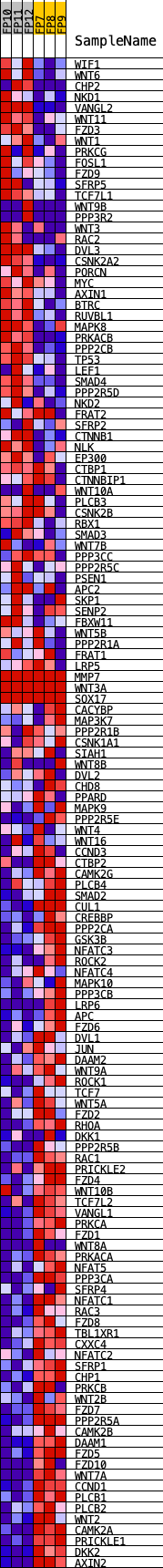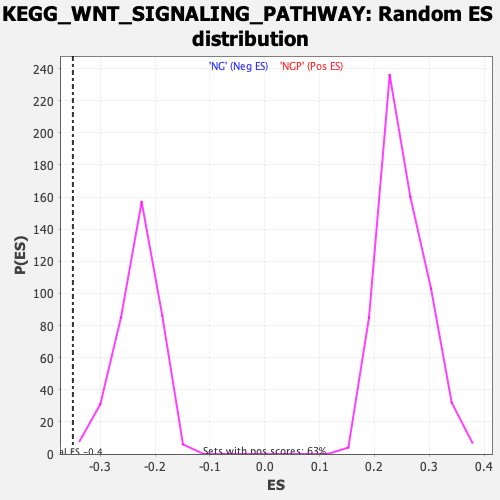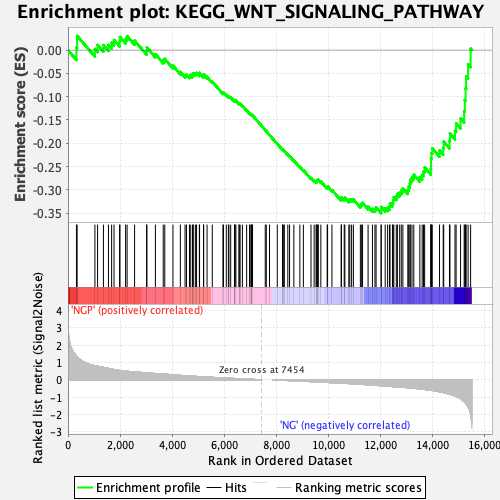
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_NG.phenotype.cls#NGP_versus_NG_repos |
| Phenotype | phenotype.cls#NGP_versus_NG_repos |
| Upregulated in class | NG |
| GeneSet | KEGG_WNT_SIGNALING_PATHWAY |
| Enrichment Score (ES) | -0.35001454 |
| Normalized Enrichment Score (NES) | -1.5041187 |
| Nominal p-value | 0.0026809652 |
| FDR q-value | 0.14791079 |
| FWER p-Value | 0.83 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | WIF1 | Wif1 | 325 | 1.369 | 0.0058 | No | ||
| 2 | WNT6 | Wnt6 | 350 | 1.331 | 0.0305 | No | ||
| 3 | CHP2 | Chp2 | 1035 | 0.819 | 0.0022 | No | ||
| 4 | NKD1 | Nkd1 | 1137 | 0.786 | 0.0111 | No | ||
| 5 | VANGL2 | Vangl2 | 1362 | 0.721 | 0.0108 | No | ||
| 6 | WNT11 | Wnt11 | 1556 | 0.660 | 0.0113 | No | ||
| 7 | FZD3 | Fzd3 | 1677 | 0.626 | 0.0158 | No | ||
| 8 | WNT1 | Wnt1 | 1769 | 0.596 | 0.0216 | No | ||
| 9 | PRKCG | Prkcg | 1985 | 0.540 | 0.0183 | No | ||
| 10 | FOSL1 | Fosl1 | 1997 | 0.539 | 0.0282 | No | ||
| 11 | FZD9 | Fzd9 | 2215 | 0.500 | 0.0239 | No | ||
| 12 | SFRP5 | Sfrp5 | 2272 | 0.492 | 0.0300 | No | ||
| 13 | TCF7L1 | Tcf7l1 | 2560 | 0.450 | 0.0202 | No | ||
| 14 | WNT9B | Wnt9b | 3022 | 0.401 | -0.0018 | No | ||
| 15 | PPP3R2 | Ppp3r2 | 3033 | 0.401 | 0.0054 | No | ||
| 16 | WNT3 | Wnt3 | 3360 | 0.359 | -0.0087 | No | ||
| 17 | RAC2 | Rac2 | 3662 | 0.338 | -0.0216 | No | ||
| 18 | DVL3 | Dvl3 | 3721 | 0.328 | -0.0189 | No | ||
| 19 | CSNK2A2 | Csnk2a2 | 4035 | 0.289 | -0.0335 | No | ||
| 20 | PORCN | Porcn | 4319 | 0.257 | -0.0469 | No | ||
| 21 | MYC | Myc | 4500 | 0.237 | -0.0539 | No | ||
| 22 | AXIN1 | Axin1 | 4555 | 0.232 | -0.0528 | No | ||
| 23 | BTRC | Btrc | 4669 | 0.220 | -0.0558 | No | ||
| 24 | RUVBL1 | Ruvbl1 | 4700 | 0.217 | -0.0535 | No | ||
| 25 | MAPK8 | Mapk8 | 4781 | 0.209 | -0.0546 | No | ||
| 26 | PRKACB | Prkacb | 4785 | 0.209 | -0.0507 | No | ||
| 27 | PPP2CB | Ppp2cb | 4832 | 0.205 | -0.0496 | No | ||
| 28 | TP53 | Trp53 | 4907 | 0.199 | -0.0505 | No | ||
| 29 | LEF1 | Lef1 | 4943 | 0.195 | -0.0489 | No | ||
