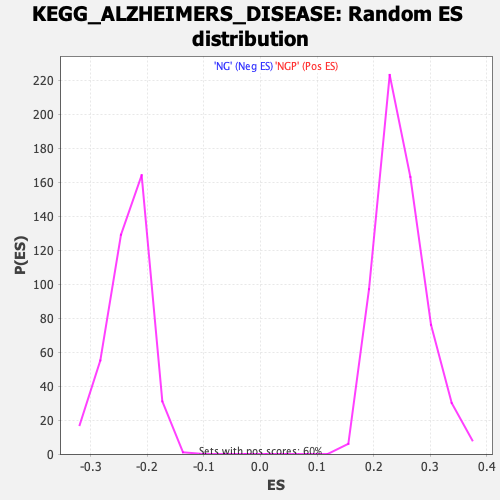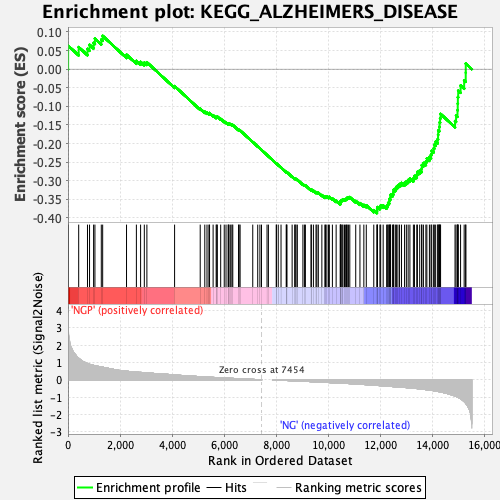
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_NG.phenotype.cls#NGP_versus_NG_repos |
| Phenotype | phenotype.cls#NGP_versus_NG_repos |
| Upregulated in class | NG |
| GeneSet | KEGG_ALZHEIMERS_DISEASE |
| Enrichment Score (ES) | -0.38708347 |
| Normalized Enrichment Score (NES) | -1.6622669 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.07719004 |
| FWER p-Value | 0.352 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CACNA1D | Cacna1d | 5 | 3.519 | 0.0632 | No | ||
| 2 | ATP2A1 | Atp2a1 | 411 | 1.242 | 0.0593 | No | ||
| 3 | CASP7 | Casp7 | 749 | 0.942 | 0.0544 | No | ||
| 4 | GRIN2B | Grin2b | 828 | 0.898 | 0.0656 | No | ||
| 5 | COX6B2 | Cox6b2 | 984 | 0.837 | 0.0706 | No | ||
| 6 | CHP2 | Chp2 | 1035 | 0.819 | 0.0821 | No | ||
| 7 | COX8C | Cox8b | 1280 | 0.741 | 0.0796 | No | ||
| 8 | COX4I2 | Cox4i2 | 1333 | 0.734 | 0.0895 | No | ||
| 9 | CDK5R1 | Cdk5r1 | 2250 | 0.495 | 0.0389 | No | ||
| 10 | ITPR1 | Itpr1 | 2629 | 0.441 | 0.0223 | No | ||
| 11 | IL1B | Il1b | 2791 | 0.428 | 0.0195 | No | ||
| 12 | LPL | Lpl | 2934 | 0.410 | 0.0177 | No | ||
| 13 | PPP3R2 | Ppp3r2 | 3033 | 0.401 | 0.0186 | No | ||
| 14 | GRIN2D | Grin2d | 4098 | 0.282 | -0.0455 | No | ||
| 15 | BACE2 | Bace2 | 5083 | 0.182 | -0.1062 | No | ||
| 16 | ITPR2 | Itpr2 | 5259 | 0.167 | -0.1146 | No | ||
| 17 | NDUFA6 | Ndufa6 | 5344 | 0.161 | -0.1171 | No | ||
| 18 | UQCRFS1 | Uqcrfs1 | 5418 | 0.155 | -0.1191 | No | ||
| 19 | HSD17B10 | Hsd17b10 | 5442 | 0.153 | -0.1178 | No | ||
| 20 | MAPK3 | Mapk3 | 5579 | 0.140 | -0.1241 | No | ||
| 21 | NDUFB2 | Ndufb2 | 5696 | 0.131 | -0.1293 | No | ||
| 22 | NDUFA10 | Ndufa10 | 5721 | 0.129 | -0.1285 | No | ||
| 23 | CYC1 | Cyc1 | 5740 | 0.126 | -0.1274 | No | ||
| 24 | NDUFV1 | Ndufv1 | 5870 | 0.117 | -0.1337 | No | ||
| 25 | CASP3 | Casp3 | 6007 | 0.107 | -0.1406 | No | ||
| 26 | APAF1 | Apaf1 | 6075 | 0.101 | -0.1431 | No | ||
| 27 | NDUFS3 | Ndufs3 | 6158 | 0.093 | -0.1468 | No | ||
| 28 | COX7A2L | Cox7a2l | 6167 | 0.093 | -0.1456 | No | ||
| 29 | APH1A | Aph1a | 6193 | 0.091 | -0.1456 | No | ||
| 30 | PLCB3 | Plcb3 | 6255 | 0.087 | -0.1480 | No | ||
| 31 | GRIN2A | Grin2a | 6303 | 0.084 | -0.1495 | No | ||
