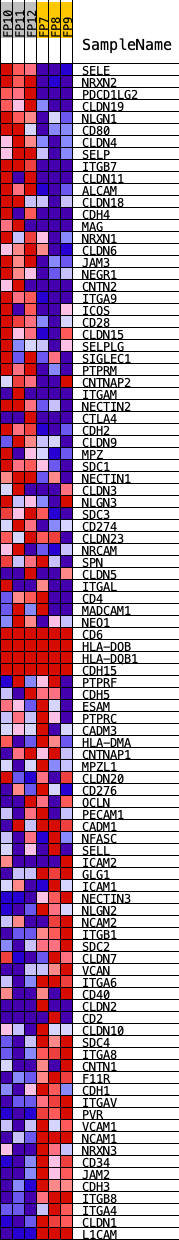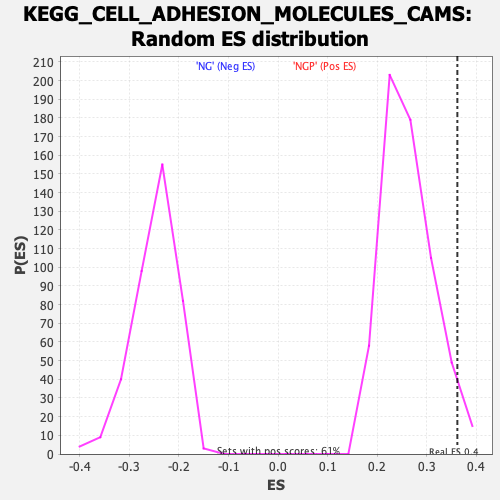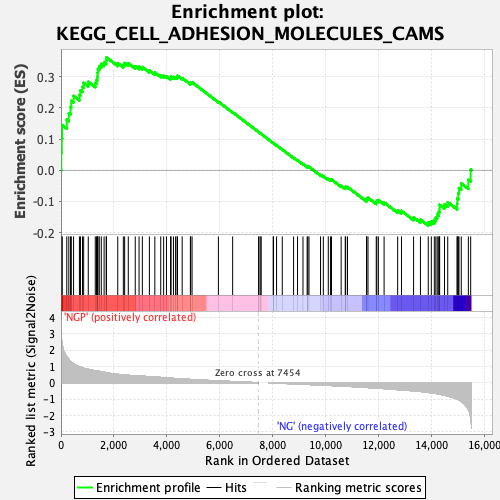
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_NG.phenotype.cls#NGP_versus_NG_repos |
| Phenotype | phenotype.cls#NGP_versus_NG_repos |
| Upregulated in class | NGP |
| GeneSet | KEGG_CELL_ADHESION_MOLECULES_CAMS |
| Enrichment Score (ES) | 0.36188367 |
| Normalized Enrichment Score (NES) | 1.3782372 |
| Nominal p-value | 0.036124796 |
| FDR q-value | 0.37496504 |
| FWER p-Value | 0.998 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SELE | Sele | 11 | 3.257 | 0.0575 | Yes | ||
| 2 | NRXN2 | Nrxn2 | 27 | 2.579 | 0.1026 | Yes | ||
| 3 | PDCD1LG2 | Pdcd1lg2 | 46 | 2.424 | 0.1448 | Yes | ||
| 4 | CLDN19 | Cldn19 | 218 | 1.583 | 0.1620 | Yes | ||
| 5 | NLGN1 | Nlgn1 | 295 | 1.438 | 0.1828 | Yes | ||
| 6 | CD80 | Cd80 | 361 | 1.311 | 0.2020 | Yes | ||
| 7 | CLDN4 | Cldn4 | 389 | 1.262 | 0.2228 | Yes | ||
| 8 | SELP | Selp | 475 | 1.170 | 0.2382 | Yes | ||
| 9 | ITGB7 | Itgb7 | 697 | 0.982 | 0.2414 | Yes | ||
| 10 | CLDN11 | Cldn11 | 736 | 0.950 | 0.2559 | Yes | ||
| 11 | ALCAM | Alcam | 813 | 0.906 | 0.2672 | Yes | ||
| 12 | CLDN18 | Cldn18 | 857 | 0.886 | 0.2803 | Yes | ||
| 13 | CDH4 | Cdh4 | 1031 | 0.820 | 0.2837 | Yes | ||
| 14 | MAG | Mag | 1296 | 0.741 | 0.2798 | Yes | ||
| 15 | NRXN1 | Nrxn1 | 1343 | 0.729 | 0.2899 | Yes | ||
| 16 | CLDN6 | Cldn6 | 1378 | 0.714 | 0.3004 | Yes | ||
| 17 | JAM3 | Jam3 | 1384 | 0.713 | 0.3128 | Yes | ||
| 18 | NEGR1 | Negr1 | 1397 | 0.711 | 0.3248 | Yes | ||
| 19 | CNTN2 | Cntn2 | 1448 | 0.692 | 0.3339 | Yes | ||
| 20 | ITGA9 | Itga9 | 1524 | 0.669 | 0.3410 | Yes | ||
| 21 | ICOS | Icos | 1630 | 0.639 | 0.3456 | Yes | ||
| 22 | CD28 | Cd28 | 1709 | 0.617 | 0.3516 | Yes | ||
| 23 | CLDN15 | Cldn15 | 1720 | 0.612 | 0.3619 | Yes | ||
| 24 | SELPLG | Selplg | 2146 | 0.513 | 0.3435 | No | ||
| 25 | SIGLEC1 | Siglec1 | 2357 | 0.480 | 0.3385 | No | ||
| 26 | PTPRM | Ptprm | 2407 | 0.476 | 0.3438 | No | ||