| 30 | SMAD4 | Smad4 | 5051 | 0.185 | -0.0522 | No | ||
| 31 | PPP2R5D | Ppp2r5d | 5058 | 0.185 | -0.0490 | No | ||
| 32 | NKD2 | Nkd2 | 5213 | 0.171 | -0.0556 | No | ||
| 33 | FRAT2 | Frat2 | 5214 | 0.171 | -0.0522 | No | ||
| 34 | SFRP2 | Sfrp2 | 5346 | 0.160 | -0.0576 | No | ||
| 35 | CTNNB1 | Ctnnb1 | 5547 | 0.143 | -0.0678 | No | ||
| 36 | NLK | Nlk | 5957 | 0.111 | -0.0922 | No | ||
| 37 | EP300 | Ep300 | 5976 | 0.109 | -0.0912 | No | ||
| 38 | CTBP1 | Ctbp1 | 6085 | 0.101 | -0.0962 | No | ||
| 39 | CTNNBIP1 | Ctnnbip1 | 6166 | 0.093 | -0.0996 | No | ||
| 40 | WNT10A | Wnt10a | 6186 | 0.091 | -0.0990 | No | ||
| 41 | PLCB3 | Plcb3 | 6255 | 0.087 | -0.1017 | No | ||
| 42 | CSNK2B | Csnk2b | 6402 | 0.078 | -0.1097 | No | ||
| 43 | RBX1 | Rbx1 | 6414 | 0.077 | -0.1089 | No | ||
| 44 | SMAD3 | Smad3 | 6415 | 0.077 | -0.1074 | No | ||
| 45 | WNT7B | Wnt7b | 6455 | 0.074 | -0.1085 | No | ||
| 46 | PPP3CC | Ppp3cc | 6577 | 0.065 | -0.1150 | No | ||
| 47 | PPP2R5C | Ppp2r5c | 6579 | 0.065 | -0.1138 | No | ||
| 48 | PSEN1 | Psen1 | 6617 | 0.063 | -0.1150 | No | ||
| 49 | APC2 | Apc2 | 6705 | 0.056 | -0.1196 | No | ||
| 50 | SKP1 | Skp1a | 6868 | 0.043 | -0.1292 | No | ||
| 51 | SENP2 | Senp2 | 6980 | 0.035 | -0.1357 | No | ||
| 52 | FBXW11 | Fbxw11 | 7033 | 0.032 | -0.1385 | No | ||
| 53 | WNT5B | Wnt5b | 7043 | 0.031 | -0.1385 | No | ||
| 54 | PPP2R1A | Ppp2r1a | 7070 | 0.029 | -0.1396 | No | ||
| 55 | FRAT1 | Frat1 | 7071 | 0.029 | -0.1390 | No | ||
| 56 | LRP5 | Lrp5 | 7094 | 0.027 | -0.1399 | No | ||
| 57 | MMP7 | Mmp7 | 7583 | 0.000 | -0.1716 | No | ||
| 58 | WNT3A | Wnt3a | 7623 | 0.000 | -0.1742 | No | ||
| 59 | SOX17 | Sox17 | 7748 | 0.000 | -0.1822 | No | ||
| 60 | CACYBP | Cacybp | 8046 | -0.014 | -0.2013 | No | ||
| 61 | MAP3K7 | Map3k7 | 8246 | -0.027 | -0.2137 | No | ||
| 62 | PPP2R1B | Ppp2r1b | 8265 | -0.028 | -0.2143 | No | ||
| 63 | CSNK1A1 | Csnk1a1 | 8272 | -0.029 | -0.2141 | No | ||
| 64 | SIAH1 | Siah1a | 8317 | -0.033 | -0.2163 | No | ||
| 65 | WNT8B | Wnt8b | 8451 | -0.041 | -0.2242 | No | ||
| 66 | DVL2 | Dvl2 | 8518 | -0.046 | -0.2275 | No | ||