| 32 | NDUFB5 | Ndufb5 | 6335 | 0.082 | -0.1501 | No | ||
| 33 | GAPDH | Gapdh | 6557 | 0.067 | -0.1632 | No | ||
| 34 | PPP3CC | Ppp3cc | 6577 | 0.065 | -0.1633 | No | ||
| 35 | PSEN1 | Psen1 | 6617 | 0.063 | -0.1647 | No | ||
| 36 | CYCS | Cycs | 7103 | 0.027 | -0.1957 | No | ||
| 37 | TNF | Tnf | 7294 | 0.011 | -0.2079 | No | ||
| 38 | NDUFS2 | Ndufs2 | 7378 | 0.006 | -0.2132 | No | ||
| 39 | UQCRC2 | Uqcrc2 | 7439 | 0.002 | -0.2171 | No | ||
| 40 | COX7B2 | Cox7b2 | 7652 | 0.000 | -0.2308 | No | ||
| 41 | MT-CO2 | mt-Co2 | 7705 | 0.000 | -0.2342 | No | ||
| 42 | NDUFA4L2 | Ndufa4l2 | 8009 | -0.011 | -0.2537 | No | ||
| 43 | NDUFS7 | Ndufs7 | 8011 | -0.011 | -0.2536 | No | ||
| 44 | ATP5F1C | Atp5c1 | 8083 | -0.016 | -0.2579 | No | ||
| 45 | TNFRSF1A | Tnfrsf1a | 8190 | -0.024 | -0.2644 | No | ||
| 46 | COX6A2 | Cox6a2 | 8390 | -0.037 | -0.2767 | No | ||
| 47 | ATP5F1B | Atp5b | 8399 | -0.037 | -0.2765 | No | ||
| 48 | ATP2A2 | Atp2a2 | 8419 | -0.038 | -0.2771 | No | ||
| 49 | GRIN2C | Grin2c | 8617 | -0.052 | -0.2889 | No | ||
| 50 | ERN1 | Ern1 | 8717 | -0.060 | -0.2943 | No | ||
| 51 | NAE1 | Nae1 | 8734 | -0.061 | -0.2942 | No | ||
| 52 | PSEN2 | Psen2 | 8767 | -0.062 | -0.2952 | No | ||
| 53 | NDUFB9 | Ndufb9 | 8817 | -0.066 | -0.2972 | No | ||
| 54 | NDUFAB1 | Ndufab1 | 9031 | -0.081 | -0.3096 | No | ||
| 55 | FADD | Fadd | 9090 | -0.085 | -0.3118 | No | ||
| 56 | NDUFA9 | Ndufa9 | 9127 | -0.088 | -0.3125 | No | ||
| 57 | NDUFS4 | Ndufs4 | 9344 | -0.103 | -0.3247 | No | ||
| 58 | COX5B | Cox5b | 9355 | -0.103 | -0.3235 | No | ||
| 59 | SDHC | Sdhc | 9435 | -0.110 | -0.3267 | No | ||
| 60 | COX4I1 | Cox4i1 | 9535 | -0.118 | -0.3310 | No | ||
| 61 | SNCA | Snca | 9568 | -0.120 | -0.3309 | No | ||
| 62 | PLCB4 | Plcb4 | 9624 | -0.125 | -0.3322 | No | ||
| 63 | ATP5F1A | Atp5a1 | 9760 | -0.135 | -0.3385 | No | ||
| 64 | COX7C | Cox7c | 9876 | -0.145 | -0.3434 | No | ||
| 65 | CAPN2 | Capn2 | 9901 | -0.147 | -0.3423 | No | ||
| 66 | NDUFS1 | Ndufs1 | 9952 | -0.151 | -0.3428 | No | ||
| 67 | COX6B1 | Cox6b1 | 10014 | -0.155 | -0.3440 | No | ||
| 68 | NDUFV2 | Ndufv2 | 10048 | -0.157 | -0.3433 | No | ||
| 69 | NOS1 | Nos1 | 10165 | -0.166 | -0.3479 | No | ||
| 70 | BACE1 | Bace1 | 10309 | -0.178 | -0.3539 | No | ||