| 27 | CNTNAP2 | Cntnap2 | 2545 | 0.454 | 0.3430 | No | ||
| 28 | ITGAM | Itgam | 2808 | 0.428 | 0.3337 | No | ||
| 29 | NECTIN2 | Nectin2 | 2952 | 0.407 | 0.3317 | No | ||
| 30 | CTLA4 | Ctla4 | 3077 | 0.401 | 0.3308 | No | ||
| 31 | CDH2 | Cdh2 | 3343 | 0.362 | 0.3201 | No | ||
| 32 | CLDN9 | Cldn9 | 3549 | 0.352 | 0.3131 | No | ||
| 33 | MPZ | Mpz | 3772 | 0.324 | 0.3045 | No | ||
| 34 | SDC1 | Sdc1 | 3882 | 0.309 | 0.3030 | No | ||
| 35 | NECTIN1 | Nectin1 | 3989 | 0.295 | 0.3014 | No | ||
| 36 | CLDN3 | Cldn3 | 4145 | 0.277 | 0.2963 | No | ||
| 37 | NLGN3 | Nlgn3 | 4162 | 0.275 | 0.3002 | No | ||
| 38 | SDC3 | Sdc3 | 4253 | 0.265 | 0.2991 | No | ||
| 39 | CD274 | Cd274 | 4334 | 0.255 | 0.2984 | No | ||
| 40 | CLDN23 | Cldn23 | 4390 | 0.250 | 0.2993 | No | ||
| 41 | NRCAM | Nrcam | 4406 | 0.248 | 0.3028 | No | ||
| 42 | SPN | Spn | 4583 | 0.229 | 0.2955 | No | ||
| 43 | CLDN5 | Cldn5 | 4893 | 0.201 | 0.2790 | No | ||
| 44 | ITGAL | Itgal | 4903 | 0.200 | 0.2820 | No | ||
| 45 | CD4 | Cd4 | 4957 | 0.194 | 0.2820 | No | ||
| 46 | MADCAM1 | Madcam1 | 5953 | 0.111 | 0.2195 | No | ||
| 47 | NEO1 | Neo1 | 6492 | 0.071 | 0.1859 | No | ||
| 48 | CD6 | Cd6 | 7473 | 0.000 | 0.1224 | No | ||
| 49 | HLA-DOB | H2-Ob | 7489 | 0.000 | 0.1214 | No | ||
| 50 | HLA-DQB1 | H2-Ab1 | 7560 | 0.000 | 0.1169 | No | ||
| 51 | CDH15 | Cdh15 | 7565 | 0.000 | 0.1166 | No | ||
| 52 | PTPRF | Ptprf | 8025 | -0.012 | 0.0871 | No | ||
| 53 | CDH5 | Cdh5 | 8038 | -0.013 | 0.0866 | No | ||
| 54 | ESAM | Esam | 8153 | -0.021 | 0.0795 | No | ||
| 55 | PTPRC | Ptprc | 8369 | -0.036 | 0.0662 | No | ||
| 56 | CADM3 | Cadm3 | 8797 | -0.064 | 0.0397 | No | ||
| 57 | HLA-DMA | H2-DMa | 8946 | -0.075 | 0.0315 | No | ||
| 58 | CNTNAP1 | Cntnap1 | 9152 | -0.090 | 0.0198 | No | ||
| 59 | MPZL1 | Mpzl1 | 9309 | -0.101 | 0.0115 | No | ||
| 60 | CLDN20 | Cldn20 | 9336 | -0.102 | 0.0116 | No | ||
| 61 | CD276 | Cd276 | 9379 | -0.106 | 0.0108 | No | ||
| 62 | OCLN | Ocln | 9816 | -0.141 | -0.0150 | No | ||
| 63 | PECAM1 | Pecam1 | 9922 | -0.149 | -0.0191 | No | ||
