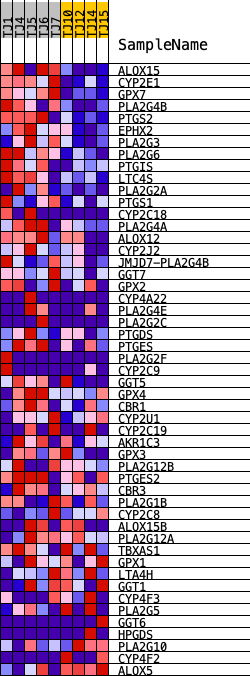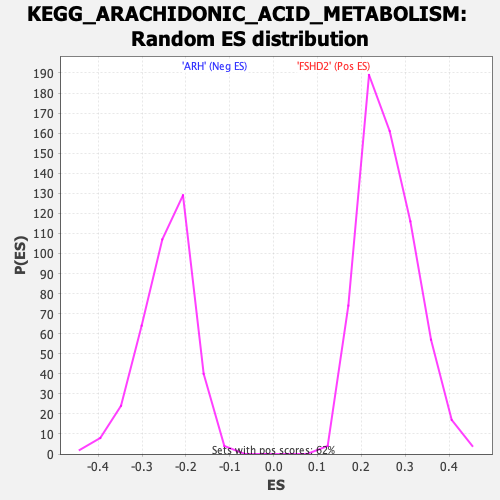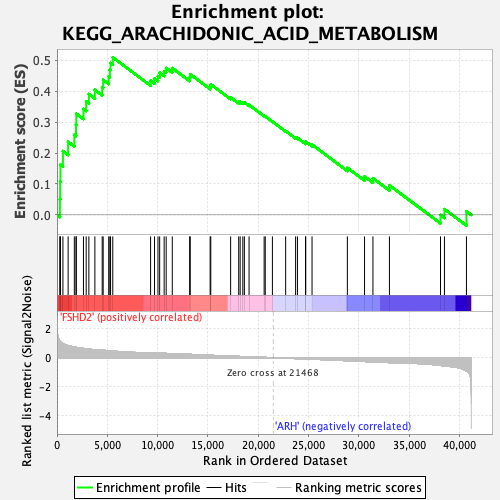
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | FSHD2 |
| GeneSet | KEGG_ARACHIDONIC_ACID_METABOLISM |
| Enrichment Score (ES) | 0.5096648 |
| Normalized Enrichment Score (NES) | 1.9520618 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.069167845 |
| FWER p-Value | 0.279 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ALOX15 | ENSG00000161905.12 | 287 | 1.141 | 0.0510 | Yes | ||
| 2 | CYP2E1 | ENSG00000130649.10 | 315 | 1.122 | 0.1074 | Yes | ||
| 3 | GPX7 | ENSG00000116157.6 | 342 | 1.095 | 0.1624 | Yes | ||
| 4 | PLA2G4B | ENSG00000243708.10 | 588 | 0.984 | 0.2064 | Yes | ||
| 5 | PTGS2 | ENSG00000073756.12 | 1105 | 0.837 | 0.2364 | Yes | ||
| 6 | EPHX2 | ENSG00000120915.14 | 1723 | 0.744 | 0.2592 | Yes | ||
| 7 | PLA2G3 | ENSG00000100078.4 | 1889 | 0.718 | 0.2917 | Yes | ||
| 8 | PLA2G6 | ENSG00000184381.20 | 1918 | 0.714 | 0.3274 | Yes | ||
| 9 | PTGIS | ENSG00000124212.6 | 2629 | 0.640 | 0.3427 | Yes | ||
| 10 | LTC4S | ENSG00000213316.10 | 2899 | 0.614 | 0.3673 | Yes | ||
| 11 | PLA2G2A | ENSG00000188257.11 | 3165 | 0.594 | 0.3911 | Yes | ||
| 12 | PTGS1 | ENSG00000095303.17 | 3763 | 0.553 | 0.4046 | Yes | ||
| 13 | CYP2C18 | ENSG00000108242.13 | 4489 | 0.518 | 0.4134 | Yes | ||
| 14 | PLA2G4A | ENSG00000116711.10 | 4587 | 0.514 | 0.4371 | Yes | ||
| 15 | ALOX12 | ENSG00000108839.12 | 5137 | 0.481 | 0.4482 | Yes | ||
| 16 | CYP2J2 | ENSG00000134716.11 | 5250 | 0.474 | 0.4696 | Yes | ||
| 17 | JMJD7-PLA2G4B | ENSG00000168970.22 | 5321 | 0.470 | 0.4918 | Yes | ||
| 18 | GGT7 | ENSG00000131067.17 | 5542 | 0.457 | 0.5097 | Yes | ||
| 19 | GPX2 | ENSG00000176153.12 | 9299 | 0.323 | 0.4347 | No | ||
| 20 | CYP4A22 | ENSG00000162365.12 | 9690 | 0.319 | 0.4415 | No | ||
| 21 | PLA2G4E | ENSG00000188089.13 | 10031 | 0.310 | 0.4490 | No | ||
| 22 | PLA2G2C | ENSG00000187980.6 | 10200 | 0.305 | 0.4604 | No | ||
