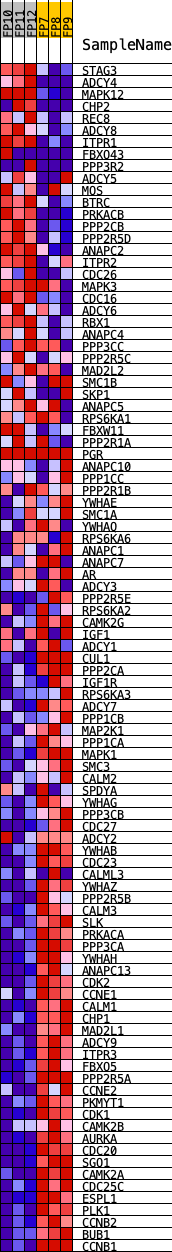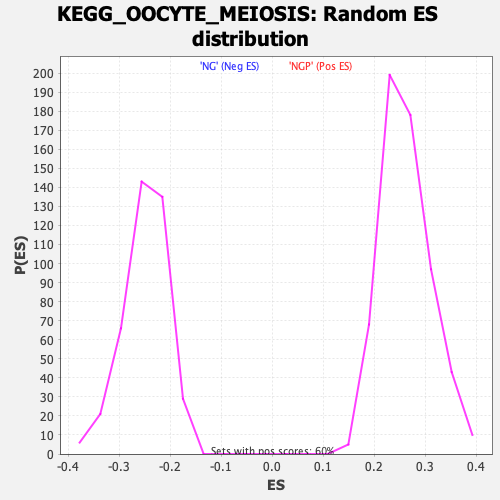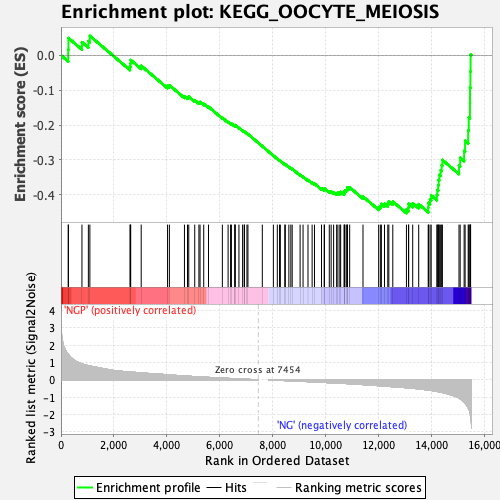
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_NG.phenotype.cls#NGP_versus_NG_repos |
| Phenotype | phenotype.cls#NGP_versus_NG_repos |
| Upregulated in class | NG |
| GeneSet | KEGG_OOCYTE_MEIOSIS |
| Enrichment Score (ES) | -0.45283353 |
| Normalized Enrichment Score (NES) | -1.8222893 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.025496654 |
| FWER p-Value | 0.077 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | STAG3 | Stag3 | 273 | 1.469 | 0.0162 | No | ||
| 2 | ADCY4 | Adcy4 | 281 | 1.467 | 0.0496 | No | ||
| 3 | MAPK12 | Mapk12 | 792 | 0.920 | 0.0377 | No | ||
| 4 | CHP2 | Chp2 | 1035 | 0.819 | 0.0409 | No | ||
| 5 | REC8 | Rec8 | 1092 | 0.801 | 0.0558 | No | ||
| 6 | ADCY8 | Adcy8 | 2609 | 0.445 | -0.0322 | No | ||
| 7 | ITPR1 | Itpr1 | 2629 | 0.441 | -0.0233 | No | ||
| 8 | FBXO43 | Fbxo43 | 2636 | 0.440 | -0.0135 | No | ||
| 9 | PPP3R2 | Ppp3r2 | 3033 | 0.401 | -0.0300 | No | ||
| 10 | ADCY5 | Adcy5 | 4030 | 0.290 | -0.0879 | No | ||
| 11 | MOS | Mos | 4099 | 0.282 | -0.0858 | No | ||
| 12 | BTRC | Btrc | 4669 | 0.220 | -0.1176 | No | ||
| 13 | PRKACB | Prkacb | 4785 | 0.209 | -0.1202 | No | ||
| 14 | PPP2CB | Ppp2cb | 4832 | 0.205 | -0.1185 | No | ||
| 15 | PPP2R5D | Ppp2r5d | 5058 | 0.185 | -0.1288 | No | ||
| 16 | ANAPC2 | Anapc2 | 5216 | 0.171 | -0.1350 | No | ||
| 17 | ITPR2 | Itpr2 | 5259 | 0.167 | -0.1339 | No | ||
| 18 | CDC26 | Cdc26 | 5398 | 0.156 | -0.1392 | No | ||
| 19 | MAPK3 | Mapk3 | 5579 | 0.140 | -0.1477 | No | ||
| 20 | CDC16 | Cdc16 | 6103 | 0.099 | -0.1793 | No | ||
| 21 | ADCY6 | Adcy6 | 6320 | 0.083 | -0.1914 | No | ||
| 22 | RBX1 | Rbx1 | 6414 | 0.077 | -0.1956 | No | ||
| 23 | ANAPC4 | Anapc4 | 6445 | 0.075 | -0.1959 | No | ||
| 24 | PPP3CC | Ppp3cc | 6577 | 0.065 | -0.2028 | No | ||
| 25 | PPP2R5C | Ppp2r5c | 6579 | 0.065 | -0.2014 | No | ||
| 26 | MAD2L2 | Mad2l2 | 6585 | 0.065 | -0.2002 | No | ||
| 27 | SMC1B | Smc1b | 6730 | 0.054 | -0.2083 | No | ||
| 28 | SKP1 | Skp1a | 6868 | 0.043 | -0.2162 | No | ||
