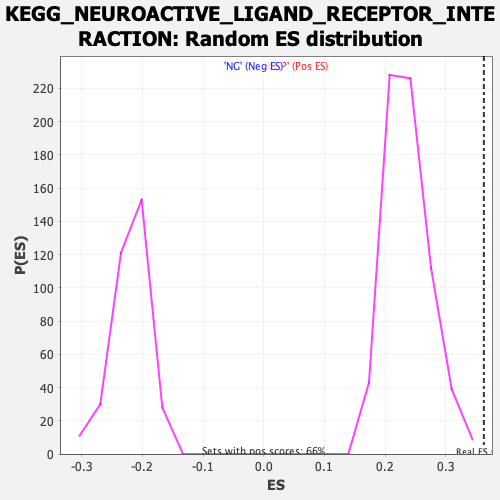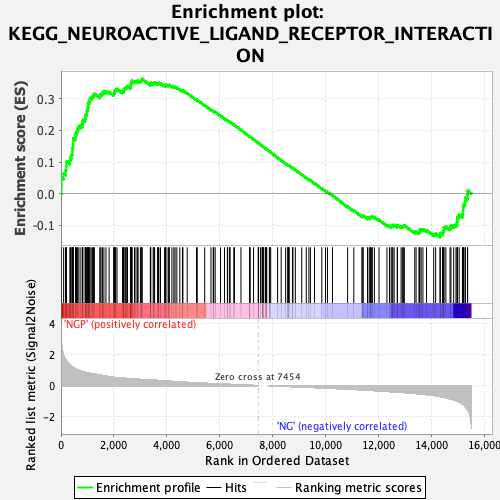
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_NG.phenotype.cls#NGP_versus_NG_repos |
| Phenotype | phenotype.cls#NGP_versus_NG_repos |
| Upregulated in class | NGP |
| GeneSet | KEGG_NEUROACTIVE_LIGAND_RECEPTOR_INTERACTION |
| Enrichment Score (ES) | 0.3627522 |
| Normalized Enrichment Score (NES) | 1.5343941 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.3589833 |
| FWER p-Value | 0.847 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | HTR1F | Htr1f | 16 | 2.890 | 0.0252 | Yes | ||
| 2 | OPRL1 | Oprl1 | 19 | 2.825 | 0.0507 | Yes | ||
| 3 | PRL | Prl | 100 | 1.955 | 0.0632 | Yes | ||
| 4 | GPR50 | Gpr50 | 174 | 1.724 | 0.0740 | Yes | ||
| 5 | P2RX7 | P2rx7 | 198 | 1.627 | 0.0873 | Yes | ||
| 6 | CHRNA10 | Chrna10 | 204 | 1.613 | 0.1016 | Yes | ||
| 7 | S1PR4 | S1pr4 | 337 | 1.348 | 0.1052 | Yes | ||
| 8 | DRD4 | Drd4 | 359 | 1.314 | 0.1158 | Yes | ||
| 9 | P2RY10 | P2ry10 | 410 | 1.242 | 0.1238 | Yes | ||
| 10 | CHRND | Chrnd | 430 | 1.228 | 0.1337 | Yes | ||
| 11 | GRID1 | Grid1 | 431 | 1.226 | 0.1448 | Yes | ||
| 12 | CHRNB2 | Chrnb2 | 454 | 1.188 | 0.1541 | Yes | ||
| 13 | HTR6 | Htr6 | 460 | 1.185 | 0.1646 | Yes | ||
| 14 | GABRB2 | Gabrb2 | 462 | 1.183 | 0.1752 | Yes | ||
| 15 | VIPR2 | Vipr2 | 548 | 1.088 | 0.1795 | Yes | ||
| 16 | CHRNA5 | Chrna5 | 552 | 1.081 | 0.1891 | Yes | ||
| 17 | F2RL3 | F2rl3 | 587 | 1.052 | 0.1965 | Yes | ||
| 18 | RXFP2 | Rxfp2 | 619 | 1.029 | 0.2038 | Yes | ||
| 19 | GNRHR | Gnrhr | 651 | 1.008 | 0.2109 | Yes | ||
| 20 | FPR1 | Fpr1 | 721 | 0.961 | 0.2151 | Yes | ||
| 21 | PTGER2 | Ptger2 | 800 | 0.912 | 0.2183 | Yes | ||
| 22 | SSTR5 | Sstr5 | 811 | 0.907 | 0.2259 | Yes | ||
| 23 | GRIN2B | Grin2b | 828 | 0.898 | 0.2330 | Yes | ||
| 24 | CHRNB4 | Chrnb4 | 909 | 0.866 | 0.2356 | Yes | ||
| 25 | GABRA6 | Gabra6 | 930 | 0.859 | 0.2421 | Yes | ||
| 26 | GRIK3 | Grik3 | 933 | 0.859 | 0.2497 | Yes | ||
| 27 | GABRQ | Gabrq | 976 | 0.839 | 0.2546 | Yes | ||
| 28 | GALR3 | Galr3 | 978 | 0.839 | 0.2622 | Yes | ||
| 29 | ADRA2B | Adra2b | 1005 | 0.825 | 0.2679 | Yes | ||
