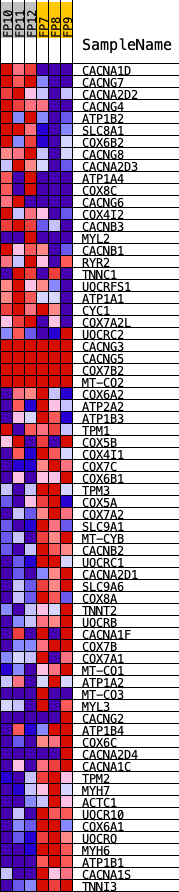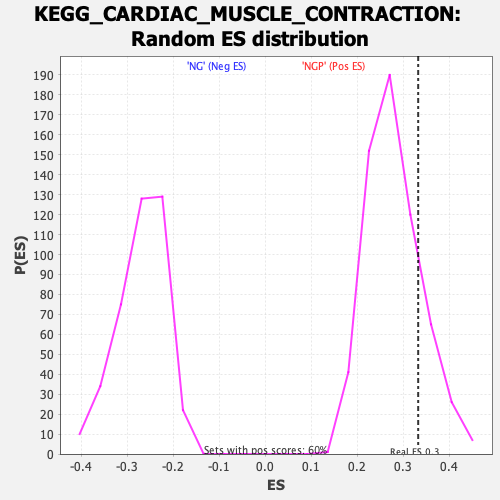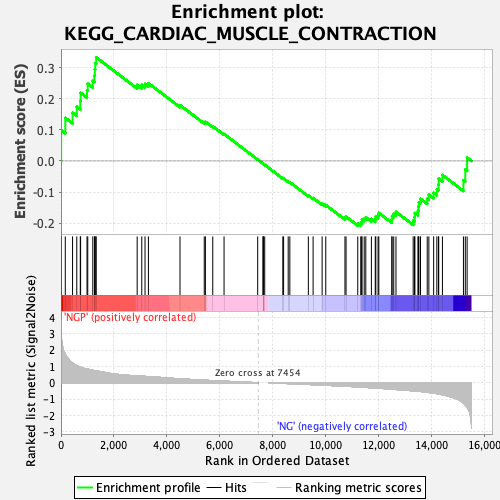
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_NG.phenotype.cls#NGP_versus_NG_repos |
| Phenotype | phenotype.cls#NGP_versus_NG_repos |
| Upregulated in class | NGP |
| GeneSet | KEGG_CARDIAC_MUSCLE_CONTRACTION |
| Enrichment Score (ES) | 0.33289823 |
| Normalized Enrichment Score (NES) | 1.190799 |
| Nominal p-value | 0.179402 |
| FDR q-value | 0.74799955 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CACNA1D | Cacna1d | 5 | 3.519 | 0.0990 | Yes | ||
| 2 | CACNG7 | Cacng7 | 162 | 1.757 | 0.1385 | Yes | ||
| 3 | CACNA2D2 | Cacna2d2 | 437 | 1.215 | 0.1551 | Yes | ||
| 4 | CACNG4 | Cacng4 | 597 | 1.047 | 0.1744 | Yes | ||
| 5 | ATP1B2 | Atp1b2 | 729 | 0.958 | 0.1930 | Yes | ||
| 6 | SLC8A1 | Slc8a1 | 742 | 0.945 | 0.2188 | Yes | ||
| 7 | COX6B2 | Cox6b2 | 984 | 0.837 | 0.2269 | Yes | ||
| 8 | CACNG8 | Cacng8 | 1012 | 0.822 | 0.2483 | Yes | ||
| 9 | CACNA2D3 | Cacna2d3 | 1202 | 0.763 | 0.2576 | Yes | ||
| 10 | ATP1A4 | Atp1a4 | 1265 | 0.741 | 0.2746 | Yes | ||
| 11 | COX8C | Cox8b | 1280 | 0.741 | 0.2946 | Yes | ||
| 12 | CACNG6 | Cacng6 | 1297 | 0.741 | 0.3144 | Yes | ||
| 13 | COX4I2 | Cox4i2 | 1333 | 0.734 | 0.3329 | Yes | ||
| 14 | CACNB3 | Cacnb3 | 2881 | 0.418 | 0.2446 | No | ||
| 15 | MYL2 | Myl2 | 3055 | 0.401 | 0.2447 | No | ||
| 16 | CACNB1 | Cacnb1 | 3178 | 0.388 | 0.2478 | No | ||
| 17 | RYR2 | Ryr2 | 3306 | 0.368 | 0.2500 | No | ||
| 18 | TNNC1 | Tnnc1 | 4501 | 0.237 | 0.1794 | No | ||
| 19 | UQCRFS1 | Uqcrfs1 | 5418 | 0.155 | 0.1245 | No | ||
| 20 | ATP1A1 | Atp1a1 | 5461 | 0.150 | 0.1260 | No | ||
| 21 | CYC1 | Cyc1 | 5740 | 0.126 | 0.1116 | No | ||
| 22 | COX7A2L | Cox7a2l | 6167 | 0.093 | 0.0867 | No | ||
| 23 | UQCRC2 | Uqcrc2 | 7439 | 0.002 | 0.0045 | No | ||
| 24 | CACNG3 | Cacng3 | 7645 | 0.000 | -0.0088 | No | ||
| 25 | CACNG5 | Cacng5 | 7646 | 0.000 | -0.0088 | No | ||
| 26 | COX7B2 | Cox7b2 | 7652 | 0.000 | -0.0091 | No | ||
| 27 | MT-CO2 | mt-Co2 | 7705 | 0.000 | -0.0125 | No | ||
| 28 | COX6A2 | Cox6a2 | 8390 | -0.037 | -0.0557 | No | ||
| 29 | ATP2A2 | Atp2a2 | 8419 | -0.038 | -0.0564 | No | ||
| 30 | ATP1B3 | Atp1b3 | 8593 | -0.051 | -0.0662 | No | ||
