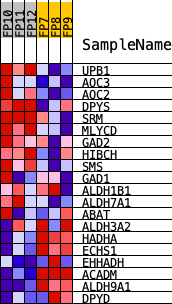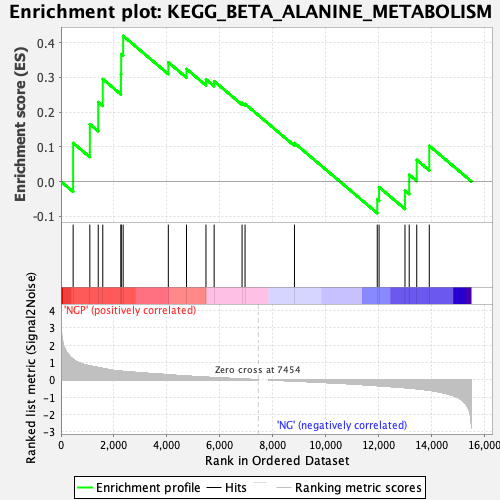
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_NG.phenotype.cls#NGP_versus_NG_repos |
| Phenotype | phenotype.cls#NGP_versus_NG_repos |
| Upregulated in class | NGP |
| GeneSet | KEGG_BETA_ALANINE_METABOLISM |
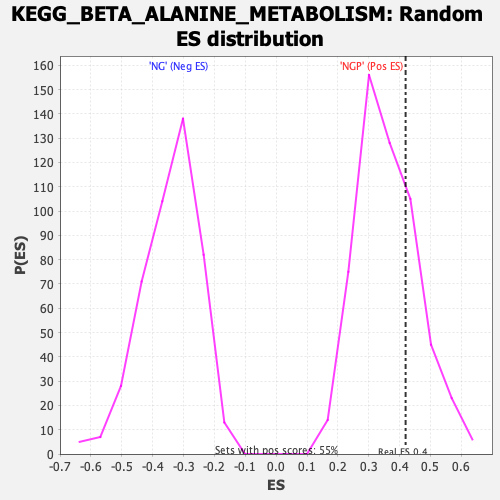
| Enrichment Score (ES) | 0.41939315 |
| Normalized Enrichment Score (NES) | 1.1631091 |
| Nominal p-value | 0.26449275 |
| FDR q-value | 0.7559915 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | UPB1 | Upb1 | 459 | 1.186 | 0.1110 | Yes | ||
| 2 | AOC3 | Aoc3 | 1091 | 0.801 | 0.1652 | Yes | ||
| 3 | AOC2 | Aoc2 | 1408 | 0.708 | 0.2288 | Yes | ||
| 4 | DPYS | Dpys | 1580 | 0.652 | 0.2951 | Yes | ||
| 5 | SRM | Srm | 2267 | 0.493 | 0.3093 | Yes | ||
| 6 | MLYCD | Mlycd | 2276 | 0.492 | 0.3670 | Yes | ||
| 7 | GAD2 | Gad2 | 2351 | 0.482 | 0.4194 | Yes | ||
| 8 | HIBCH | Hibch | 4059 | 0.286 | 0.3433 | No | ||
| 9 | SMS | Sms | 4747 | 0.212 | 0.3241 | No | ||
| 10 | GAD1 | Gad1 | 5483 | 0.149 | 0.2944 | No | ||
| 11 | ALDH1B1 | Aldh1b1 | 5792 | 0.122 | 0.2890 | No | ||
| 12 | ALDH7A1 | Aldh7a1 | 6848 | 0.045 | 0.2262 | No | ||
| 13 | ABAT | Abat | 6961 | 0.037 | 0.2234 | No | ||
| 14 | ALDH3A2 | Aldh3a2 | 8830 | -0.067 | 0.1108 | No | ||
| 15 | HADHA | Hadha | 11961 | -0.331 | -0.0518 | No | ||
| 16 | ECHS1 | Echs1 | 12030 | -0.339 | -0.0160 | No | ||
| 17 | EHHADH | Ehhadh | 13010 | -0.448 | -0.0260 | No | ||
| 18 | ACADM | Acadm | 13170 | -0.472 | 0.0198 | No | ||
| 19 | ALDH9A1 | Aldh9a1 | 13455 | -0.514 | 0.0624 | No | ||
| 20 | DPYD | Dpyd | 13929 | -0.599 | 0.1029 | No |