| 30 | NPFFR1 | Npffr1 | 1022 | 0.820 | 0.2743 | Yes | ||
| 31 | GPR156 | Gpr156 | 1032 | 0.820 | 0.2812 | Yes | ||
| 32 | GRM2 | Grm2 | 1034 | 0.819 | 0.2885 | Yes | ||
| 33 | FPR2 | Fpr2 | 1087 | 0.802 | 0.2924 | Yes | ||
| 34 | ADORA1 | Adora1 | 1090 | 0.801 | 0.2996 | Yes | ||
| 35 | S1PR5 | S1pr5 | 1141 | 0.784 | 0.3034 | Yes | ||
| 36 | P2RX1 | P2rx1 | 1189 | 0.770 | 0.3073 | Yes | ||
| 37 | GCGR | Gcgr | 1229 | 0.751 | 0.3116 | Yes | ||
| 38 | GPR83 | Gpr83 | 1263 | 0.741 | 0.3162 | Yes | ||
| 39 | OXTR | Oxtr | 1472 | 0.683 | 0.3088 | Yes | ||
| 40 | LPAR4 | Lpar4 | 1483 | 0.680 | 0.3143 | Yes | ||
| 41 | GRM5 | Grm5 | 1553 | 0.661 | 0.3158 | Yes | ||
| 42 | GRIN3A | Grin3a | 1563 | 0.658 | 0.3212 | Yes | ||
| 43 | TRPV1 | Trpv1 | 1623 | 0.641 | 0.3231 | Yes | ||
| 44 | GRIN3B | Grin3b | 1701 | 0.620 | 0.3237 | Yes | ||
| 45 | APLNR | Aplnr | 1819 | 0.582 | 0.3213 | Yes | ||
| 46 | ADCYAP1R1 | Adcyap1r1 | 1991 | 0.539 | 0.3151 | Yes | ||
| 47 | PRLR | Prlr | 2014 | 0.536 | 0.3185 | Yes | ||
| 48 | HRH1 | Hrh1 | 2025 | 0.533 | 0.3227 | Yes | ||
| 49 | LEPR | Lepr | 2041 | 0.531 | 0.3265 | Yes | ||
| 50 | C3AR1 | C3ar1 | 2066 | 0.525 | 0.3297 | Yes | ||
| 51 | NTSR1 | Ntsr1 | 2117 | 0.517 | 0.3311 | Yes | ||
| 52 | LPAR2 | Lpar2 | 2327 | 0.486 | 0.3219 | Yes | ||
| 53 | MTNR1A | Mtnr1a | 2367 | 0.478 | 0.3237 | Yes | ||
| 54 | F2 | F2 | 2369 | 0.478 | 0.3280 | Yes | ||
| 55 | PRLHR | Prlhr | 2380 | 0.478 | 0.3317 | Yes | ||
| 56 | CHRNB3 | Chrnb3 | 2430 | 0.473 | 0.3327 | Yes | ||
| 57 | CNR1 | Cnr1 | 2455 | 0.470 | 0.3354 | Yes | ||
| 58 | HTR2B | Htr2b | 2490 | 0.463 | 0.3374 | Yes | ||
| 59 | PTH1R | Pth1r | 2515 | 0.459 | 0.3400 | Yes | ||
| 60 | AGTR1 | Agtr1a | 2633 | 0.440 | 0.3364 | Yes | ||
| 61 | MC1R | Mc1r | 2637 | 0.440 | 0.3402 | Yes | ||
| 62 | GRM8 | Grm8 | 2638 | 0.440 | 0.3442 | Yes | ||
| 63 | ADRA1A | Adra1a | 2644 | 0.440 | 0.3478 | Yes | ||
| 64 | GLP2R | Glp2r | 2671 | 0.440 | 0.3501 | Yes | ||
| 65 | TACR1 | Tacr1 | 2674 | 0.440 | 0.3540 | Yes | ||
| 66 | ADORA2A | Adora2a | 2688 | 0.440 | 0.3571 | Yes | ||
| 67 | CHRNA3 | Chrna3 | 2781 | 0.428 | 0.3550 | Yes | ||