| 67 | CHD8 | Chd8 | 8683 | -0.057 | -0.2371 | No | ||
| 68 | PPARD | Ppard | 8914 | -0.073 | -0.2506 | No | ||
| 69 | MAPK9 | Mapk9 | 9049 | -0.083 | -0.2577 | No | ||
| 70 | PPP2R5E | Ppp2r5e | 9342 | -0.103 | -0.2746 | No | ||
| 71 | WNT4 | Wnt4 | 9463 | -0.111 | -0.2802 | No | ||
| 72 | WNT16 | Wnt16 | 9537 | -0.118 | -0.2826 | No | ||
| 73 | CCND3 | Ccnd3 | 9564 | -0.120 | -0.2820 | No | ||
| 74 | CTBP2 | Ctbp2 | 9575 | -0.121 | -0.2802 | No | ||
| 75 | CAMK2G | Camk2g | 9587 | -0.121 | -0.2786 | No | ||
| 76 | PLCB4 | Plcb4 | 9624 | -0.125 | -0.2784 | No | ||
| 77 | SMAD2 | Smad2 | 9721 | -0.132 | -0.2821 | No | ||
| 78 | CUL1 | Cul1 | 9964 | -0.152 | -0.2948 | No | ||
| 79 | CREBBP | Crebbp | 9980 | -0.152 | -0.2928 | No | ||
| 80 | PPP2CA | Ppp2ca | 10147 | -0.165 | -0.3003 | No | ||
| 81 | GSK3B | Gsk3b | 10504 | -0.192 | -0.3197 | No | ||
| 82 | NFATC3 | Nfatc3 | 10510 | -0.192 | -0.3162 | No | ||
| 83 | ROCK2 | Rock2 | 10619 | -0.200 | -0.3193 | No | ||
| 84 | NFATC4 | Nfatc4 | 10645 | -0.202 | -0.3169 | No | ||
| 85 | MAPK10 | Mapk10 | 10797 | -0.215 | -0.3225 | No | ||
| 86 | PPP3CB | Ppp3cb | 10834 | -0.218 | -0.3206 | No | ||
| 87 | LRP6 | Lrp6 | 10904 | -0.223 | -0.3207 | No | ||
| 88 | APC | Apc | 10972 | -0.230 | -0.3205 | No | ||
| 89 | FZD6 | Fzd6 | 11240 | -0.257 | -0.3328 | No | ||
| 90 | DVL1 | Dvl1 | 11277 | -0.260 | -0.3300 | No | ||
| 91 | JUN | Jun | 11324 | -0.264 | -0.3278 | No | ||
| 92 | DAAM2 | Daam2 | 11537 | -0.284 | -0.3360 | No | ||
| 93 | WNT9A | Wnt9a | 11705 | -0.301 | -0.3409 | No | ||
| 94 | ROCK1 | Rock1 | 11801 | -0.311 | -0.3409 | No | ||
| 95 | TCF7 | Tcf7 | 11844 | -0.317 | -0.3374 | No | ||
| 96 | WNT5A | Wnt5a | 12039 | -0.341 | -0.3433 | Yes | ||
| 97 | FZD2 | Fzd2 | 12041 | -0.341 | -0.3367 | Yes | ||
| 98 | RHOA | Rhoa | 12195 | -0.354 | -0.3396 | Yes | ||
| 99 | DKK1 | Dkk1 | 12288 | -0.365 | -0.3384 | Yes | ||
| 100 | PPP2R5B | Ppp2r5b | 12360 | -0.373 | -0.3357 | Yes | ||
| 101 | RAC1 | Rac1 | 12380 | -0.375 | -0.3295 | Yes | ||
| 102 | PRICKLE2 | Prickle2 | 12474 | -0.388 | -0.3279 | Yes | ||
| 103 | FZD4 | Fzd4 | 12501 | -0.391 | -0.3219 | Yes | ||
