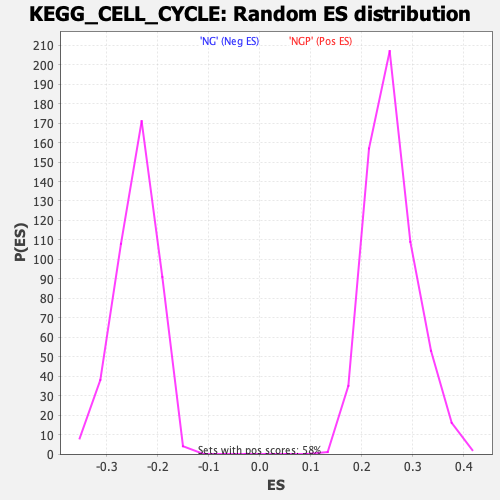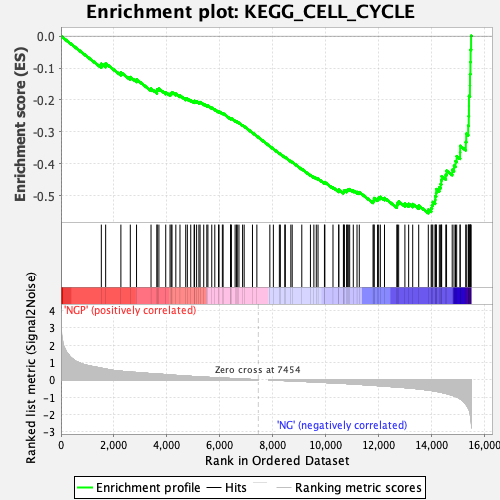
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_NG.phenotype.cls#NGP_versus_NG_repos |
| Phenotype | phenotype.cls#NGP_versus_NG_repos |
| Upregulated in class | NG |
| GeneSet | KEGG_CELL_CYCLE |
| Enrichment Score (ES) | -0.55503106 |
| Normalized Enrichment Score (NES) | -2.2941787 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CDKN1C | Cdkn1c | 1525 | 0.668 | -0.0865 | No | ||
| 2 | MDM2 | Mdm2 | 1689 | 0.623 | -0.0855 | No | ||
| 3 | CDK4 | Cdk4 | 2265 | 0.493 | -0.1136 | No | ||
| 4 | TGFB2 | Tgfb2 | 2621 | 0.442 | -0.1284 | No | ||
| 5 | MCM7 | Mcm7 | 2857 | 0.420 | -0.1358 | No | ||
| 6 | WEE2 | Wee2 | 3405 | 0.356 | -0.1647 | No | ||
| 7 | CDKN1A | Cdkn1a | 3630 | 0.341 | -0.1729 | No | ||
| 8 | ORC5 | Orc5 | 3635 | 0.341 | -0.1668 | No | ||
| 9 | CDC25A | Cdc25a | 3703 | 0.331 | -0.1650 | No | ||
| 10 | ORC3 | Orc3 | 3961 | 0.299 | -0.1761 | No | ||
| 11 | MAD1L1 | Mad1l1 | 4124 | 0.279 | -0.1814 | No | ||
| 12 | ZBTB17 | Zbtb17 | 4168 | 0.274 | -0.1791 | No | ||
| 13 | ATR | Atr | 4198 | 0.271 | -0.1759 | No | ||
| 14 | CCNB3 | Ccnb3 | 4343 | 0.254 | -0.1805 | No | ||
| 15 | MYC | Myc | 4500 | 0.237 | -0.1862 | No | ||
| 16 | PRKDC | Prkdc | 4717 | 0.216 | -0.1962 | No | ||
| 17 | E2F3 | E2f3 | 4778 | 0.209 | -0.1962 | No | ||
| 18 | TP53 | Trp53 | 4907 | 0.199 | -0.2008 | No | ||
| 19 | ABL1 | Abl1 | 5035 | 0.187 | -0.2056 | No | ||
| 20 | SMAD4 | Smad4 | 5051 | 0.185 | -0.2031 | No | ||
| 21 | HDAC2 | Hdac2 | 5129 | 0.179 | -0.2048 | No | ||
| 22 | ANAPC2 | Anapc2 | 5216 | 0.171 | -0.2072 | No | ||
| 23 | SFN | Sfn | 5260 | 0.167 | -0.2068 | No | ||
| 24 | CDC26 | Cdc26 | 5398 | 0.156 | -0.2128 | No | ||
| 25 | ATM | Atm | 5515 | 0.145 | -0.2176 | No | ||
| 26 | RB1 | Rb1 | 5556 | 0.143 | -0.2176 | No | ||
| 27 | SKP2 | Skp2 | 5702 | 0.131 | -0.2246 | No | ||
| 28 | E2F5 | E2f5 | 5817 | 0.121 | -0.2297 | No | ||
| 29 | CDK7 | Cdk7 | 5960 | 0.110 | -0.2369 | No | ||
| 30 | EP300 | Ep300 | 5976 | 0.109 | -0.2358 | No | ||
| 31 | CDC16 | Cdc16 | 6103 | 0.099 | -0.2421 | No | ||
| 32 | E2F4 | E2f4 | 6131 | 0.096 | -0.2421 | No | ||
| 33 | RBX1 | Rbx1 | 6414 | 0.077 | -0.2590 | No | ||
| 34 | SMAD3 | Smad3 | 6415 | 0.077 | -0.2576 | No | ||
| 35 | ANAPC4 | Anapc4 | 6445 | 0.075 | -0.2580 | No | ||