| 68 | GRIK4 | Grik4 | 2837 | 0.424 | 0.3553 | Yes | ||
| 69 | ADRA2C | Adra2c | 2902 | 0.415 | 0.3548 | Yes | ||
| 70 | CHRNE | Chrne | 2912 | 0.414 | 0.3580 | Yes | ||
| 71 | GABRD | Gabrd | 3004 | 0.401 | 0.3557 | Yes | ||
| 72 | GALR1 | Galr1 | 3034 | 0.401 | 0.3574 | Yes | ||
| 73 | OPRK1 | Oprk1 | 3061 | 0.401 | 0.3594 | Yes | ||
| 74 | F2RL2 | F2rl2 | 3066 | 0.401 | 0.3628 | Yes | ||
| 75 | MC4R | Mc4r | 3381 | 0.356 | 0.3455 | No | ||
| 76 | MC3R | Mc3r | 3382 | 0.356 | 0.3487 | No | ||
| 77 | AVPR1B | Avpr1b | 3396 | 0.356 | 0.3511 | No | ||
| 78 | NPY2R | Npy2r | 3487 | 0.356 | 0.3484 | No | ||
| 79 | GHRHR | Ghrhr | 3515 | 0.356 | 0.3499 | No | ||
| 80 | GABRA1 | Gabra1 | 3541 | 0.353 | 0.3515 | No | ||
| 81 | GABRR2 | Gabrr2 | 3652 | 0.339 | 0.3474 | No | ||
| 82 | ADRA2A | Adra2a | 3688 | 0.334 | 0.3481 | No | ||
| 83 | NMUR1 | Nmur1 | 3700 | 0.332 | 0.3504 | No | ||
| 84 | GABRE | Gabre | 3763 | 0.324 | 0.3493 | No | ||
| 85 | GABBR2 | Gabbr2 | 3918 | 0.303 | 0.3420 | No | ||
| 86 | GIPR | Gipr | 3953 | 0.300 | 0.3425 | No | ||
| 87 | HRH3 | Hrh3 | 3972 | 0.297 | 0.3440 | No | ||
| 88 | KISS1R | Kiss1r | 4051 | 0.287 | 0.3415 | No | ||
| 89 | TSHR | Tshr | 4075 | 0.284 | 0.3426 | No | ||
| 90 | GRIN2D | Grin2d | 4098 | 0.282 | 0.3437 | No | ||
| 91 | P2RX2 | P2rx2 | 4195 | 0.271 | 0.3399 | No | ||
| 92 | LTB4R2 | Ltb4r2 | 4269 | 0.263 | 0.3375 | No | ||
| 93 | GLRA1 | Glra1 | 4314 | 0.257 | 0.3370 | No | ||
| 94 | GRIA3 | Gria3 | 4381 | 0.251 | 0.3349 | No | ||
| 95 | THRB | Thrb | 4495 | 0.237 | 0.3297 | No | ||
| 96 | GRPR | Grpr | 4594 | 0.228 | 0.3254 | No | ||
| 97 | P2RY1 | P2ry1 | 4611 | 0.226 | 0.3264 | No | ||
| 98 | EDNRA | Ednra | 4774 | 0.210 | 0.3177 | No | ||
| 99 | TRHR | Trhr | 5126 | 0.179 | 0.2964 | No | ||
| 100 | HCRTR1 | Hcrtr1 | 5153 | 0.177 | 0.2963 | No | ||
| 101 | GABRA3 | Gabra3 | 5435 | 0.153 | 0.2793 | No | ||
| 102 | OPRD1 | Oprd1 | 5672 | 0.132 | 0.2651 | No | ||
| 103 | CHRM1 | Chrm1 | 5755 | 0.125 | 0.2609 | No | ||
| 104 | PLG | Plg | 5776 | 0.123 | 0.2607 | No | ||
| 105 | VIPR1 | Vipr1 | 5831 | 0.119 | 0.2583 | No | ||
| 106 | GLP1R | Glp1r | 6034 | 0.105 | 0.2460 | No | ||