| 29 | ANAPC5 | Anapc5 | 6928 | 0.039 | -0.2191 | No | ||
| 30 | RPS6KA1 | Rps6ka1 | 6940 | 0.038 | -0.2189 | No | ||
| 31 | FBXW11 | Fbxw11 | 7033 | 0.032 | -0.2242 | No | ||
| 32 | PPP2R1A | Ppp2r1a | 7070 | 0.029 | -0.2259 | No | ||
| 33 | PGR | Pgr | 7614 | 0.000 | -0.2610 | No | ||
| 34 | ANAPC10 | Anapc10 | 8032 | -0.012 | -0.2878 | No | ||
| 35 | PPP1CC | Ppp1cc | 8181 | -0.023 | -0.2969 | No | ||
| 36 | PPP2R1B | Ppp2r1b | 8265 | -0.028 | -0.3016 | No | ||
| 37 | YWHAE | Ywhae | 8296 | -0.031 | -0.3028 | No | ||
| 38 | SMC1A | Smc1a | 8464 | -0.041 | -0.3127 | No | ||
| 39 | YWHAQ | Ywhaq | 8484 | -0.043 | -0.3129 | No | ||
| 40 | RPS6KA6 | Rps6ka6 | 8618 | -0.052 | -0.3204 | No | ||
| 41 | ANAPC1 | Anapc1 | 8693 | -0.058 | -0.3238 | No | ||
| 42 | ANAPC7 | Anapc7 | 8744 | -0.061 | -0.3257 | No | ||
| 43 | AR | Ar | 9043 | -0.082 | -0.3431 | No | ||
| 44 | ADCY3 | Adcy3 | 9155 | -0.091 | -0.3482 | No | ||
| 45 | PPP2R5E | Ppp2r5e | 9342 | -0.103 | -0.3579 | No | ||
| 46 | RPS6KA2 | Rps6ka2 | 9498 | -0.114 | -0.3653 | No | ||
| 47 | CAMK2G | Camk2g | 9587 | -0.121 | -0.3682 | No | ||
| 48 | IGF1 | Igf1 | 9853 | -0.143 | -0.3820 | No | ||
| 49 | ADCY1 | Adcy1 | 9955 | -0.151 | -0.3851 | No | ||
| 50 | CUL1 | Cul1 | 9964 | -0.152 | -0.3821 | No | ||
| 51 | PPP2CA | Ppp2ca | 10147 | -0.165 | -0.3901 | No | ||
| 52 | IGF1R | Igf1r | 10217 | -0.170 | -0.3907 | No | ||
| 53 | RPS6KA3 | Rps6ka3 | 10311 | -0.178 | -0.3926 | No | ||
| 54 | ADCY7 | Adcy7 | 10425 | -0.186 | -0.3957 | No | ||
| 55 | PPP1CB | Ppp1cb | 10473 | -0.190 | -0.3943 | No | ||
| 56 | MAP2K1 | Map2k1 | 10546 | -0.196 | -0.3945 | No | ||
| 57 | PPP1CA | Ppp1ca | 10577 | -0.198 | -0.3919 | No | ||
| 58 | MAPK1 | Mapk1 | 10708 | -0.207 | -0.3955 | No | ||
| 59 | SMC3 | Smc3 | 10718 | -0.208 | -0.3913 | No | ||
| 60 | CALM2 | Calm2 | 10735 | -0.209 | -0.3875 | No | ||
| 61 | SPDYA | Spdya | 10791 | -0.214 | -0.3861 | No | ||
| 62 | YWHAG | Ywhag | 10830 | -0.218 | -0.3836 | No | ||
| 63 | PPP3CB | Ppp3cb | 10834 | -0.218 | -0.3787 | No | ||