| 36 | MAD2L2 | Mad2l2 | 6585 | 0.065 | -0.2659 | No | ||
| 37 | CDC25B | Cdc25b | 6635 | 0.062 | -0.2679 | No | ||
| 38 | GADD45B | Gadd45b | 6667 | 0.059 | -0.2688 | No | ||
| 39 | SMC1B | Smc1b | 6730 | 0.054 | -0.2718 | No | ||
| 40 | SKP1 | Skp1a | 6868 | 0.043 | -0.2799 | No | ||
| 41 | ANAPC5 | Anapc5 | 6928 | 0.039 | -0.2830 | No | ||
| 42 | BUB3 | Bub3 | 7242 | 0.016 | -0.3030 | No | ||
| 43 | ORC2 | Orc2 | 7408 | 0.004 | -0.3136 | No | ||
| 44 | CDKN2C | Cdkn2c | 7899 | -0.003 | -0.3454 | No | ||
| 45 | ANAPC10 | Anapc10 | 8032 | -0.012 | -0.3537 | No | ||
| 46 | ORC4 | Orc4 | 8259 | -0.028 | -0.3679 | No | ||
| 47 | YWHAE | Ywhae | 8296 | -0.031 | -0.3696 | No | ||
| 48 | SMC1A | Smc1a | 8464 | -0.041 | -0.3797 | No | ||
| 49 | YWHAQ | Ywhaq | 8484 | -0.043 | -0.3801 | No | ||
| 50 | ANAPC1 | Anapc1 | 8693 | -0.058 | -0.3926 | No | ||
| 51 | ANAPC7 | Anapc7 | 8744 | -0.061 | -0.3947 | No | ||
| 52 | CCNH | Ccnh | 9107 | -0.087 | -0.4165 | No | ||
| 53 | GADD45A | Gadd45a | 9434 | -0.110 | -0.4356 | No | ||
| 54 | CCND3 | Ccnd3 | 9564 | -0.120 | -0.4418 | No | ||
| 55 | RAD21 | Rad21 | 9655 | -0.128 | -0.4452 | No | ||
| 56 | SMAD2 | Smad2 | 9721 | -0.132 | -0.4470 | No | ||
| 57 | CUL1 | Cul1 | 9964 | -0.152 | -0.4599 | No | ||
| 58 | CREBBP | Crebbp | 9980 | -0.152 | -0.4580 | No | ||
| 59 | DBF4 | Dbf4 | 10287 | -0.175 | -0.4746 | No | ||
| 60 | GSK3B | Gsk3b | 10504 | -0.192 | -0.4850 | No | ||
| 61 | FZR1 | Fzr1 | 10507 | -0.192 | -0.4816 | No | ||
| 62 | TFDP2 | Tfdp2 | 10681 | -0.205 | -0.4890 | No | ||
| 63 | RBL2 | Rbl2 | 10696 | -0.206 | -0.4861 | No | ||
| 64 | SMC3 | Smc3 | 10718 | -0.208 | -0.4835 | No | ||
| 65 | MCM2 | Mcm2 | 10802 | -0.215 | -0.4849 | No | ||
| 66 | YWHAG | Ywhag | 10830 | -0.218 | -0.4826 | No | ||
| 67 | TFDP1 | Tfdp1 | 10866 | -0.220 | -0.4808 | No | ||
| 68 | CDC27 | Cdc27 | 10918 | -0.225 | -0.4799 | No | ||
| 69 | PCNA | Pcna | 11055 | -0.238 | -0.4843 | No | ||
| 70 | STAG2 | Stag2 | 11196 | -0.252 | -0.4887 | No | ||
| 71 | E2F1 | E2f1 | 11282 | -0.260 | -0.4894 | No | ||
| 72 | ORC6 | Orc6 | 11802 | -0.311 | -0.5172 | No | ||
| 73 | CDC14A | Cdc14a | 11838 | -0.316 | -0.5136 | No | ||
| 74 | E2F2 | E2f2 | 11841 | -0.317 | -0.5078 | No | ||
| 75 | STAG1 | Stag1 | 11973 | -0.332 | -0.5102 | No | ||
| 76 | YWHAB | Ywhab | 12014 | -0.337 | -0.5065 | No | ||
