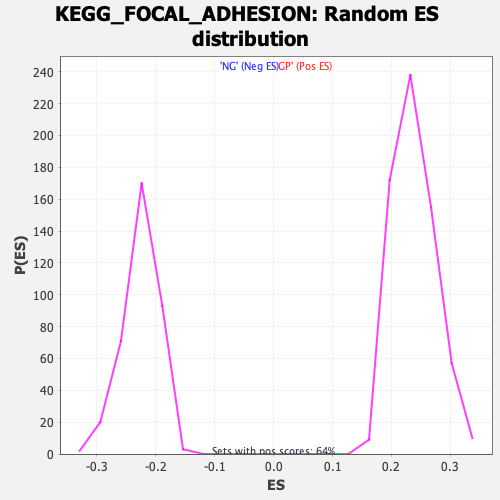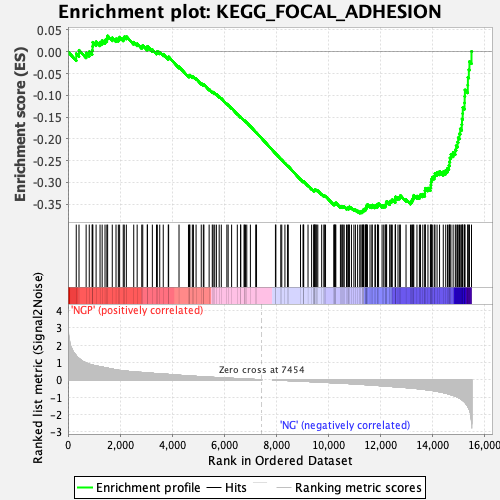
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_NG.phenotype.cls#NGP_versus_NG_repos |
| Phenotype | phenotype.cls#NGP_versus_NG_repos |
| Upregulated in class | NG |
| GeneSet | KEGG_FOCAL_ADHESION |
| Enrichment Score (ES) | -0.3705011 |
| Normalized Enrichment Score (NES) | -1.6393081 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.07905926 |
| FWER p-Value | 0.425 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | COMP | Comp | 317 | 1.391 | -0.0042 | No | ||
| 2 | PIK3CD | Pik3cd | 423 | 1.233 | 0.0035 | No | ||
| 3 | ITGB7 | Itgb7 | 697 | 0.982 | -0.0027 | No | ||
| 4 | ZYX | Zyx | 817 | 0.904 | 0.0003 | No | ||
| 5 | KDR | Kdr | 927 | 0.859 | 0.0033 | No | ||
| 6 | SHC3 | Shc3 | 945 | 0.853 | 0.0123 | No | ||
| 7 | ITGA2B | Itga2b | 953 | 0.849 | 0.0219 | No | ||
| 8 | ACTG1 | Actg1 | 1080 | 0.805 | 0.0232 | No | ||
| 9 | BCAR1 | Bcar1 | 1232 | 0.750 | 0.0222 | No | ||
| 10 | ACTB | Actb | 1313 | 0.740 | 0.0258 | No | ||
| 11 | THBS1 | Thbs1 | 1423 | 0.703 | 0.0270 | No | ||
| 12 | PAK5 | Pak7 | 1494 | 0.678 | 0.0304 | No | ||
| 13 | ITGA9 | Itga9 | 1524 | 0.669 | 0.0364 | No | ||
| 14 | ILK | Ilk | 1702 | 0.620 | 0.0322 | No | ||
| 15 | PARVG | Parvg | 1846 | 0.574 | 0.0297 | No | ||
| 16 | PDGFRA | Pdgfra | 1936 | 0.552 | 0.0304 | No | ||
| 17 | PRKCG | Prkcg | 1985 | 0.540 | 0.0337 | No | ||
| 18 | LAMC3 | Lamc3 | 2124 | 0.517 | 0.0308 | No | ||
| 19 | COL11A2 | Col11a2 | 2163 | 0.509 | 0.0343 | No | ||
| 20 | LAMB1 | Lamb1 | 2243 | 0.497 | 0.0350 | No | ||
| 21 | VAV3 | Vav3 | 2527 | 0.457 | 0.0220 | No | ||
| 22 | COL6A6 | Col6a6 | 2659 | 0.440 | 0.0187 | No | ||
| 23 | RELN | Reln | 2836 | 0.424 | 0.0122 | No | ||
| 24 | VCL | Vcl | 2874 | 0.419 | 0.0147 | No | ||
| 25 | VTN | Vtn | 3047 | 0.401 | 0.0083 | No | ||
| 26 | MYL2 | Myl2 | 3055 | 0.401 | 0.0125 | No | ||
| 27 | TNXB | Tnxb | 3243 | 0.378 | 0.0048 | No | ||
| 28 | VAV1 | Vav1 | 3402 | 0.356 | -0.0013 | No | ||
| 29 | IBSP | Ibsp | 3434 | 0.356 | 0.0009 | No | ||
| 30 | MYL10 | Myl10 | 3527 | 0.356 | -0.0009 | No | ||
| 31 | RAC2 | Rac2 | 3662 | 0.338 | -0.0056 | No | ||
| 32 | FLNA | Flna | 3855 | 0.312 | -0.0145 | No | ||
| 33 | ITGA3 | Itga3 | 3869 | 0.310 | -0.0116 | No | ||
