 | -0.3148 | No | ||
| 12 | ACAA1 | Acaa1a | 8076 | -0.031 | -0.3149 | No | ||
| 13 | ACADSB | Acadsb | 8658 | -0.117 | -0.3491 | No | ||
| 14 | HSD17B10 | Hsd17b10 | 9253 | -0.202 | -0.3817 | No | ||
| 15 | ALDH7A1 | Aldh7a1 | 9275 | -0.206 | -0.3771 | No | ||
| 16 | MCEE | Mcee | 9880 | -0.304 | -0.4075 | No | ||
| 17 | OXCT2 | Oxct2a | 10006 | -0.323 | -0.4063 | No | ||
| 18 | DLD | Dld | 11358 | -0.528 | -0.4785 | Yes | ||
| 19 | AUH | Auh | 11412 | -0.539 | -0.4665 | Yes | ||
| 20 | OXCT1 | Oxct1 | 11433 | -0.542 | -0.4523 | Yes | ||
| 21 | ALDH9A1 | Aldh9a1 | 11469 | -0.550 | -0.4388 | Yes | ||
| 22 | ACADM | Acadm | 11509 | -0.559 | -0.4254 | Yes | ||
| 23 | ALDH6A1 | Aldh6a1 | 11647 | -0.589 | -0.4174 | Yes | ||
| 24 | ECHS1 | Echs1 | 11893 | -0.641 | -0.4149 | Yes | ||
| 25 | ACAA2 | Acaa2 | 12062 | -0.681 | -0.4063 | Yes | ||
| 26 | HIBCH | Hibch | 12246 | -0.725 | -0.3974 | Yes | ||
| 27 | HMGCL | Hmgcl | 12366 | -0.755 | -0.3835 | Yes | ||
| 28 | HIBADH | Hibadh | 12709 | -0.837 | -0.3816 | Yes | ||
| 29 | HADH | Hadh | 12884 | -0.884 | -0.3676 | Yes | ||
| 30 | HADHA | Hadha | 12903 | -0.892 | -0.3433 | Yes | ||
| 31 | IVD | Ivd | 13188 | -0.985 | -0.3335 | Yes | ||
| 32 | BCKDHB | Bckdhb | 13202 | -0.992 | -0.3060 | Yes | ||
| 33 | HADHB | Hadhb | 13557 | -1.128 | -0.2966 | Yes | ||
| 34 | ACAD8 | Acad8 | 13668 | -1.174 | -0.2702 | Yes | ||
| 35 | MCCC2 | Mccc2 | 13773 | -1.217 | -0.2421 | Yes | ||
| 36 | PCCA | Pcca | 13783 | -1.220 | -0.2078 | Yes | ||
| 37 | DBT | Dbt | 14240 | -1.483 | -0.1949 | Yes | ||
| 38 | PCCB | Pccb | 14280 | -1.512 | -0.1542 | Yes | ||
| 39 | BCAT2 | Bcat2 | 14364 | -1.577 | -0.1145 | Yes | ||
| 40 | BCAT1 | Bcat1 | 14893 | -2.125 | -0.0878 | Yes | ||
| 41 | ACAT1 | Acat1 | 14924 | -2.181 | -0.0275 | Yes | ||
| 42 | EHHADH | Ehhadh | 14995 | -2.317 | 0.0342 | Yes |