| 23 | PTGDS | ENSG00000107317.13 | 10645 | 0.305 | 0.4651 | No | ||
| 24 | PTGES | ENSG00000148344.11 | 10848 | 0.297 | 0.4753 | No | ||
| 25 | PLA2G2F | ENSG00000158786.4 | 11453 | 0.285 | 0.4751 | No | ||
| 26 | CYP2C9 | ENSG00000138109.11 | 13179 | 0.233 | 0.4449 | No | ||
| 27 | GGT5 | ENSG00000099998.17 | 13238 | 0.231 | 0.4553 | No | ||
| 28 | GPX4 | ENSG00000167468.17 | 15215 | 0.168 | 0.4157 | No | ||
| 29 | CBR1 | ENSG00000159228.13 | 15279 | 0.165 | 0.4226 | No | ||
| 30 | CYP2U1 | ENSG00000155016.18 | 17251 | 0.111 | 0.3802 | No | ||
| 31 | CYP2C19 | ENSG00000165841.11 | 18054 | 0.089 | 0.3653 | No | ||
| 32 | AKR1C3 | ENSG00000196139.14 | 18197 | 0.085 | 0.3662 | No | ||
| 33 | GPX3 | ENSG00000211445.12 | 18462 | 0.080 | 0.3638 | No | ||
| 34 | PLA2G12B | ENSG00000138308.6 | 18616 | 0.076 | 0.3640 | No | ||
| 35 | PTGES2 | ENSG00000148334.15 | 19082 | 0.063 | 0.3558 | No | ||
| 36 | CBR3 | ENSG00000159231.6 | 20580 | 0.023 | 0.3206 | No | ||
| 37 | PLA2G1B | ENSG00000170890.14 | 20692 | 0.020 | 0.3189 | No | ||
| 38 | CYP2C8 | ENSG00000138115.14 | 21401 | 0.001 | 0.3017 | No | ||
| 39 | ALOX15B | ENSG00000179593.16 | 22718 | -0.031 | 0.2713 | No | ||
| 40 | PLA2G12A | ENSG00000123739.11 | 23706 | -0.058 | 0.2502 | No | ||
| 41 | TBXAS1 | ENSG00000059377.17 | 23889 | -0.063 | 0.2490 | No | ||
| 42 | GPX1 | ENSG00000233276.5 | 24689 | -0.082 | 0.2338 | No | ||
| 43 | LTA4H | ENSG00000111144.10 | 24714 | -0.083 | 0.2374 | No | ||
| 44 | GGT1 | ENSG00000100031.19 | 25339 | -0.100 | 0.2273 | No | ||
| 45 | CYP4F3 | ENSG00000186529.16 | 28849 | -0.207 | 0.1525 | No | ||
| 46 | PLA2G5 | ENSG00000127472.11 | 30555 | -0.261 | 0.1243 | No | ||
| 47 | GGT6 | ENSG00000167741.10 | 31389 | -0.285 | 0.1185 | No | ||
| 48 | HPGDS | ENSG00000163106.11 | 33029 | -0.336 | 0.0958 | No | ||
| 49 | PLA2G10 | ENSG00000069764.9 | 38102 | -0.535 | -0.0004 | No | ||
| 50 | CYP4F2 | ENSG00000186115.13 | 38494 | -0.558 | 0.0184 | No | ||
| 51 | ALOX5 | ENSG00000012779.11 | 40680 | -0.913 | 0.0117 | No |