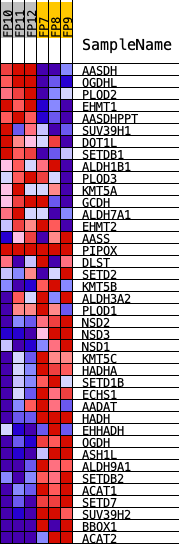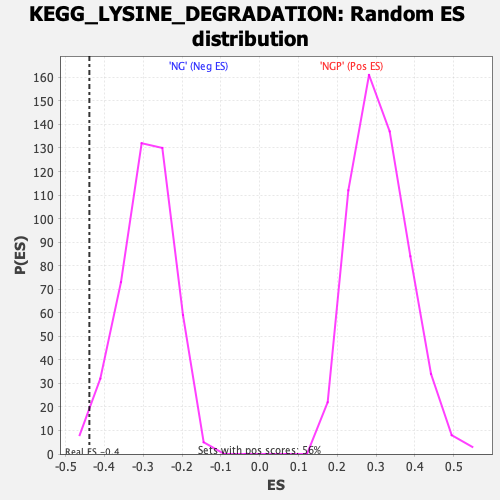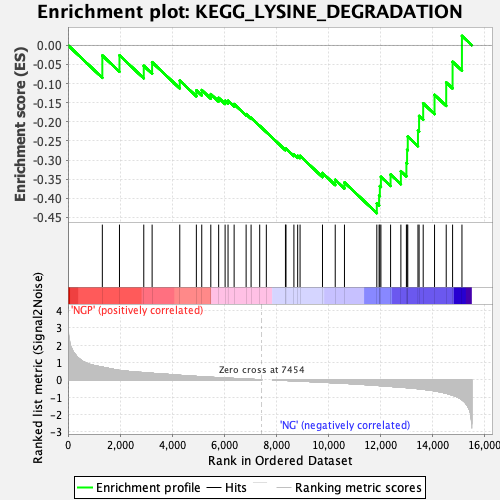
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_NG.phenotype.cls#NGP_versus_NG_repos |
| Phenotype | phenotype.cls#NGP_versus_NG_repos |
| Upregulated in class | NG |
| GeneSet | KEGG_LYSINE_DEGRADATION |
| Enrichment Score (ES) | -0.43886203 |
| Normalized Enrichment Score (NES) | -1.5033094 |
| Nominal p-value | 0.01594533 |
| FDR q-value | 0.14171186 |
| FWER p-Value | 0.833 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | AASDH | Aasdh | 1320 | 0.738 | -0.0267 | No | ||
| 2 | OGDHL | Ogdhl | 1979 | 0.542 | -0.0263 | No | ||
| 3 | PLOD2 | Plod2 | 2913 | 0.413 | -0.0538 | No | ||
| 4 | EHMT1 | Ehmt1 | 3234 | 0.380 | -0.0443 | No | ||
| 5 | AASDHPPT | Aasdhppt | 4297 | 0.259 | -0.0924 | No | ||
| 6 | SUV39H1 | Suv39h1 | 4935 | 0.196 | -0.1180 | No | ||
| 7 | DOT1L | Dot1l | 5142 | 0.178 | -0.1172 | No | ||
| 8 | SETDB1 | Setdb1 | 5490 | 0.148 | -0.1278 | No | ||
| 9 | ALDH1B1 | Aldh1b1 | 5792 | 0.122 | -0.1376 | No | ||
| 10 | PLOD3 | Plod3 | 6040 | 0.105 | -0.1453 | No | ||
| 11 | KMT5A | Kmt5a | 6153 | 0.094 | -0.1451 | No | ||
| 12 | GCDH | Gcdh | 6387 | 0.078 | -0.1539 | No | ||
| 13 | ALDH7A1 | Aldh7a1 | 6848 | 0.045 | -0.1801 | No | ||
| 14 | EHMT2 | Ehmt2 | 7039 | 0.031 | -0.1898 | No | ||
| 15 | AASS | Aass | 7370 | 0.007 | -0.2106 | No | ||
| 16 | PIPOX | Pipox | 7625 | 0.000 | -0.2270 | No | ||
| 17 | DLST | Dlst | 8360 | -0.035 | -0.2717 | No | ||
| 18 | SETD2 | Setd2 | 8381 | -0.036 | -0.2701 | No | ||
| 19 | KMT5B | Kmt5b | 8685 | -0.057 | -0.2851 | No | ||
| 20 | ALDH3A2 | Aldh3a2 | 8830 | -0.067 | -0.2891 | No | ||
| 21 | PLOD1 | Plod1 | 8922 | -0.073 | -0.2892 | No | ||
| 22 | NSD2 | Nsd2 | 9784 | -0.137 | -0.3339 | No | ||
| 23 | NSD3 | Nsd3 | 10274 | -0.175 | -0.3516 | No | ||
| 24 | NSD1 | Nsd1 | 10629 | -0.200 | -0.3586 | No | ||
| 25 | KMT5C | Kmt5c | 11873 | -0.320 | -0.4135 | Yes | ||
| 26 | HADHA | Hadha | 11961 | -0.331 | -0.3929 | Yes | ||
| 27 | SETD1B | Setd1b | 11990 | -0.334 | -0.3682 | Yes | ||
| 28 | ECHS1 | Echs1 | 12030 | -0.339 | -0.3439 | Yes | ||
| 29 | AADAT | Aadat | 12402 | -0.379 | -0.3378 | Yes | ||
| 30 | HADH | Hadh | 12798 | -0.430 | -0.3293 | Yes | ||
| 31 | EHHADH | Ehhadh | 13010 | -0.448 | -0.3074 | Yes | ||
| 32 | OGDH | Ogdh | 13037 | -0.452 | -0.2732 | Yes | ||
| 33 | ASH1L | Ash1l | 13063 | -0.455 | -0.2387 | Yes | ||
| 34 | ALDH9A1 | Aldh9a1 | 13455 | -0.514 | -0.2233 | Yes | ||
| 35 | SETDB2 | Setdb2 | 13498 | -0.520 | -0.1848 | Yes | ||
| 36 | ACAT1 | Acat1 | 13656 | -0.543 | -0.1519 | Yes | ||
| 37 | SETD7 | Setd7 | 14092 | -0.635 | -0.1297 | Yes | ||
| 38 | SUV39H2 | Suv39h2 | 14541 | -0.774 | -0.0972 | Yes | ||
| 39 | BBOX1 | Bbox1 | 14786 | -0.880 | -0.0432 | Yes | ||
| 40 | ACAT2 | Acat2 | 15146 | -1.147 | 0.0245 | Yes |