| 104 | WNT10B | Wnt10b | 12526 | -0.394 | -0.3157 | Yes | ||
| 105 | TCF7L2 | Tcf7l2 | 12621 | -0.404 | -0.3139 | Yes | ||
| 106 | VANGL1 | Vangl1 | 12665 | -0.410 | -0.3086 | Yes | ||
| 107 | PRKCA | Prkca | 12751 | -0.423 | -0.3058 | Yes | ||
| 108 | FZD1 | Fzd1 | 12813 | -0.432 | -0.3012 | Yes | ||
| 109 | WNT8A | Wnt8a | 12875 | -0.438 | -0.2966 | Yes | ||
| 110 | PRKACA | Prkaca | 13065 | -0.456 | -0.2999 | Yes | ||
| 111 | NFAT5 | Nfat5 | 13098 | -0.460 | -0.2929 | Yes | ||
| 112 | PPP3CA | Ppp3ca | 13145 | -0.468 | -0.2867 | Yes | ||
| 113 | SFRP4 | Sfrp4 | 13160 | -0.471 | -0.2783 | Yes | ||
| 114 | NFATC1 | Nfatc1 | 13225 | -0.480 | -0.2730 | Yes | ||
| 115 | RAC3 | Rac3 | 13294 | -0.495 | -0.2677 | Yes | ||
| 116 | FZD8 | Fzd8 | 13528 | -0.523 | -0.2725 | Yes | ||
| 117 | TBL1XR1 | Tbl1xr1 | 13617 | -0.535 | -0.2677 | Yes | ||
| 118 | CXXC4 | Cxxc4 | 13669 | -0.546 | -0.2602 | Yes | ||
| 119 | NFATC2 | Nfatc2 | 13711 | -0.555 | -0.2520 | Yes | ||
| 120 | SFRP1 | Sfrp1 | 13949 | -0.603 | -0.2555 | Yes | ||
| 121 | CHP1 | Chp1 | 13957 | -0.605 | -0.2440 | Yes | ||
| 122 | PRKCB | Prkcb | 13958 | -0.605 | -0.2321 | Yes | ||
| 123 | WNT2B | Wnt2b | 13977 | -0.609 | -0.2213 | Yes | ||
| 124 | FZD7 | Fzd7 | 14004 | -0.614 | -0.2109 | Yes | ||
| 125 | PPP2R5A | Ppp2r5a | 14284 | -0.689 | -0.2154 | Yes | ||
| 126 | CAMK2B | Camk2b | 14425 | -0.735 | -0.2100 | Yes | ||
| 127 | DAAM1 | Daam1 | 14445 | -0.743 | -0.1966 | Yes | ||
| 128 | FZD5 | Fzd5 | 14666 | -0.828 | -0.1946 | Yes | ||
| 129 | FZD10 | Fzd10 | 14685 | -0.842 | -0.1792 | Yes | ||
| 130 | WNT7A | Wnt7a | 14879 | -0.933 | -0.1733 | Yes | ||
| 131 | CCND1 | Ccnd1 | 14921 | -0.960 | -0.1571 | Yes | ||
| 132 | PLCB1 | Plcb1 | 15093 | -1.088 | -0.1468 | Yes | ||
| 133 | PLCB2 | Plcb2 | 15231 | -1.247 | -0.1311 | Yes | ||
| 134 | WNT2 | Wnt2 | 15256 | -1.292 | -0.1072 | Yes | ||
| 135 | CAMK2A | Camk2a | 15284 | -1.351 | -0.0824 | Yes | ||
| 136 | PRICKLE1 | Prickle1 | 15307 | -1.389 | -0.0564 | Yes | ||
| 137 | DKK2 | Dkk2 | 15382 | -1.564 | -0.0304 | Yes | ||
| 138 | AXIN2 | Axin2 | 15479 | -2.011 | 0.0030 | Yes |