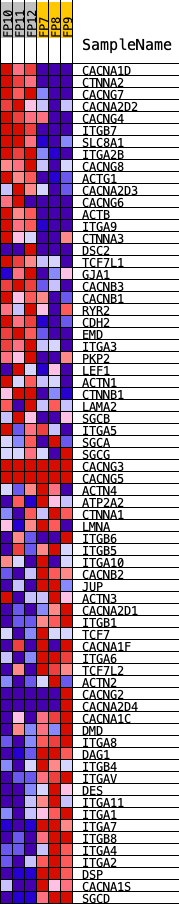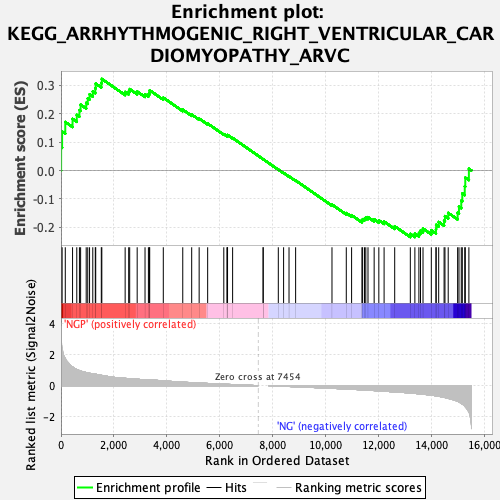
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_NG.phenotype.cls#NGP_versus_NG_repos |
| Phenotype | phenotype.cls#NGP_versus_NG_repos |
| Upregulated in class | NGP |
| GeneSet | KEGG_ARRHYTHMOGENIC_RIGHT_VENTRICULAR_CARDIOMYOPATHY_ARVC |
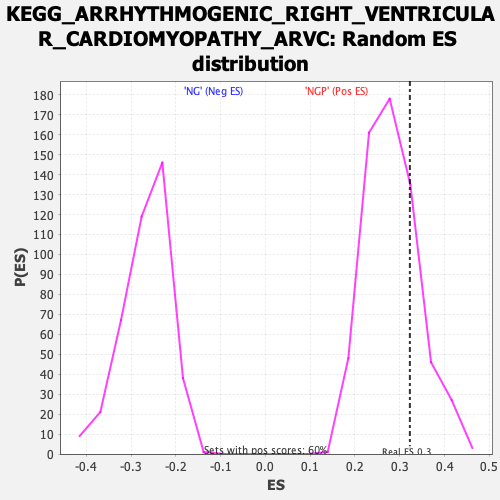
| Enrichment Score (ES) | 0.32255676 |
| Normalized Enrichment Score (NES) | 1.1456864 |
| Nominal p-value | 0.21035059 |
| FDR q-value | 0.7760984 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CACNA1D | Cacna1d | 5 | 3.519 | 0.0822 | Yes | ||
| 2 | CTNNA2 | Ctnna2 | 42 | 2.455 | 0.1375 | Yes | ||
| 3 | CACNG7 | Cacng7 | 162 | 1.757 | 0.1710 | Yes | ||
| 4 | CACNA2D2 | Cacna2d2 | 437 | 1.215 | 0.1818 | Yes | ||
| 5 | CACNG4 | Cacng4 | 597 | 1.047 | 0.1960 | Yes | ||
| 6 | ITGB7 | Itgb7 | 697 | 0.982 | 0.2127 | Yes | ||
| 7 | SLC8A1 | Slc8a1 | 742 | 0.945 | 0.2320 | Yes | ||
| 8 | ITGA2B | Itga2b | 953 | 0.849 | 0.2383 | Yes | ||
| 9 | CACNG8 | Cacng8 | 1012 | 0.822 | 0.2538 | Yes | ||
| 10 | ACTG1 | Actg1 | 1080 | 0.805 | 0.2684 | Yes | ||
| 11 | CACNA2D3 | Cacna2d3 | 1202 | 0.763 | 0.2784 | Yes | ||
| 12 | CACNG6 | Cacng6 | 1297 | 0.741 | 0.2897 | Yes | ||
| 13 | ACTB | Actb | 1313 | 0.740 | 0.3061 | Yes | ||
| 14 | ITGA9 | Itga9 | 1524 | 0.669 | 0.3082 | Yes | ||
| 15 | CTNNA3 | Ctnna3 | 1544 | 0.663 | 0.3226 | Yes | ||
| 16 | DSC2 | Dsc2 | 2426 | 0.473 | 0.2767 | No | ||
| 17 | TCF7L1 | Tcf7l1 | 2560 | 0.450 | 0.2786 | No | ||
| 18 | GJA1 | Gja6 | 2598 | 0.446 | 0.2867 | No | ||
| 19 | CACNB3 | Cacnb3 | 2881 | 0.418 | 0.2782 | No | ||
| 20 | CACNB1 | Cacnb1 | 3178 | 0.388 | 0.2682 | No | ||
| 21 | RYR2 | Ryr2 | 3306 | 0.368 | 0.2686 | No | ||
| 22 | CDH2 | Cdh2 | 3343 | 0.362 | 0.2748 | No | ||
| 23 | EMD | Emd | 3355 | 0.360 | 0.2825 | No | ||
| 24 | ITGA3 | Itga3 | 3869 | 0.310 | 0.2566 | No | ||
| 25 | PKP2 | Pkp2 | 4605 | 0.227 | 0.2143 | No | ||
| 26 | LEF1 | Lef1 | 4943 | 0.195 | 0.1971 | No | ||
| 27 | ACTN1 | Actn1 | 5225 | 0.170 | 0.1829 | No | ||
| 28 | CTNNB1 | Ctnnb1 | 5547 | 0.143 | 0.1655 | No | ||
| 29 | LAMA2 | Lama2 | 6157 | 0.094 | 0.1283 | No | ||
| 30 | SGCB | Sgcb | 6279 | 0.085 | 0.1225 | No | ||
| 31 | ITGA5 | Itga5 | 6290 | 0.084 | 0.1238 | No | ||