| 107 | HTR7 | Htr7 | 6183 | 0.091 | 0.2372 | No | ||
| 108 | CHRM3 | Chrm3 | 6292 | 0.084 | 0.2309 | No | ||
| 109 | GRIN2A | Grin2a | 6303 | 0.084 | 0.2310 | No | ||
| 110 | SSTR3 | Sstr3 | 6377 | 0.079 | 0.2270 | No | ||
| 111 | AVPR2 | Avpr2 | 6395 | 0.078 | 0.2266 | No | ||
| 112 | CNR2 | Cnr2 | 6536 | 0.068 | 0.2180 | No | ||
| 113 | PARD3 | Pard3 | 6561 | 0.066 | 0.2171 | No | ||
| 114 | PTGIR | Ptgir | 6807 | 0.048 | 0.2015 | No | ||
| 115 | HTR1B | Htr1b | 7140 | 0.024 | 0.1800 | No | ||
| 116 | EDNRB | Ednrb | 7141 | 0.024 | 0.1802 | No | ||
| 117 | PTGER4 | Ptger4 | 7147 | 0.023 | 0.1801 | No | ||
| 118 | GALR2 | Galr2 | 7283 | 0.013 | 0.1714 | No | ||
| 119 | HTR5A | Htr5a | 7459 | 0.000 | 0.1600 | No | ||
| 120 | GRM1 | Grm1 | 7468 | 0.000 | 0.1595 | No | ||
| 121 | GRM3 | Grm3 | 7469 | 0.000 | 0.1595 | No | ||
| 122 | GRM6 | Grm6 | 7470 | 0.000 | 0.1595 | No | ||
| 123 | MCHR1 | Mchr1 | 7543 | 0.000 | 0.1548 | No | ||
| 124 | GABRG2 | Gabrg2 | 7609 | 0.000 | 0.1505 | No | ||
| 125 | GABRG1 | Gabrg1 | 7610 | 0.000 | 0.1505 | No | ||
| 126 | CHRNA9 | Chrna9 | 7622 | 0.000 | 0.1498 | No | ||
| 127 | CTSG | Ctsg | 7631 | 0.000 | 0.1493 | No | ||
| 128 | CCKBR | Cckbr | 7648 | 0.000 | 0.1482 | No | ||
| 129 | C5AR1 | C5ar1 | 7651 | 0.000 | 0.1481 | No | ||
| 130 | P2RY4 | P2ry4 | 7725 | 0.000 | 0.1433 | No | ||
| 131 | CALCR | Calcr | 7746 | 0.000 | 0.1420 | No | ||
| 132 | GHSR | Ghsr | 7756 | 0.000 | 0.1414 | No | ||
| 133 | HCRTR2 | Hcrtr2 | 7764 | 0.000 | 0.1410 | No | ||
| 134 | P2RY13 | P2ry13 | 7782 | 0.000 | 0.1399 | No | ||
| 135 | GRIK5 | Grik5 | 7882 | -0.002 | 0.1334 | No | ||
| 136 | CHRNA2 | Chrna2 | 7892 | -0.003 | 0.1329 | No | ||
| 137 | GRM4 | Grm4 | 7931 | -0.005 | 0.1304 | No | ||
| 138 | BDKRB2 | Bdkrb2 | 8193 | -0.024 | 0.1136 | No | ||
| 139 | GRIA2 | Gria2 | 8328 | -0.034 | 0.1051 | No | ||
| 140 | DRD1 | Drd1 | 8497 | -0.044 | 0.0946 | No | ||
| 141 | GPR35 | Gpr35 | 8583 | -0.050 | 0.0895 | No | ||
| 142 | ADORA2B | Adora2b | 8584 | -0.050 | 0.0899 | No | ||
| 143 | PTAFR | Ptafr | 8596 | -0.051 | 0.0897 | No | ||
