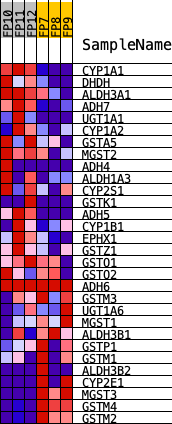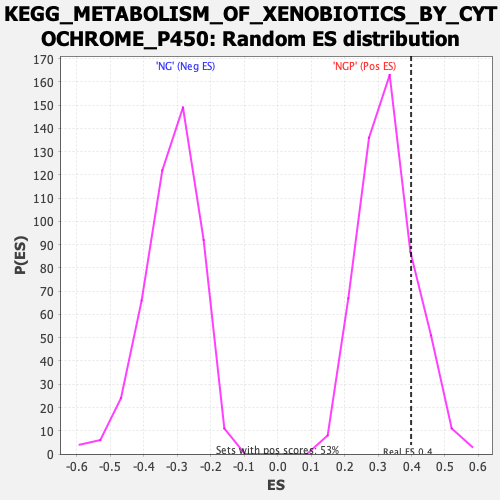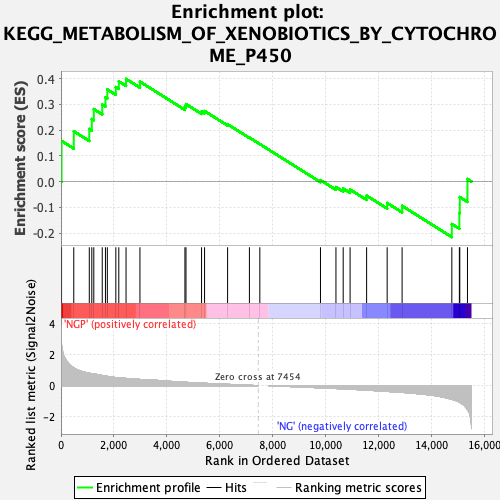
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_NG.phenotype.cls#NGP_versus_NG_repos |
| Phenotype | phenotype.cls#NGP_versus_NG_repos |
| Upregulated in class | NGP |
| GeneSet | KEGG_METABOLISM_OF_XENOBIOTICS_BY_CYTOCHROME_P450 |
| Enrichment Score (ES) | 0.3987882 |
| Normalized Enrichment Score (NES) | 1.2160983 |
| Nominal p-value | 0.19771864 |
| FDR q-value | 0.70198184 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CYP1A1 | Cyp1a1 | 21 | 2.759 | 0.1580 | Yes | ||
| 2 | DHDH | Dhdh | 483 | 1.163 | 0.1955 | Yes | ||
| 3 | ALDH3A1 | Aldh3a1 | 1070 | 0.808 | 0.2043 | Yes | ||
| 4 | ADH7 | Adh7 | 1165 | 0.776 | 0.2431 | Yes | ||
| 5 | UGT1A1 | Ugt1a1 | 1240 | 0.748 | 0.2815 | Yes | ||
| 6 | CYP1A2 | Cyp1a2 | 1560 | 0.659 | 0.2990 | Yes | ||
| 7 | GSTA5 | Gsta3 | 1681 | 0.625 | 0.3274 | Yes | ||
| 8 | MGST2 | Mgst2 | 1752 | 0.604 | 0.3578 | Yes | ||
| 9 | ADH4 | Adh4 | 2074 | 0.523 | 0.3673 | Yes | ||
| 10 | ALDH1A3 | Aldh1a3 | 2186 | 0.505 | 0.3893 | Yes | ||
| 11 | CYP2S1 | Cyp2s1 | 2460 | 0.469 | 0.3988 | Yes | ||
| 12 | GSTK1 | Gstk1 | 2986 | 0.401 | 0.3881 | No | ||
| 13 | ADH5 | Adh5 | 4687 | 0.218 | 0.2910 | No | ||
| 14 | CYP1B1 | Cyp1b1 | 4727 | 0.215 | 0.3009 | No | ||
| 15 | EPHX1 | Ephx1 | 5316 | 0.162 | 0.2723 | No | ||
| 16 | GSTZ1 | Gstz1 | 5430 | 0.154 | 0.2739 | No | ||
| 17 | GSTO1 | Gsto1 | 6301 | 0.084 | 0.2226 | No | ||
| 18 | GSTO2 | Gsto2 | 7126 | 0.025 | 0.1709 | No | ||
| 19 | ADH6 | Adh6b | 7520 | 0.000 | 0.1455 | No | ||
| 20 | GSTM3 | Gstm5 | 9814 | -0.141 | 0.0056 | No | ||
| 21 | UGT1A6 | Ugt1a6a | 10399 | -0.184 | -0.0214 | No | ||
| 22 | MGST1 | Mgst1 | 10673 | -0.204 | -0.0272 | No | ||
| 23 | ALDH3B1 | Aldh3b1 | 10934 | -0.226 | -0.0309 | No | ||
| 24 | GSTP1 | Gstp1 | 11559 | -0.286 | -0.0547 | No | ||
| 25 | GSTM1 | Gstm1 | 12336 | -0.371 | -0.0833 | No | ||
| 26 | ALDH3B2 | Aldh3b3 | 12901 | -0.438 | -0.0944 | No | ||
| 27 | CYP2E1 | Cyp2e1 | 14782 | -0.880 | -0.1649 | No | ||
| 28 | MGST3 | Mgst3 | 15067 | -1.063 | -0.1218 | No | ||
| 29 | GSTM4 | Gstm4 | 15083 | -1.081 | -0.0604 | No | ||
| 30 | GSTM2 | Gstm7 | 15373 | -1.537 | 0.0098 | No |