| 34 | PGF | Pgf | 4268 | 0.263 | -0.0345 | No | ||
| 35 | MYLK2 | Mylk2 | 4632 | 0.225 | -0.0555 | No | ||
| 36 | VASP | Vasp | 4673 | 0.220 | -0.0555 | No | ||
| 37 | VAV2 | Vav2 | 4685 | 0.219 | -0.0536 | No | ||
| 38 | MAPK8 | Mapk8 | 4781 | 0.209 | -0.0574 | No | ||
| 39 | VEGFB | Vegfb | 4822 | 0.206 | -0.0575 | No | ||
| 40 | ERBB2 | Erbb2 | 4924 | 0.197 | -0.0618 | No | ||
| 41 | PAK4 | Pak4 | 5123 | 0.179 | -0.0726 | No | ||
| 42 | VWF | Vwf | 5203 | 0.173 | -0.0757 | No | ||
| 43 | ACTN1 | Actn1 | 5225 | 0.170 | -0.0750 | No | ||
| 44 | TNN | Tnn | 5426 | 0.154 | -0.0863 | No | ||
| 45 | CTNNB1 | Ctnnb1 | 5547 | 0.143 | -0.0924 | No | ||
| 46 | MAPK3 | Mapk3 | 5579 | 0.140 | -0.0928 | No | ||
| 47 | LAMA1 | Lama1 | 5640 | 0.135 | -0.0951 | No | ||
| 48 | MYL9 | Myl9 | 5705 | 0.130 | -0.0977 | No | ||
| 49 | PDGFC | Pdgfc | 5812 | 0.121 | -0.1032 | No | ||
| 50 | PARVB | Parvb | 5894 | 0.116 | -0.1071 | No | ||
| 51 | AKT1 | Akt1 | 6108 | 0.098 | -0.1198 | No | ||
| 52 | LAMA2 | Lama2 | 6157 | 0.094 | -0.1218 | No | ||
| 53 | ITGA5 | Itga5 | 6290 | 0.084 | -0.1295 | No | ||
| 54 | AKT2 | Akt2 | 6509 | 0.070 | -0.1428 | No | ||
| 55 | XIAP | Xiap | 6636 | 0.061 | -0.1503 | No | ||
| 56 | COL2A1 | Col2a1 | 6640 | 0.061 | -0.1498 | No | ||
| 57 | ARHGAP35 | Arhgap35 | 6771 | 0.051 | -0.1577 | No | ||
| 58 | PAK6 | Pak6 | 6781 | 0.050 | -0.1577 | No | ||
| 59 | PAK1 | Pak1 | 6823 | 0.047 | -0.1598 | No | ||
| 60 | SRC | Src | 6873 | 0.043 | -0.1625 | No | ||
| 61 | BIRC2 | Birc2 | 7012 | 0.033 | -0.1711 | No | ||
| 62 | LAMB2 | Lamb2 | 7217 | 0.018 | -0.1842 | No | ||
| 63 | MYLPF | Mylpf | 7248 | 0.015 | -0.1860 | No | ||
| 64 | FLNC | Flnc | 7975 | -0.008 | -0.2332 | No | ||
| 65 | EGF | Egf | 7992 | -0.009 | -0.2342 | No | ||
| 66 | PPP1CC | Ppp1cc | 8181 | -0.023 | -0.2461 | No | ||
| 67 | ACTN4 | Actn4 | 8219 | -0.025 | -0.2483 | No | ||
| 68 | PIK3CA | Pik3ca | 8340 | -0.034 | -0.2557 | No | ||
| 69 | SOS1 | Sos1 | 8438 | -0.040 | -0.2615 | No | ||
| 70 | PTK2 | Ptk2 | 8469 | -0.041 | -0.2630 | No | ||
| 71 | CAV3 | Cav3 | 8939 | -0.075 | -0.2927 | No | ||
| 72 | CRKL | Crkl | 9039 | -0.082 | -0.2982 | No | ||
| 73 | MAPK9 | Mapk9 | 9049 | -0.083 | -0.2978 | No | ||
| 74 | SOS2 | Sos2 | 9062 | -0.084 | -0.2976 | No | ||
| 75 | PPP1R12A | Ppp1r12a | 9228 | -0.094 | -0.3072 | No | ||