| 144 | GRIN2C | Grin2c | 8617 | -0.052 | 0.0888 | No | ||
| 145 | TSHB | Tshb | 8629 | -0.053 | 0.0886 | No | ||
| 146 | SSTR4 | Sstr4 | 8753 | -0.062 | 0.0811 | No | ||
| 147 | CHRNA4 | Chrna4 | 8772 | -0.063 | 0.0805 | No | ||
| 148 | P2RX4 | P2rx4 | 8863 | -0.069 | 0.0753 | No | ||
| 149 | GABRA4 | Gabra4 | 9105 | -0.087 | 0.0603 | No | ||
| 150 | LPAR3 | Lpar3 | 9276 | -0.098 | 0.0501 | No | ||
| 151 | MAS1 | Mas1 | 9376 | -0.105 | 0.0446 | No | ||
| 152 | SSTR1 | Sstr1 | 9433 | -0.110 | 0.0419 | No | ||
| 153 | HRH4 | Hrh4 | 9588 | -0.122 | 0.0330 | No | ||
| 154 | BDKRB1 | Bdkrb1 | 9865 | -0.145 | 0.0163 | No | ||
| 155 | GRIA1 | Gria1 | 10005 | -0.154 | 0.0086 | No | ||
| 156 | S1PR2 | S1pr2 | 10078 | -0.159 | 0.0053 | No | ||
| 157 | GABRB3 | Gabrb3 | 10267 | -0.174 | -0.0054 | No | ||
| 158 | CHRM5 | Chrm5 | 10841 | -0.218 | -0.0408 | No | ||
| 159 | LPAR1 | Lpar1 | 11074 | -0.240 | -0.0538 | No | ||
| 160 | LPAR6 | Lpar6 | 11378 | -0.270 | -0.0711 | No | ||
| 161 | GABRR1 | Gabrr1 | 11417 | -0.273 | -0.0711 | No | ||
| 162 | TBXA2R | Tbxa2r | 11426 | -0.274 | -0.0692 | No | ||
| 163 | NPY5R | Npy5r | 11597 | -0.290 | -0.0777 | No | ||
| 164 | AGTR2 | Agtr2 | 11603 | -0.291 | -0.0753 | No | ||
| 165 | F2R | F2r | 11677 | -0.299 | -0.0774 | No | ||
| 166 | HTR4 | Htr4 | 11702 | -0.301 | -0.0762 | No | ||
| 167 | PTGFR | Ptgfr | 11709 | -0.302 | -0.0739 | No | ||
| 168 | CHRNA1 | Chrna1 | 11738 | -0.305 | -0.0730 | No | ||
| 169 | NMBR | Nmbr | 11775 | -0.308 | -0.0725 | No | ||
| 170 | SCTR | Sctr | 11852 | -0.318 | -0.0746 | No | ||
| 171 | CCKAR | Cckar | 12028 | -0.339 | -0.0829 | No | ||
| 172 | HTR2A | Htr2a | 12327 | -0.370 | -0.0991 | No | ||
| 173 | P2RX3 | P2rx3 | 12424 | -0.382 | -0.1019 | No | ||
| 174 | CHRM4 | Chrm4 | 12488 | -0.389 | -0.1024 | No | ||
| 175 | GABBR1 | Gabbr1 | 12524 | -0.394 | -0.1012 | No | ||
| 176 | S1PR1 | S1pr1 | 12551 | -0.397 | -0.0993 | No | ||
| 177 | GRIA4 | Gria4 | 12601 | -0.402 | -0.0988 | No | ||
| 178 | CHRNB1 | Chrnb1 | 12717 | -0.418 | -0.1025 | No | ||
| 179 | RXFP1 | Rxfp1 | 12726 | -0.420 | -0.0992 | No | ||
| 180 | AVPR1A | Avpr1a | 12863 | -0.438 | -0.1042 | No | ||