| 71 | APBB1 | Apbb1 | 10467 | -0.189 | -0.3607 | No | ||
| 72 | SDHB | Sdhb | 10474 | -0.190 | -0.3577 | No | ||
| 73 | CASP9 | Casp9 | 10476 | -0.190 | -0.3543 | No | ||
| 74 | GSK3B | Gsk3b | 10504 | -0.192 | -0.3526 | No | ||
| 75 | NCSTN | Ncstn | 10540 | -0.195 | -0.3514 | No | ||
| 76 | NDUFB10 | Ndufb10 | 10571 | -0.197 | -0.3498 | No | ||
| 77 | NDUFB3 | Ndufb3 | 10631 | -0.200 | -0.3500 | No | ||
| 78 | NDUFA8 | Ndufa8 | 10676 | -0.204 | -0.3492 | No | ||
| 79 | MAPK1 | Mapk1 | 10708 | -0.207 | -0.3474 | No | ||
| 80 | CALM2 | Calm2 | 10735 | -0.209 | -0.3453 | No | ||
| 81 | COX5A | Cox5a | 10781 | -0.213 | -0.3444 | No | ||
| 82 | PPP3CB | Ppp3cb | 10834 | -0.218 | -0.3439 | No | ||
| 83 | APOE | Apoe | 11062 | -0.239 | -0.3543 | No | ||
| 84 | COX7A2 | Cox7a2 | 11224 | -0.255 | -0.3602 | No | ||
| 85 | MT-CYB | mt-Cytb | 11374 | -0.270 | -0.3650 | No | ||
| 86 | UQCRC1 | Uqcrc1 | 11468 | -0.278 | -0.3660 | No | ||
| 87 | NDUFB1 | Ndufb1-ps | 11756 | -0.306 | -0.3792 | No | ||
| 88 | ADAM17 | Adam17 | 11879 | -0.320 | -0.3813 | Yes | ||
| 89 | COX8A | Cox8a | 11885 | -0.321 | -0.3758 | Yes | ||
| 90 | ATF6 | Atf6 | 11898 | -0.324 | -0.3708 | Yes | ||
| 91 | UQCRB | Uqcrb | 11992 | -0.334 | -0.3708 | Yes | ||
| 92 | CACNA1F | Cacna1f | 12021 | -0.338 | -0.3665 | Yes | ||
| 93 | CALML3 | Calml3 | 12117 | -0.346 | -0.3664 | Yes | ||
| 94 | BAD | Bad | 12251 | -0.360 | -0.3686 | Yes | ||
| 95 | ATP5MC3 | Atp5g3 | 12291 | -0.365 | -0.3645 | Yes | ||
| 96 | IDE | Ide | 12311 | -0.368 | -0.3591 | Yes | ||
| 97 | NDUFB6 | Ndufb6 | 12354 | -0.372 | -0.3551 | Yes | ||
| 98 | NDUFA3 | Ndufa3 | 12359 | -0.373 | -0.3486 | Yes | ||
| 99 | CALM3 | Calm3 | 12399 | -0.378 | -0.3443 | Yes | ||
| 100 | ATP5F1D | Atp5d | 12403 | -0.379 | -0.3377 | Yes | ||
| 101 | ATP5PD | Atp5h | 12479 | -0.388 | -0.3356 | Yes | ||
| 102 | COX7B | Cox7b | 12512 | -0.392 | -0.3306 | Yes | ||
| 103 | COX7A1 | Cox7a1 | 12522 | -0.394 | -0.3240 | Yes | ||
| 104 | MT-CO1 | mt-Co1 | 12585 | -0.401 | -0.3208 | Yes | ||
| 105 | CASP8 | Casp8 | 12626 | -0.405 | -0.3161 | Yes | ||
| 106 | CAPN1 | Capn1 | 12683 | -0.413 | -0.3123 | Yes | ||
| 107 | CDK5 | Cdk5 | 12747 | -0.423 | -0.3088 | Yes | ||
| 108 | FAS | Fas | 12823 | -0.434 | -0.3058 | Yes | ||