| 31 | TPM1 | Tpm1 | 8655 | -0.055 | -0.0686 | No | ||
| 32 | COX5B | Cox5b | 9355 | -0.103 | -0.1109 | No | ||
| 33 | COX4I1 | Cox4i1 | 9535 | -0.118 | -0.1191 | No | ||
| 34 | COX7C | Cox7c | 9876 | -0.145 | -0.1370 | No | ||
| 35 | COX6B1 | Cox6b1 | 10014 | -0.155 | -0.1415 | No | ||
| 36 | TPM3 | Tpm3 | 10740 | -0.210 | -0.1825 | No | ||
| 37 | COX5A | Cox5a | 10781 | -0.213 | -0.1791 | No | ||
| 38 | COX7A2 | Cox7a2 | 11224 | -0.255 | -0.2005 | No | ||
| 39 | SLC9A1 | Slc9a1 | 11337 | -0.265 | -0.2002 | No | ||
| 40 | MT-CYB | mt-Cytb | 11374 | -0.270 | -0.1949 | No | ||
| 41 | CACNB2 | Cacnb2 | 11382 | -0.270 | -0.1878 | No | ||
| 42 | UQCRC1 | Uqcrc1 | 11468 | -0.278 | -0.1854 | No | ||
| 43 | CACNA2D1 | Cacna2d1 | 11528 | -0.283 | -0.1812 | No | ||
| 44 | SLC9A6 | Slc9a6 | 11740 | -0.305 | -0.1863 | No | ||
| 45 | COX8A | Cox8a | 11885 | -0.321 | -0.1865 | No | ||
| 46 | TNNT2 | Tnnt2 | 11907 | -0.324 | -0.1787 | No | ||
| 47 | UQCRB | Uqcrb | 11992 | -0.334 | -0.1747 | No | ||
| 48 | CACNA1F | Cacna1f | 12021 | -0.338 | -0.1670 | No | ||
| 49 | COX7B | Cox7b | 12512 | -0.392 | -0.1876 | No | ||
| 50 | COX7A1 | Cox7a1 | 12522 | -0.394 | -0.1771 | No | ||
| 51 | MT-CO1 | mt-Co1 | 12585 | -0.401 | -0.1698 | No | ||
| 52 | ATP1A2 | Atp1a2 | 12670 | -0.411 | -0.1636 | No | ||
| 53 | MT-CO3 | mt-Co3 | 13320 | -0.498 | -0.1916 | No | ||
| 54 | MYL3 | Myl3 | 13372 | -0.501 | -0.1807 | No | ||
| 55 | CACNG2 | Cacng2 | 13384 | -0.501 | -0.1673 | No | ||
| 56 | ATP1B4 | Atp1b4 | 13500 | -0.520 | -0.1600 | No | ||
| 57 | COX6C | Cox6c | 13521 | -0.523 | -0.1466 | No | ||
| 58 | CACNA2D4 | Cacna2d4 | 13536 | -0.524 | -0.1327 | No | ||
| 59 | CACNA1C | Cacna1c | 13597 | -0.532 | -0.1215 | No | ||
| 60 | TPM2 | Tpm2 | 13851 | -0.583 | -0.1215 | No | ||
| 61 | MYH7 | Myh7 | 13913 | -0.596 | -0.1086 | No | ||
| 62 | ACTC1 | Actc1 | 14094 | -0.635 | -0.1023 | No | ||
| 63 | UQCR10 | Uqcr10 | 14215 | -0.666 | -0.0913 | No | ||
| 64 | COX6A1 | Cox6a1 | 14276 | -0.686 | -0.0758 | No | ||
| 65 | UQCRQ | Uqcrq | 14286 | -0.690 | -0.0569 | No | ||
| 66 | MYH6 | Myh6 | 14426 | -0.735 | -0.0451 | No | ||
| 67 | ATP1B1 | Atp1b1 | 15221 | -1.235 | -0.0616 | No | ||
| 68 | CACNA1S | Cacna1s | 15291 | -1.364 | -0.0276 | No | ||
| 69 | TNNI3 | Tnni3 | 15360 | -1.511 | 0.0107 | No |