| 181 | SSTR2 | Sstr2 | 12910 | -0.438 | -0.1032 | No | ||
| 182 | GLRA3 | Glra3 | 12950 | -0.443 | -0.1017 | No | ||
| 183 | PTH2R | Pth2r | 12986 | -0.446 | -0.1000 | No | ||
| 184 | NTSR2 | Ntsr2 | 13379 | -0.501 | -0.1210 | No | ||
| 185 | NPY1R | Npy1r | 13430 | -0.507 | -0.1197 | No | ||
| 186 | TACR2 | Tacr2 | 13539 | -0.525 | -0.1220 | No | ||
| 187 | NR3C1 | Nr3c1 | 13565 | -0.527 | -0.1188 | No | ||
| 188 | ADRB1 | Adrb1 | 13566 | -0.528 | -0.1140 | No | ||
| 189 | GLRB | Glrb | 13621 | -0.536 | -0.1127 | No | ||
| 190 | GHR | Ghr | 13688 | -0.550 | -0.1120 | No | ||
| 191 | PTGER1 | Ptger1 | 13825 | -0.578 | -0.1157 | No | ||
| 192 | GRM7 | Grm7 | 14096 | -0.636 | -0.1275 | No | ||
| 193 | THRA | Thra | 14170 | -0.654 | -0.1264 | No | ||
| 194 | TACR3 | Tacr3 | 14338 | -0.704 | -0.1309 | No | ||
| 195 | ADRA1D | Adra1d | 14341 | -0.705 | -0.1246 | No | ||
| 196 | P2RY6 | P2ry6 | 14431 | -0.737 | -0.1238 | No | ||
| 197 | TSPO | Tspo | 14451 | -0.745 | -0.1182 | No | ||
| 198 | ADRB2 | Adrb2 | 14481 | -0.754 | -0.1133 | No | ||
| 199 | S1PR3 | S1pr3 | 14482 | -0.755 | -0.1064 | No | ||
| 200 | GRID2 | Grid2 | 14557 | -0.780 | -0.1042 | No | ||
| 201 | MC5R | Mc5r | 14715 | -0.853 | -0.1067 | No | ||
| 202 | P2RX6 | P2rx6 | 14748 | -0.865 | -0.1010 | No | ||
| 203 | PTGER3 | Ptger3 | 14852 | -0.919 | -0.0994 | No | ||
| 204 | HRH2 | Hrh2 | 14939 | -0.969 | -0.0962 | No | ||
| 205 | P2RY2 | P2ry2 | 14979 | -0.994 | -0.0897 | No | ||
| 206 | GRIN1 | Grin1 | 14989 | -0.997 | -0.0813 | No | ||
| 207 | CALCRL | Calcrl | 14996 | -1.002 | -0.0726 | No | ||
| 208 | CYSLTR1 | Cysltr1 | 15059 | -1.058 | -0.0670 | No | ||
| 209 | CHRNG | Chrng | 15182 | -1.191 | -0.0642 | No | ||
| 210 | GRIK1 | Grik1 | 15203 | -1.217 | -0.0544 | No | ||
| 211 | ADRA1B | Adra1b | 15210 | -1.225 | -0.0437 | No | ||
| 212 | GABRB1 | Gabrb1 | 15227 | -1.245 | -0.0335 | No | ||
| 213 | GRIK2 | Grik2 | 15282 | -1.346 | -0.0248 | No | ||
| 214 | GABRA2 | Gabra2 | 15292 | -1.364 | -0.0130 | No | ||
| 215 | P2RY14 | P2ry14 | 15374 | -1.538 | -0.0043 | No | ||
| 216 | F2RL1 | F2rl1 | 15379 | -1.559 | 0.0095 | No |