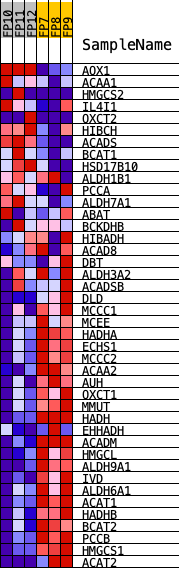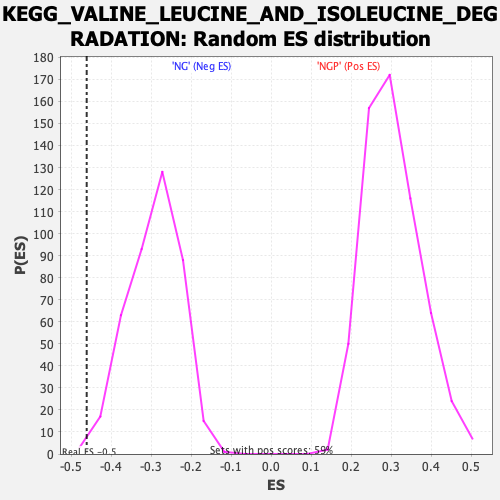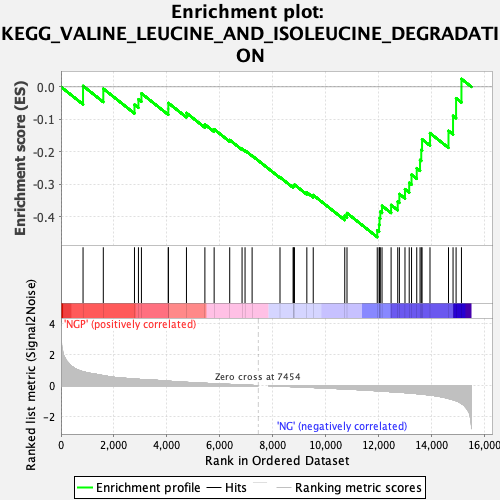
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_NG.phenotype.cls#NGP_versus_NG_repos |
| Phenotype | phenotype.cls#NGP_versus_NG_repos |
| Upregulated in class | NG |
| GeneSet | KEGG_VALINE_LEUCINE_AND_ISOLEUCINE_DEGRADATION |
| Enrichment Score (ES) | -0.46189716 |
| Normalized Enrichment Score (NES) | -1.5778615 |
| Nominal p-value | 0.0024509805 |
| FDR q-value | 0.102789275 |
| FWER p-Value | 0.637 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | AOX1 | Aox1 | 834 | 0.897 | 0.0029 | No | ||
| 2 | ACAA1 | Acaa1a | 1600 | 0.647 | -0.0056 | No | ||
| 3 | HMGCS2 | Hmgcs2 | 2779 | 0.428 | -0.0546 | No | ||
| 4 | IL4I1 | Il4i1 | 2929 | 0.412 | -0.0382 | No | ||
| 5 | OXCT2 | Oxct2a | 3042 | 0.401 | -0.0201 | No | ||
| 6 | HIBCH | Hibch | 4059 | 0.286 | -0.0676 | No | ||
| 7 | ACADS | Acads | 4060 | 0.286 | -0.0495 | No | ||
| 8 | BCAT1 | Bcat1 | 4745 | 0.213 | -0.0802 | No | ||
| 9 | HSD17B10 | Hsd17b10 | 5442 | 0.153 | -0.1155 | No | ||
| 10 | ALDH1B1 | Aldh1b1 | 5792 | 0.122 | -0.1303 | No | ||
| 11 | PCCA | Pcca | 6381 | 0.079 | -0.1633 | No | ||
| 12 | ALDH7A1 | Aldh7a1 | 6848 | 0.045 | -0.1906 | No | ||
| 13 | ABAT | Abat | 6961 | 0.037 | -0.1955 | No | ||
| 14 | BCKDHB | Bckdhb | 7228 | 0.017 | -0.2116 | No | ||
| 15 | HIBADH | Hibadh | 8283 | -0.029 | -0.2778 | No | ||
| 16 | ACAD8 | Acad8 | 8778 | -0.063 | -0.3057 | No | ||
| 17 | DBT | Dbt | 8796 | -0.064 | -0.3028 | No | ||
| 18 | ALDH3A2 | Aldh3a2 | 8830 | -0.067 | -0.3006 | No | ||
| 19 | ACADSB | Acadsb | 9298 | -0.099 | -0.3245 | No | ||
| 20 | DLD | Dld | 9540 | -0.118 | -0.3326 | No | ||
| 21 | MCCC1 | Mccc1 | 10732 | -0.209 | -0.3963 | No | ||
| 22 | MCEE | Mcee | 10817 | -0.216 | -0.3881 | No | ||
| 23 | HADHA | Hadha | 11961 | -0.331 | -0.4410 | Yes | ||
| 24 | ECHS1 | Echs1 | 12030 | -0.339 | -0.4239 | Yes | ||
| 25 | MCCC2 | Mccc2 | 12048 | -0.341 | -0.4034 | Yes | ||
| 26 | ACAA2 | Acaa2 | 12077 | -0.344 | -0.3835 | Yes | ||
| 27 | AUH | Auh | 12142 | -0.349 | -0.3656 | Yes | ||
| 28 | OXCT1 | Oxct1 | 12485 | -0.389 | -0.3631 | Yes | ||
| 29 | MMUT | Mmut | 12736 | -0.421 | -0.3526 | Yes | ||
| 30 | HADH | Hadh | 12798 | -0.430 | -0.3294 | Yes | ||
| 31 | EHHADH | Ehhadh | 13010 | -0.448 | -0.3147 | Yes | ||
| 32 | ACADM | Acadm | 13170 | -0.472 | -0.2951 | Yes | ||
| 33 | HMGCL | Hmgcl | 13258 | -0.487 | -0.2699 | Yes | ||
| 34 | ALDH9A1 | Aldh9a1 | 13455 | -0.514 | -0.2501 | Yes | ||
| 35 | IVD | Ivd | 13580 | -0.529 | -0.2246 | Yes | ||
| 36 | ALDH6A1 | Aldh6a1 | 13626 | -0.537 | -0.1936 | Yes | ||
| 37 | ACAT1 | Acat1 | 13656 | -0.543 | -0.1612 | Yes | ||
| 38 | HADHB | Hadhb | 13956 | -0.605 | -0.1422 | Yes | ||
| 39 | BCAT2 | Bcat2 | 14656 | -0.825 | -0.1352 | Yes | ||
| 40 | PCCB | Pccb | 14828 | -0.906 | -0.0890 | Yes | ||
| 41 | HMGCS1 | Hmgcs1 | 14943 | -0.970 | -0.0350 | Yes | ||
| 42 | ACAT2 | Acat2 | 15146 | -1.147 | 0.0245 | Yes |