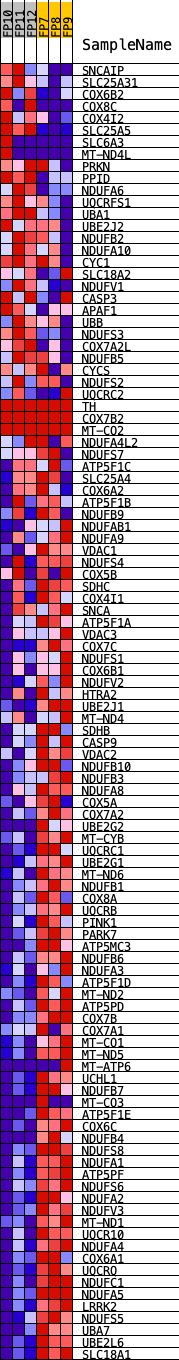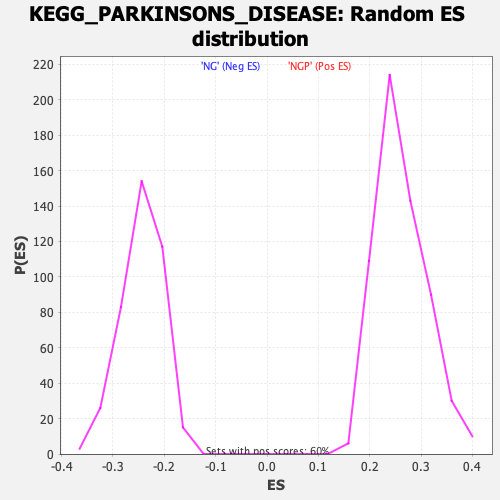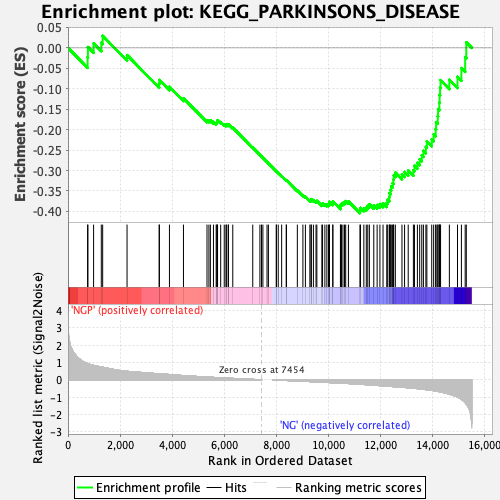
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_NG.phenotype.cls#NGP_versus_NG_repos |
| Phenotype | phenotype.cls#NGP_versus_NG_repos |
| Upregulated in class | NG |
| GeneSet | KEGG_PARKINSONS_DISEASE |
| Enrichment Score (ES) | -0.40436158 |
| Normalized Enrichment Score (NES) | -1.6546981 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0749144 |
| FWER p-Value | 0.377 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SNCAIP | Sncaip | 753 | 0.941 | -0.0229 | No | ||
| 2 | SLC25A31 | Slc25a31 | 766 | 0.933 | 0.0021 | No | ||
| 3 | COX6B2 | Cox6b2 | 984 | 0.837 | 0.0112 | No | ||
| 4 | COX8C | Cox8b | 1280 | 0.741 | 0.0125 | No | ||
| 5 | COX4I2 | Cox4i2 | 1333 | 0.734 | 0.0294 | No | ||
| 6 | SLC25A5 | Slc25a5 | 2270 | 0.493 | -0.0177 | No | ||
| 7 | SLC6A3 | Slc6a3 | 3504 | 0.356 | -0.0878 | No | ||
| 8 | MT-ND4L | mt-Nd4l | 3514 | 0.356 | -0.0785 | No | ||
| 9 | PRKN | Prkn | 3900 | 0.306 | -0.0951 | No | ||
| 10 | PPID | Ppid | 4439 | 0.244 | -0.1232 | No | ||
| 11 | NDUFA6 | Ndufa6 | 5344 | 0.161 | -0.1774 | No | ||
| 12 | UQCRFS1 | Uqcrfs1 | 5418 | 0.155 | -0.1779 | No | ||
| 13 | UBA1 | Uba1 | 5474 | 0.149 | -0.1773 | No | ||
| 14 | UBE2J2 | Ube2j2 | 5592 | 0.139 | -0.1811 | No | ||
| 15 | NDUFB2 | Ndufb2 | 5696 | 0.131 | -0.1841 | No | ||
| 16 | NDUFA10 | Ndufa10 | 5721 | 0.129 | -0.1821 | No | ||
| 17 | CYC1 | Cyc1 | 5740 | 0.126 | -0.1798 | No | ||
| 18 | SLC18A2 | Slc18a2 | 5750 | 0.125 | -0.1769 | No | ||
| 19 | NDUFV1 | Ndufv1 | 5870 | 0.117 | -0.1814 | No | ||
| 20 | CASP3 | Casp3 | 6007 | 0.107 | -0.1873 | No | ||
| 21 | APAF1 | Apaf1 | 6075 | 0.101 | -0.1888 | No | ||
| 22 | UBB | Ubb | 6088 | 0.100 | -0.1868 | No | ||
| 23 | NDUFS3 | Ndufs3 | 6158 | 0.093 | -0.1887 | No | ||
| 24 | COX7A2L | Cox7a2l | 6167 | 0.093 | -0.1867 | No | ||
| 25 | NDUFB5 | Ndufb5 | 6335 | 0.082 | -0.1952 | No | ||
| 26 | CYCS | Cycs | 7103 | 0.027 | -0.2443 | No | ||
| 27 | NDUFS2 | Ndufs2 | 7378 | 0.006 | -0.2619 | No | ||
| 28 | UQCRC2 | Uqcrc2 | 7439 | 0.002 | -0.2657 | No | ||
| 29 | TH | Th | 7493 | 0.000 | -0.2692 | No | ||
| 30 | COX7B2 | Cox7b2 | 7652 | 0.000 | -0.2794 | No | ||
| 31 | MT-CO2 | mt-Co2 | 7705 | 0.000 | -0.2828 | No | ||