| 32 | SGCA | Sgca | 6299 | 0.084 | 0.1253 | No | ||
| 33 | SGCG | Sgcg | 6491 | 0.071 | 0.1146 | No | ||
| 34 | CACNG3 | Cacng3 | 7645 | 0.000 | 0.0400 | No | ||
| 35 | CACNG5 | Cacng5 | 7646 | 0.000 | 0.0400 | No | ||
| 36 | ACTN4 | Actn4 | 8219 | -0.025 | 0.0036 | No | ||
| 37 | ATP2A2 | Atp2a2 | 8419 | -0.038 | -0.0084 | No | ||
| 38 | CTNNA1 | Ctnna1 | 8624 | -0.052 | -0.0204 | No | ||
| 39 | LMNA | Lmna | 8874 | -0.070 | -0.0348 | No | ||
| 40 | ITGB6 | Itgb6 | 10249 | -0.172 | -0.1197 | No | ||
| 41 | ITGB5 | Itgb5 | 10790 | -0.214 | -0.1496 | No | ||
| 42 | ITGA10 | Gm42957 | 10991 | -0.232 | -0.1571 | No | ||
| 43 | CACNB2 | Cacnb2 | 11382 | -0.270 | -0.1760 | No | ||
| 44 | JUP | Jup | 11408 | -0.273 | -0.1712 | No | ||
| 45 | ACTN3 | Actn3 | 11480 | -0.279 | -0.1693 | No | ||
| 46 | CACNA2D1 | Cacna2d1 | 11528 | -0.283 | -0.1657 | No | ||
| 47 | ITGB1 | Itgb1 | 11611 | -0.292 | -0.1641 | No | ||
| 48 | TCF7 | Tcf7 | 11844 | -0.317 | -0.1717 | No | ||
| 49 | CACNA1F | Cacna1f | 12021 | -0.338 | -0.1752 | No | ||
| 50 | ITGA6 | Itga6 | 12220 | -0.357 | -0.1796 | No | ||
| 51 | TCF7L2 | Tcf7l2 | 12621 | -0.404 | -0.1960 | No | ||
| 52 | ACTN2 | Actn2 | 13212 | -0.478 | -0.2230 | No | ||
| 53 | CACNG2 | Cacng2 | 13384 | -0.501 | -0.2223 | No | ||
| 54 | CACNA2D4 | Cacna2d4 | 13536 | -0.524 | -0.2197 | No | ||
| 55 | CACNA1C | Cacna1c | 13597 | -0.532 | -0.2111 | No | ||
| 56 | DMD | Dmd | 13692 | -0.551 | -0.2043 | No | ||
| 57 | ITGA8 | Itga8 | 14006 | -0.615 | -0.2101 | No | ||
| 58 | DAG1 | Dag1 | 14182 | -0.657 | -0.2060 | No | ||
| 59 | ITGB4 | Itgb4 | 14189 | -0.658 | -0.1910 | No | ||
| 60 | ITGAV | Itgav | 14285 | -0.689 | -0.1810 | No | ||
| 61 | DES | Des | 14489 | -0.757 | -0.1764 | No | ||
| 62 | ITGA11 | Itga11 | 14522 | -0.769 | -0.1604 | No | ||
| 63 | ITGA1 | Itga1 | 14645 | -0.821 | -0.1491 | No | ||
| 64 | ITGA7 | Itga7 | 15001 | -1.007 | -0.1484 | No | ||
| 65 | ITGB8 | Itgb8 | 15052 | -1.054 | -0.1269 | No | ||
| 66 | ITGA4 | Itga4 | 15139 | -1.134 | -0.1059 | No | ||
| 67 | ITGA2 | Itga2 | 15178 | -1.187 | -0.0805 | No | ||
| 68 | DSP | Dsp | 15271 | -1.317 | -0.0556 | No | ||
| 69 | CACNA1S | Cacna1s | 15291 | -1.364 | -0.0248 | No | ||
| 70 | SGCD | Sgcd | 15427 | -1.700 | 0.0063 | No |