| 64 | CADM1 | Cadm1 | 10109 | -0.161 | -0.0283 | No | ||
| 65 | NFASC | Nfasc | 10191 | -0.168 | -0.0305 | No | ||
| 66 | SELL | Sell | 10231 | -0.170 | -0.0300 | No | ||
| 67 | ICAM2 | Icam2 | 10596 | -0.199 | -0.0500 | No | ||
| 68 | GLG1 | Glg1 | 10750 | -0.211 | -0.0562 | No | ||
| 69 | ICAM1 | Icam1 | 10767 | -0.212 | -0.0534 | No | ||
| 70 | NECTIN3 | Nectin3 | 10836 | -0.218 | -0.0540 | No | ||
| 71 | NLGN2 | Nlgn2 | 11557 | -0.286 | -0.0955 | No | ||
| 72 | NCAM2 | Ncam2 | 11560 | -0.286 | -0.0905 | No | ||
| 73 | ITGB1 | Itgb1 | 11611 | -0.292 | -0.0885 | No | ||
| 74 | SDC2 | Sdc2 | 11923 | -0.326 | -0.1029 | No | ||
| 75 | CLDN7 | Cldn7 | 11945 | -0.329 | -0.0984 | No | ||
| 76 | VCAN | Vcan | 12004 | -0.336 | -0.0961 | No | ||
| 77 | ITGA6 | Itga6 | 12220 | -0.357 | -0.1037 | No | ||
| 78 | CD40 | Cd40 | 12733 | -0.421 | -0.1293 | No | ||
| 79 | CLDN2 | Cldn2 | 12878 | -0.438 | -0.1308 | No | ||
| 80 | CD2 | Cd2 | 13335 | -0.499 | -0.1515 | No | ||
| 81 | CLDN10 | Cldn10 | 13595 | -0.532 | -0.1588 | No | ||
| 82 | SDC4 | Sdc4 | 13890 | -0.590 | -0.1673 | No | ||
| 83 | ITGA8 | Itga8 | 14006 | -0.615 | -0.1637 | No | ||
| 84 | CNTN1 | Cntn1 | 14120 | -0.640 | -0.1596 | No | ||
| 85 | F11R | F11r | 14177 | -0.656 | -0.1515 | No | ||
| 86 | CDH1 | Cdh1 | 14230 | -0.671 | -0.1429 | No | ||
| 87 | ITGAV | Itgav | 14285 | -0.689 | -0.1341 | No | ||
| 88 | PVR | Pvr | 14318 | -0.699 | -0.1237 | No | ||
| 89 | VCAM1 | Vcam1 | 14319 | -0.699 | -0.1112 | No | ||
| 90 | NCAM1 | Ncam1 | 14505 | -0.764 | -0.1095 | No | ||
| 91 | NRXN3 | Nrxn3 | 14627 | -0.812 | -0.1029 | No | ||
| 92 | CD34 | Cd34 | 14982 | -0.995 | -0.1080 | No | ||
| 93 | JAM2 | Jam2 | 14988 | -0.996 | -0.0905 | No | ||
| 94 | CDH3 | Cdh3 | 15028 | -1.030 | -0.0747 | No | ||
| 95 | ITGB8 | Itgb8 | 15052 | -1.054 | -0.0573 | No | ||
| 96 | ITGA4 | Itga4 | 15139 | -1.134 | -0.0426 | No | ||
| 97 | CLDN1 | Cldn1 | 15407 | -1.638 | -0.0306 | No | ||
| 98 | L1CAM | L1cam | 15495 | -2.139 | 0.0019 | No |