| 64 | CDC27 | Cdc27 | 10918 | -0.225 | -0.3789 | No | ||
| 65 | ADCY2 | Adcy2 | 11422 | -0.274 | -0.4052 | No | ||
| 66 | YWHAB | Ywhab | 12014 | -0.337 | -0.4358 | No | ||
| 67 | CDC23 | Cdc23 | 12078 | -0.344 | -0.4319 | No | ||
| 68 | CALML3 | Calml3 | 12117 | -0.346 | -0.4264 | No | ||
| 69 | YWHAZ | Ywhaz | 12235 | -0.358 | -0.4257 | No | ||
| 70 | PPP2R5B | Ppp2r5b | 12360 | -0.373 | -0.4252 | No | ||
| 71 | CALM3 | Calm3 | 12399 | -0.378 | -0.4189 | No | ||
| 72 | SLK | Slk | 12549 | -0.396 | -0.4195 | No | ||
| 73 | PRKACA | Prkaca | 13065 | -0.456 | -0.4423 | Yes | ||
| 74 | PPP3CA | Ppp3ca | 13145 | -0.468 | -0.4367 | Yes | ||
| 75 | YWHAH | Ywhah | 13149 | -0.468 | -0.4261 | Yes | ||
| 76 | ANAPC13 | Anapc13 | 13302 | -0.496 | -0.4245 | Yes | ||
| 77 | CDK2 | Cdk2 | 13529 | -0.524 | -0.4270 | Yes | ||
| 78 | CCNE1 | Ccne1 | 13892 | -0.591 | -0.4369 | Yes | ||
| 79 | CALM1 | Calm1 | 13895 | -0.591 | -0.4234 | Yes | ||
| 80 | CHP1 | Chp1 | 13957 | -0.605 | -0.4134 | Yes | ||
| 81 | MAD2L1 | Mad2l1 | 14001 | -0.614 | -0.4020 | Yes | ||
| 82 | ADCY9 | Adcy9 | 14216 | -0.667 | -0.4005 | Yes | ||
| 83 | ITPR3 | Itpr3 | 14234 | -0.673 | -0.3861 | Yes | ||
| 84 | FBXO5 | Fbxo5 | 14269 | -0.685 | -0.3725 | Yes | ||
| 85 | PPP2R5A | Ppp2r5a | 14284 | -0.689 | -0.3575 | Yes | ||
| 86 | CCNE2 | Ccne2 | 14309 | -0.696 | -0.3430 | Yes | ||
| 87 | PKMYT1 | Pkmyt1 | 14359 | -0.714 | -0.3297 | Yes | ||
| 88 | CDK1 | Cdk1 | 14397 | -0.727 | -0.3153 | Yes | ||
| 89 | CAMK2B | Camk2b | 14425 | -0.735 | -0.3001 | Yes | ||
| 90 | AURKA | Aurka | 15053 | -1.055 | -0.3165 | Yes | ||
| 91 | CDC20 | Cdc20 | 15097 | -1.097 | -0.2940 | Yes | ||
| 92 | SGO1 | Sgo1 | 15247 | -1.280 | -0.2741 | Yes | ||
| 93 | CAMK2A | Camk2a | 15284 | -1.351 | -0.2453 | Yes | ||
| 94 | CDC25C | Cdc25c | 15401 | -1.621 | -0.2154 | Yes | ||
| 95 | ESPL1 | Espl1 | 15422 | -1.681 | -0.1779 | Yes | ||
| 96 | PLK1 | Plk1 | 15467 | -1.900 | -0.1370 | Yes | ||
| 97 | CCNB2 | Ccnb2 | 15470 | -1.905 | -0.0932 | Yes | ||
| 98 | BUB1 | Bub1 | 15485 | -2.064 | -0.0464 | Yes | ||
| 99 | CCNB1 | Ccnb1 | 15492 | -2.124 | 0.0021 | Yes |