| 76 | RAP1A | Rap1a | 9366 | -0.105 | -0.3149 | No | ||
| 77 | PXN | Pxn | 9450 | -0.111 | -0.3190 | No | ||
| 78 | HGF | Hgf | 9454 | -0.111 | -0.3179 | No | ||
| 79 | PIP5K1C | Pip5k1c | 9486 | -0.113 | -0.3186 | No | ||
| 80 | RAF1 | Raf1 | 9494 | -0.114 | -0.3177 | No | ||
| 81 | MYL12A | Myl12a | 9495 | -0.114 | -0.3164 | No | ||
| 82 | GRB2 | Grb2 | 9534 | -0.118 | -0.3174 | No | ||
| 83 | CCND3 | Ccnd3 | 9564 | -0.120 | -0.3179 | No | ||
| 84 | VEGFC | Vegfc | 9604 | -0.123 | -0.3190 | No | ||
| 85 | RAPGEF1 | Rapgef1 | 9753 | -0.135 | -0.3271 | No | ||
| 86 | COL4A6 | Col4a6 | 9840 | -0.142 | -0.3310 | No | ||
| 87 | IGF1 | Igf1 | 9853 | -0.143 | -0.3301 | No | ||
| 88 | CAPN2 | Capn2 | 9901 | -0.147 | -0.3314 | No | ||
| 89 | IGF1R | Igf1r | 10217 | -0.170 | -0.3499 | No | ||
| 90 | ITGB6 | Itgb6 | 10249 | -0.172 | -0.3499 | No | ||
| 91 | FYN | Fyn | 10250 | -0.172 | -0.3479 | No | ||
| 92 | LAMC1 | Lamc1 | 10281 | -0.175 | -0.3478 | No | ||
| 93 | PAK2 | Pak2 | 10297 | -0.177 | -0.3467 | No | ||
| 94 | PPP1CB | Ppp1cb | 10473 | -0.190 | -0.3558 | No | ||
| 95 | GSK3B | Gsk3b | 10504 | -0.192 | -0.3555 | No | ||
| 96 | MAP2K1 | Map2k1 | 10546 | -0.196 | -0.3559 | No | ||
| 97 | PPP1CA | Ppp1ca | 10577 | -0.198 | -0.3555 | No | ||
| 98 | ROCK2 | Rock2 | 10619 | -0.200 | -0.3558 | No | ||
| 99 | MAPK1 | Mapk1 | 10708 | -0.207 | -0.3591 | No | ||
| 100 | CRK | Crk | 10736 | -0.210 | -0.3584 | No | ||
| 101 | ITGB5 | Itgb5 | 10790 | -0.214 | -0.3593 | No | ||
| 102 | MAPK10 | Mapk10 | 10797 | -0.215 | -0.3572 | No | ||
| 103 | PIK3CG | Pik3cg | 10818 | -0.216 | -0.3559 | No | ||
| 104 | PDPK1 | Pdpk1 | 10897 | -0.223 | -0.3584 | No | ||
| 105 | ITGA10 | Gm42957 | 10991 | -0.232 | -0.3617 | No | ||
| 106 | TLN1 | Tln1 | 11050 | -0.238 | -0.3626 | No | ||
| 107 | SHC4 | Shc4 | 11135 | -0.246 | -0.3652 | No | ||
| 108 | FLNB | Flnb | 11217 | -0.254 | -0.3675 | Yes | ||
| 109 | AKT3 | Akt3 | 11260 | -0.258 | -0.3672 | Yes | ||
| 110 | FN1 | Fn1 | 11306 | -0.262 | -0.3670 | Yes | ||
| 111 | JUN | Jun | 11324 | -0.264 | -0.3650 | Yes | ||
| 112 | EGFR | Egfr | 11360 | -0.268 | -0.3641 | Yes | ||
| 113 | DIAPH1 | Diaph1 | 11370 | -0.269 | -0.3615 | Yes | ||
| 114 | SPP1 | Spp1 | 11427 | -0.274 | -0.3619 | Yes | ||
| 115 | CHAD | Chad | 11439 | -0.275 | -0.3594 | Yes | ||