| 32 | NDUFA4L2 | Ndufa4l2 | 8009 | -0.011 | -0.3021 | No | ||
| 33 | NDUFS7 | Ndufs7 | 8011 | -0.011 | -0.3019 | No | ||
| 34 | ATP5F1C | Atp5c1 | 8083 | -0.016 | -0.3061 | No | ||
| 35 | SLC25A4 | Slc25a4 | 8214 | -0.025 | -0.3138 | No | ||
| 36 | COX6A2 | Cox6a2 | 8390 | -0.037 | -0.3241 | No | ||
| 37 | ATP5F1B | Atp5b | 8399 | -0.037 | -0.3236 | No | ||
| 38 | NDUFB9 | Ndufb9 | 8817 | -0.066 | -0.3488 | No | ||
| 39 | NDUFAB1 | Ndufab1 | 9031 | -0.081 | -0.3604 | No | ||
| 40 | NDUFA9 | Ndufa9 | 9127 | -0.088 | -0.3641 | No | ||
| 41 | VDAC1 | Vdac1 | 9299 | -0.099 | -0.3725 | No | ||
| 42 | NDUFS4 | Ndufs4 | 9344 | -0.103 | -0.3725 | No | ||
| 43 | COX5B | Cox5b | 9355 | -0.103 | -0.3703 | No | ||
| 44 | SDHC | Sdhc | 9435 | -0.110 | -0.3724 | No | ||
| 45 | COX4I1 | Cox4i1 | 9535 | -0.118 | -0.3755 | No | ||
| 46 | SNCA | Snca | 9568 | -0.120 | -0.3743 | No | ||
| 47 | ATP5F1A | Atp5a1 | 9760 | -0.135 | -0.3830 | No | ||
| 48 | VDAC3 | Vdac3 | 9787 | -0.138 | -0.3808 | No | ||
| 49 | COX7C | Cox7c | 9876 | -0.145 | -0.3825 | No | ||
| 50 | NDUFS1 | Ndufs1 | 9952 | -0.151 | -0.3832 | No | ||
| 51 | COX6B1 | Cox6b1 | 10014 | -0.155 | -0.3829 | No | ||
| 52 | NDUFV2 | Ndufv2 | 10048 | -0.157 | -0.3807 | No | ||
| 53 | HTRA2 | Htra2 | 10049 | -0.157 | -0.3764 | No | ||
| 54 | UBE2J1 | Ube2j1 | 10179 | -0.167 | -0.3801 | No | ||
| 55 | MT-ND4 | mt-Nd4 | 10184 | -0.167 | -0.3758 | No | ||
| 56 | SDHB | Sdhb | 10474 | -0.190 | -0.3893 | No | ||
| 57 | CASP9 | Casp9 | 10476 | -0.190 | -0.3841 | No | ||
| 58 | VDAC2 | Vdac2 | 10516 | -0.193 | -0.3813 | No | ||
| 59 | NDUFB10 | Ndufb10 | 10571 | -0.197 | -0.3793 | No | ||
| 60 | NDUFB3 | Ndufb3 | 10631 | -0.200 | -0.3776 | No | ||
| 61 | NDUFA8 | Ndufa8 | 10676 | -0.204 | -0.3748 | No | ||
| 62 | COX5A | Cox5a | 10781 | -0.213 | -0.3757 | No | ||
| 63 | COX7A2 | Cox7a2 | 11224 | -0.255 | -0.3973 | Yes | ||
| 64 | UBE2G2 | Ube2g2 | 11245 | -0.257 | -0.3915 | Yes | ||
| 65 | MT-CYB | mt-Cytb | 11374 | -0.270 | -0.3924 | Yes | ||
| 66 | UQCRC1 | Uqcrc1 | 11468 | -0.278 | -0.3907 | Yes | ||
| 67 | UBE2G1 | Ube2g1 | 11525 | -0.283 | -0.3865 | Yes | ||
| 68 | MT-ND6 | mt-Nd6 | 11587 | -0.289 | -0.3825 | Yes | ||
| 69 | NDUFB1 | Ndufb1-ps | 11756 | -0.306 | -0.3849 | Yes | ||
| 70 | COX8A | Cox8a | 11885 | -0.321 | -0.3844 | Yes | ||