| 109 | MT-ATP6 | mt-Atp6 | 12940 | -0.443 | -0.3053 | Yes | ||
| 110 | APP | App | 13008 | -0.448 | -0.3016 | Yes | ||
| 111 | ADAM10 | Adam10 | 13077 | -0.458 | -0.2978 | Yes | ||
| 112 | PPP3CA | Ppp3ca | 13145 | -0.468 | -0.2937 | Yes | ||
| 113 | NDUFB7 | Ndufb7 | 13279 | -0.492 | -0.2934 | Yes | ||
| 114 | MT-CO3 | mt-Co3 | 13320 | -0.498 | -0.2870 | Yes | ||
| 115 | BID | Bid | 13410 | -0.503 | -0.2837 | Yes | ||
| 116 | ATP5F1E | Atp5e | 13433 | -0.508 | -0.2760 | Yes | ||
| 117 | COX6C | Cox6c | 13521 | -0.523 | -0.2722 | Yes | ||
| 118 | CACNA1C | Cacna1c | 13597 | -0.532 | -0.2675 | Yes | ||
| 119 | NDUFB4 | Ndufb4 | 13601 | -0.534 | -0.2580 | Yes | ||
| 120 | NDUFS8 | Ndufs8 | 13664 | -0.545 | -0.2522 | Yes | ||
| 121 | NDUFA1 | Ndufa1 | 13759 | -0.566 | -0.2481 | Yes | ||
| 122 | ATP5PF | Atp5j | 13797 | -0.573 | -0.2402 | Yes | ||
| 123 | CALM1 | Calm1 | 13895 | -0.591 | -0.2358 | Yes | ||
| 124 | CHP1 | Chp1 | 13957 | -0.605 | -0.2288 | Yes | ||
| 125 | NDUFS6 | Ndufs6 | 13984 | -0.610 | -0.2195 | Yes | ||
| 126 | NDUFA2 | Ndufa2 | 14058 | -0.626 | -0.2130 | Yes | ||
| 127 | LRP1 | Lrp1 | 14082 | -0.633 | -0.2030 | Yes | ||
| 128 | NDUFV3 | Ndufv3 | 14135 | -0.645 | -0.1948 | Yes | ||
| 129 | UQCR10 | Uqcr10 | 14215 | -0.666 | -0.1879 | Yes | ||
| 130 | NDUFA4 | Ndufa4 | 14228 | -0.671 | -0.1766 | Yes | ||
| 131 | ITPR3 | Itpr3 | 14234 | -0.673 | -0.1647 | Yes | ||
| 132 | COX6A1 | Cox6a1 | 14276 | -0.686 | -0.1550 | Yes | ||
| 133 | UQCRQ | Uqcrq | 14286 | -0.690 | -0.1431 | Yes | ||
| 134 | NDUFC1 | Ndufc1 | 14306 | -0.695 | -0.1318 | Yes | ||
| 135 | NDUFA5 | Ndufa5 | 14324 | -0.701 | -0.1203 | Yes | ||
| 136 | ATP2A3 | Atp2a3 | 14880 | -0.934 | -0.1395 | Yes | ||
| 137 | EIF2AK3 | Eif2ak3 | 14920 | -0.959 | -0.1247 | Yes | ||
| 138 | NDUFS5 | Ndufs5 | 14975 | -0.990 | -0.1103 | Yes | ||
| 139 | MAPT | Mapt | 14984 | -0.995 | -0.0929 | Yes | ||
| 140 | GRIN1 | Grin1 | 14989 | -0.997 | -0.0751 | Yes | ||
| 141 | RYR3 | Ryr3 | 15002 | -1.010 | -0.0577 | Yes | ||
| 142 | PLCB1 | Plcb1 | 15093 | -1.088 | -0.0439 | Yes | ||
| 143 | PLCB2 | Plcb2 | 15231 | -1.247 | -0.0303 | Yes | ||
| 144 | CACNA1S | Cacna1s | 15291 | -1.364 | -0.0095 | Yes | ||
| 145 | MME | Mme | 15293 | -1.364 | 0.0151 | Yes |