| 116 | MYL7 | Myl7 | 11466 | -0.277 | -0.3578 | Yes | ||
| 117 | ACTN3 | Actn3 | 11480 | -0.279 | -0.3553 | Yes | ||
| 118 | PDGFA | Pdgfa | 11487 | -0.280 | -0.3524 | Yes | ||
| 119 | PTEN | Pten | 11511 | -0.282 | -0.3506 | Yes | ||
| 120 | ITGB1 | Itgb1 | 11611 | -0.292 | -0.3536 | Yes | ||
| 121 | BIRC3 | Birc3 | 11686 | -0.300 | -0.3549 | Yes | ||
| 122 | MET | Met | 11695 | -0.300 | -0.3518 | Yes | ||
| 123 | ROCK1 | Rock1 | 11801 | -0.311 | -0.3550 | Yes | ||
| 124 | COL4A1 | Col4a1 | 11819 | -0.313 | -0.3524 | Yes | ||
| 125 | CDC42 | Cdc42 | 11896 | -0.324 | -0.3535 | Yes | ||
| 126 | MYLK3 | Mylk3 | 11899 | -0.324 | -0.3498 | Yes | ||
| 127 | TLN2 | Tln2 | 11940 | -0.328 | -0.3486 | Yes | ||
| 128 | BCL2 | Bcl2 | 12075 | -0.344 | -0.3532 | Yes | ||
| 129 | BRAF | Braf | 12139 | -0.348 | -0.3532 | Yes | ||
| 130 | RHOA | Rhoa | 12195 | -0.354 | -0.3526 | Yes | ||
| 131 | ITGA6 | Itga6 | 12220 | -0.357 | -0.3500 | Yes | ||
| 132 | HRAS | Hras | 12233 | -0.358 | -0.3465 | Yes | ||
| 133 | BAD | Bad | 12251 | -0.360 | -0.3434 | Yes | ||
| 134 | PARVA | Parva | 12373 | -0.374 | -0.3468 | Yes | ||
| 135 | RAC1 | Rac1 | 12380 | -0.375 | -0.3428 | Yes | ||
| 136 | SHC1 | Shc1 | 12435 | -0.383 | -0.3418 | Yes | ||
| 137 | LAMC2 | Lamc2 | 12467 | -0.387 | -0.3392 | Yes | ||
| 138 | RAP1B | Rap1b | 12581 | -0.401 | -0.3419 | Yes | ||
| 139 | ELK1 | Elk1 | 12586 | -0.401 | -0.3374 | Yes | ||
| 140 | PIK3R5 | Pik3r5 | 12592 | -0.402 | -0.3330 | Yes | ||
| 141 | COL4A2 | Col4a2 | 12687 | -0.414 | -0.3342 | Yes | ||
| 142 | PRKCA | Prkca | 12751 | -0.423 | -0.3333 | Yes | ||
| 143 | DOCK1 | Dock1 | 12778 | -0.426 | -0.3300 | Yes | ||
| 144 | PIK3R1 | Pik3r1 | 12996 | -0.447 | -0.3388 | Yes | ||
| 145 | MYL12B | Myl12b | 13167 | -0.472 | -0.3443 | Yes | ||
| 146 | ACTN2 | Actn2 | 13212 | -0.478 | -0.3415 | Yes | ||
| 147 | LAMA4 | Lama4 | 13247 | -0.484 | -0.3380 | Yes | ||
| 148 | MYLK | Mylk | 13274 | -0.491 | -0.3339 | Yes | ||
| 149 | RAC3 | Rac3 | 13294 | -0.495 | -0.3293 | Yes | ||
| 150 | VEGFD | Vegfd | 13416 | -0.504 | -0.3312 | Yes | ||
| 151 | VEGFA | Vegfa | 13510 | -0.521 | -0.3311 | Yes | ||
| 152 | PAK3 | Pak3 | 13553 | -0.527 | -0.3277 | Yes | ||
| 153 | COL11A1 | Col11a1 | 13641 | -0.540 | -0.3269 | Yes | ||
| 154 | CAV1 | Cav1 | 13720 | -0.556 | -0.3255 | Yes | ||