| 71 | UQCRB | Uqcrb | 11992 | -0.334 | -0.3820 | Yes | ||
| 72 | PINK1 | Pink1 | 12112 | -0.345 | -0.3802 | Yes | ||
| 73 | PARK7 | Park7 | 12255 | -0.361 | -0.3794 | Yes | ||
| 74 | ATP5MC3 | Atp5g3 | 12291 | -0.365 | -0.3716 | Yes | ||
| 75 | NDUFB6 | Ndufb6 | 12354 | -0.372 | -0.3653 | Yes | ||
| 76 | NDUFA3 | Ndufa3 | 12359 | -0.373 | -0.3553 | Yes | ||
| 77 | ATP5F1D | Atp5d | 12403 | -0.379 | -0.3476 | Yes | ||
| 78 | MT-ND2 | mt-Nd2 | 12428 | -0.382 | -0.3386 | Yes | ||
| 79 | ATP5PD | Atp5h | 12479 | -0.388 | -0.3311 | Yes | ||
| 80 | COX7B | Cox7b | 12512 | -0.392 | -0.3224 | Yes | ||
| 81 | COX7A1 | Cox7a1 | 12522 | -0.394 | -0.3121 | Yes | ||
| 82 | MT-CO1 | mt-Co1 | 12585 | -0.401 | -0.3050 | Yes | ||
| 83 | MT-ND5 | mt-Nd5 | 12840 | -0.436 | -0.3094 | Yes | ||
| 84 | MT-ATP6 | mt-Atp6 | 12940 | -0.443 | -0.3036 | Yes | ||
| 85 | UCHL1 | Uchl1 | 13080 | -0.458 | -0.3000 | Yes | ||
| 86 | NDUFB7 | Ndufb7 | 13279 | -0.492 | -0.2992 | Yes | ||
| 87 | MT-CO3 | mt-Co3 | 13320 | -0.498 | -0.2881 | Yes | ||
| 88 | ATP5F1E | Atp5e | 13433 | -0.508 | -0.2813 | Yes | ||
| 89 | COX6C | Cox6c | 13521 | -0.523 | -0.2725 | Yes | ||
| 90 | NDUFB4 | Ndufb4 | 13601 | -0.534 | -0.2629 | Yes | ||
| 91 | NDUFS8 | Ndufs8 | 13664 | -0.545 | -0.2518 | Yes | ||
| 92 | NDUFA1 | Ndufa1 | 13759 | -0.566 | -0.2423 | Yes | ||
| 93 | ATP5PF | Atp5j | 13797 | -0.573 | -0.2288 | Yes | ||
| 94 | NDUFS6 | Ndufs6 | 13984 | -0.610 | -0.2241 | Yes | ||
| 95 | NDUFA2 | Ndufa2 | 14058 | -0.626 | -0.2115 | Yes | ||
| 96 | NDUFV3 | Ndufv3 | 14135 | -0.645 | -0.1986 | Yes | ||
| 97 | MT-ND1 | mt-Nd1 | 14152 | -0.649 | -0.1817 | Yes | ||
| 98 | UQCR10 | Uqcr10 | 14215 | -0.666 | -0.1673 | Yes | ||
| 99 | NDUFA4 | Ndufa4 | 14228 | -0.671 | -0.1496 | Yes | ||
| 100 | COX6A1 | Cox6a1 | 14276 | -0.686 | -0.1337 | Yes | ||
| 101 | UQCRQ | Uqcrq | 14286 | -0.690 | -0.1152 | Yes | ||
| 102 | NDUFC1 | Ndufc1 | 14306 | -0.695 | -0.0972 | Yes | ||
| 103 | NDUFA5 | Ndufa5 | 14324 | -0.701 | -0.0789 | Yes | ||
| 104 | LRRK2 | Lrrk2 | 14662 | -0.827 | -0.0779 | Yes | ||
| 105 | NDUFS5 | Ndufs5 | 14975 | -0.990 | -0.0708 | Yes | ||
| 106 | UBA7 | Uba7 | 15125 | -1.126 | -0.0494 | Yes | ||
| 107 | UBE2L6 | Ube2l6 | 15270 | -1.316 | -0.0223 | Yes | ||
| 108 | SLC18A1 | Slc18a1 | 15313 | -1.405 | 0.0138 | Yes |