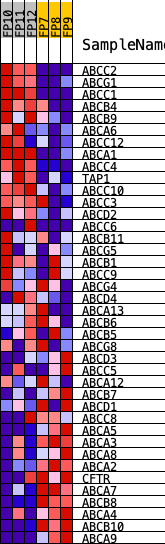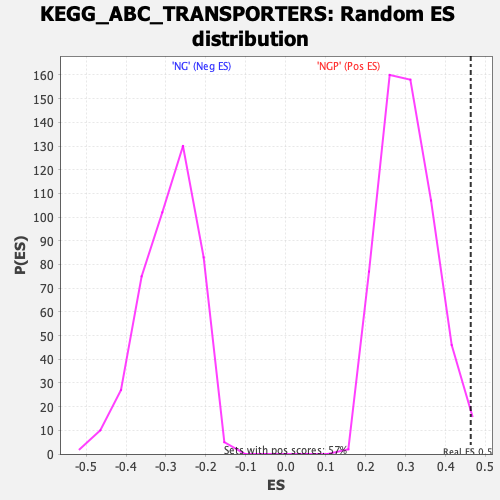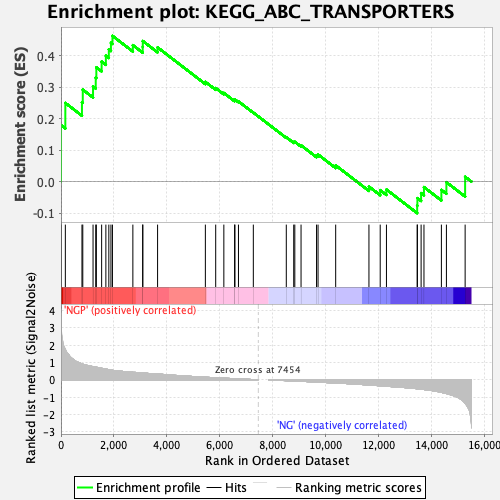
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_NG.phenotype.cls#NGP_versus_NG_repos |
| Phenotype | phenotype.cls#NGP_versus_NG_repos |
| Upregulated in class | NGP |
| GeneSet | KEGG_ABC_TRANSPORTERS |
| Enrichment Score (ES) | 0.46317863 |
| Normalized Enrichment Score (NES) | 1.5205207 |
| Nominal p-value | 0.01590106 |
| FDR q-value | 0.29892746 |
| FWER p-Value | 0.873 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ABCC2 | Abcc2 | 1 | 3.848 | 0.1792 | Yes | ||
| 2 | ABCG1 | Abcg1 | 164 | 1.744 | 0.2500 | Yes | ||
| 3 | ABCC1 | Abcc1 | 793 | 0.918 | 0.2522 | Yes | ||
| 4 | ABCB4 | Abcb4 | 824 | 0.900 | 0.2922 | Yes | ||
| 5 | ABCB9 | Abcb9 | 1212 | 0.756 | 0.3025 | Yes | ||
| 6 | ABCA6 | Abca6 | 1314 | 0.740 | 0.3305 | Yes | ||
| 7 | ABCC12 | Abcc12 | 1337 | 0.733 | 0.3632 | Yes | ||
| 8 | ABCA1 | Abca1 | 1537 | 0.666 | 0.3814 | Yes | ||
| 9 | ABCC4 | Abcc4 | 1697 | 0.621 | 0.4000 | Yes | ||
| 10 | TAP1 | Tap1 | 1809 | 0.585 | 0.4201 | Yes | ||
| 11 | ABCC10 | Abcc10 | 1889 | 0.562 | 0.4412 | Yes | ||
| 12 | ABCC3 | Abcc3 | 1946 | 0.549 | 0.4632 | Yes | ||
| 13 | ABCD2 | Abcd2 | 2718 | 0.433 | 0.4336 | No | ||
| 14 | ABCC6 | Abcc6 | 3090 | 0.401 | 0.4283 | No | ||
| 15 | ABCB11 | Abcb11 | 3094 | 0.401 | 0.4468 | No | ||
| 16 | ABCG5 | Abcg5 | 3653 | 0.339 | 0.4266 | No | ||
| 17 | ABCB1 | Abcb1a | 5460 | 0.151 | 0.3169 | No | ||
| 18 | ABCC9 | Abcc9 | 5851 | 0.118 | 0.2973 | No | ||
| 19 | ABCG4 | Abcg4 | 6159 | 0.093 | 0.2818 | No | ||
| 20 | ABCD4 | Abcd4 | 6566 | 0.066 | 0.2586 | No | ||
| 21 | ABCA13 | Abca13 | 6578 | 0.065 | 0.2610 | No | ||
| 22 | ABCB6 | Abcb6 | 6712 | 0.055 | 0.2550 | No | ||
| 23 | ABCB5 | Abcb5 | 7274 | 0.014 | 0.2194 | No | ||
| 24 | ABCG8 | Abcg8 | 8521 | -0.046 | 0.1410 | No | ||
| 25 | ABCD3 | Abcd3 | 8802 | -0.065 | 0.1260 | No | ||
| 26 | ABCC5 | Abcc5 | 8843 | -0.068 | 0.1266 | No | ||
| 27 | ABCA12 | Abca12 | 9079 | -0.085 | 0.1153 | No | ||
| 28 | ABCB7 | Abcb7 | 9665 | -0.129 | 0.0836 | No | ||
| 29 | ABCD1 | Abcd1 | 9722 | -0.133 | 0.0861 | No | ||
| 30 | ABCC8 | Abcc8 | 10389 | -0.183 | 0.0517 | No | ||
| 31 | ABCA5 | Abca5 | 11647 | -0.296 | -0.0157 | No | ||
| 32 | ABCA3 | Abca3 | 12073 | -0.343 | -0.0271 | No | ||
| 33 | ABCA8 | Abca8b | 12309 | -0.368 | -0.0252 | No | ||
| 34 | ABCA2 | Abca2 | 13471 | -0.516 | -0.0761 | No | ||
| 35 | CFTR | Cftr | 13480 | -0.517 | -0.0525 | No | ||
| 36 | ABCA7 | Abca7 | 13620 | -0.536 | -0.0365 | No | ||
| 37 | ABCB8 | Abcb8 | 13728 | -0.558 | -0.0175 | No | ||
| 38 | ABCA4 | Abca4 | 14387 | -0.722 | -0.0263 | No | ||
| 39 | ABCB10 | Abcb10 | 14577 | -0.788 | -0.0018 | No | ||
| 40 | ABCA9 | Abca9 | 15285 | -1.351 | 0.0155 | No |