| 77 | CDC23 | Cdc23 | 12078 | -0.344 | -0.5042 | No | ||
| 78 | YWHAZ | Ywhaz | 12235 | -0.358 | -0.5076 | No | ||
| 79 | CHEK2 | Chek2 | 12702 | -0.416 | -0.5301 | No | ||
| 80 | HDAC1 | Hdac1 | 12724 | -0.420 | -0.5236 | No | ||
| 81 | TGFB1 | Tgfb1 | 12771 | -0.426 | -0.5187 | No | ||
| 82 | RBL1 | Rbl1 | 13007 | -0.448 | -0.5256 | No | ||
| 83 | YWHAH | Ywhah | 13149 | -0.468 | -0.5260 | No | ||
| 84 | ANAPC13 | Anapc13 | 13302 | -0.496 | -0.5266 | No | ||
| 85 | CDK2 | Cdk2 | 13529 | -0.524 | -0.5315 | No | ||
| 86 | CCNE1 | Ccne1 | 13892 | -0.591 | -0.5440 | Yes | ||
| 87 | MAD2L1 | Mad2l1 | 14001 | -0.614 | -0.5396 | Yes | ||
| 88 | CHEK1 | Chek1 | 14031 | -0.620 | -0.5299 | Yes | ||
| 89 | GADD45G | Gadd45g | 14062 | -0.627 | -0.5202 | Yes | ||
| 90 | CDKN2D | Cdkn2d | 14151 | -0.649 | -0.5138 | Yes | ||
| 91 | CCNA1 | Ccna1 | 14156 | -0.651 | -0.5019 | Yes | ||
| 92 | CDKN1B | Cdkn1b | 14187 | -0.658 | -0.4916 | Yes | ||
| 93 | MCM6 | Mcm6 | 14196 | -0.661 | -0.4798 | Yes | ||
| 94 | CCNE2 | Ccne2 | 14309 | -0.696 | -0.4741 | Yes | ||
| 95 | PKMYT1 | Pkmyt1 | 14359 | -0.714 | -0.4640 | Yes | ||
| 96 | CDC7 | Cdc7 | 14384 | -0.721 | -0.4521 | Yes | ||
| 97 | CDK1 | Cdk1 | 14397 | -0.727 | -0.4393 | Yes | ||
| 98 | MCM4 | Mcm4 | 14554 | -0.779 | -0.4349 | Yes | ||
| 99 | ORC1 | Orc1 | 14582 | -0.790 | -0.4220 | Yes | ||
| 100 | CDK6 | Cdk6 | 14796 | -0.885 | -0.4193 | Yes | ||
| 101 | MCM3 | Mcm3 | 14865 | -0.927 | -0.4064 | Yes | ||
| 102 | CCND1 | Ccnd1 | 14921 | -0.960 | -0.3921 | Yes | ||
| 103 | WEE1 | Wee1 | 14965 | -0.985 | -0.3766 | Yes | ||
| 104 | MCM5 | Mcm5 | 15094 | -1.089 | -0.3646 | Yes | ||
| 105 | CDC20 | Cdc20 | 15097 | -1.097 | -0.3443 | Yes | ||
| 106 | TGFB3 | Tgfb3 | 15310 | -1.400 | -0.3320 | Yes | ||
| 107 | CDC45 | Cdc45 | 15327 | -1.438 | -0.3062 | Yes | ||
| 108 | CDC25C | Cdc25c | 15401 | -1.621 | -0.2807 | Yes | ||
| 109 | ESPL1 | Espl1 | 15422 | -1.681 | -0.2507 | Yes | ||
| 110 | TTK | Ttk | 15429 | -1.705 | -0.2193 | Yes | ||
| 111 | CDC6 | Cdc6 | 15430 | -1.707 | -0.1875 | Yes | ||
| 112 | PLK1 | Plk1 | 15467 | -1.900 | -0.1545 | Yes | ||
| 113 | CCNB2 | Ccnb2 | 15470 | -1.905 | -0.1191 | Yes | ||
| 114 | BUB1 | Bub1 | 15485 | -2.064 | -0.0815 | Yes | ||
| 115 | CCNB1 | Ccnb1 | 15492 | -2.124 | -0.0424 | Yes | ||
| 116 | CCNA2 | Ccna2 | 15511 | -2.386 | 0.0009 | Yes |