| 155 | PDGFRB | Pdgfrb | 13723 | -0.557 | -0.3190 | Yes | ||
| 156 | ARHGAP5 | Arhgap5 | 13738 | -0.561 | -0.3133 | Yes | ||
| 157 | COL6A1 | Col6a1 | 13839 | -0.580 | -0.3130 | Yes | ||
| 158 | LAMA5 | Lama5 | 13933 | -0.600 | -0.3119 | Yes | ||
| 159 | COL6A3 | Col6a3 | 13951 | -0.603 | -0.3059 | Yes | ||
| 160 | PRKCB | Prkcb | 13958 | -0.605 | -0.2991 | Yes | ||
| 161 | TNR | Tnr | 13971 | -0.608 | -0.2927 | Yes | ||
| 162 | ITGA8 | Itga8 | 14006 | -0.615 | -0.2877 | Yes | ||
| 163 | THBS3 | Thbs3 | 14080 | -0.632 | -0.2850 | Yes | ||
| 164 | RASGRF1 | Rasgrf1 | 14108 | -0.639 | -0.2792 | Yes | ||
| 165 | ITGB4 | Itgb4 | 14189 | -0.658 | -0.2766 | Yes | ||
| 166 | ITGAV | Itgav | 14285 | -0.689 | -0.2746 | Yes | ||
| 167 | TNC | Tnc | 14428 | -0.735 | -0.2752 | Yes | ||
| 168 | ITGA11 | Itga11 | 14522 | -0.769 | -0.2722 | Yes | ||
| 169 | COL1A2 | Col1a2 | 14592 | -0.795 | -0.2673 | Yes | ||
| 170 | ITGA1 | Itga1 | 14645 | -0.821 | -0.2610 | Yes | ||
| 171 | PDGFB | Pdgfb | 14669 | -0.829 | -0.2527 | Yes | ||
| 172 | COL6A2 | Col6a2 | 14684 | -0.839 | -0.2437 | Yes | ||
| 173 | CAV2 | Cav2 | 14721 | -0.854 | -0.2359 | Yes | ||
| 174 | FLT1 | Flt1 | 14807 | -0.893 | -0.2309 | Yes | ||
| 175 | COL5A2 | Col5a2 | 14890 | -0.944 | -0.2251 | Yes | ||
| 176 | CCND1 | Ccnd1 | 14921 | -0.960 | -0.2157 | Yes | ||
| 177 | COL3A1 | Col3a1 | 14977 | -0.993 | -0.2075 | Yes | ||
| 178 | ITGA7 | Itga7 | 15001 | -1.007 | -0.1971 | Yes | ||
| 179 | ITGB8 | Itgb8 | 15052 | -1.054 | -0.1879 | Yes | ||
| 180 | LAMA3 | Lama3 | 15079 | -1.075 | -0.1769 | Yes | ||
| 181 | ITGA4 | Itga4 | 15139 | -1.134 | -0.1673 | Yes | ||
| 182 | LAMB3 | Lamb3 | 15148 | -1.149 | -0.1542 | Yes | ||
| 183 | COL5A1 | Col5a1 | 15174 | -1.182 | -0.1419 | Yes | ||
| 184 | ITGA2 | Itga2 | 15178 | -1.187 | -0.1281 | Yes | ||
| 185 | THBS4 | Thbs4 | 15245 | -1.275 | -0.1173 | Yes | ||
| 186 | THBS2 | Thbs2 | 15251 | -1.286 | -0.1024 | Yes | ||
| 187 | PIK3CB | Pik3cb | 15260 | -1.300 | -0.0875 | Yes | ||
| 188 | COL5A3 | Col5a3 | 15370 | -1.534 | -0.0765 | Yes | ||
| 189 | FLT4 | Flt4 | 15375 | -1.541 | -0.0586 | Yes | ||
| 190 | COL4A4 | Col4a4 | 15411 | -1.647 | -0.0414 | Yes | ||
| 191 | COL1A1 | Col1a1 | 15434 | -1.720 | -0.0224 | Yes | ||
| 192 | PDGFD | Pdgfd | 15513 | -2.394 | 0.0008 | Yes |