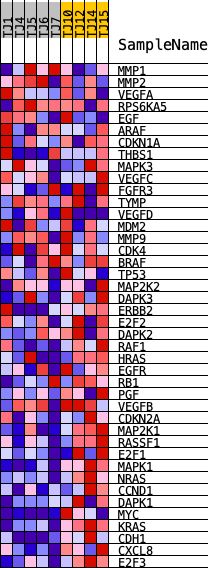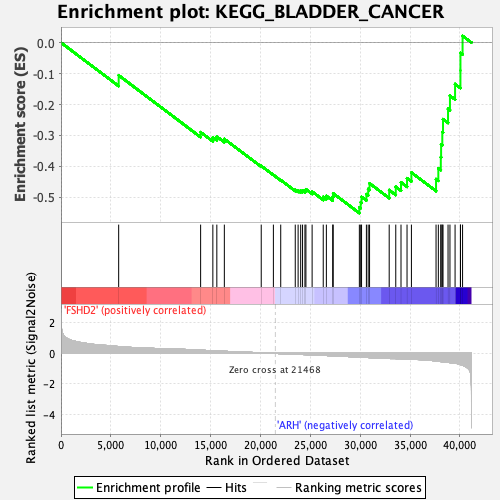
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | ARH |
| GeneSet | KEGG_BLADDER_CANCER |
| Enrichment Score (ES) | -0.55108005 |
| Normalized Enrichment Score (NES) | -2.235398 |
| Nominal p-value | 0.0 |
| FDR q-value | 4.4511445E-4 |
| FWER p-Value | 0.011 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | MMP1 | ENSG00000196611.5 | 5783 | 0.444 | -0.1058 | No | ||
| 2 | MMP2 | ENSG00000087245.13 | 14001 | 0.206 | -0.2895 | No | ||
| 3 | VEGFA | ENSG00000112715.22 | 15227 | 0.167 | -0.3061 | No | ||
| 4 | RPS6KA5 | ENSG00000100784.12 | 15626 | 0.156 | -0.3036 | No | ||
| 5 | EGF | ENSG00000138798.13 | 16379 | 0.135 | -0.3113 | No | ||
| 6 | ARAF | ENSG00000078061.13 | 20078 | 0.037 | -0.3983 | No | ||
| 7 | CDKN1A | ENSG00000124762.13 | 21297 | 0.004 | -0.4276 | No | ||
| 8 | THBS1 | ENSG00000137801.11 | 22033 | -0.015 | -0.4443 | No | ||
| 9 | MAPK3 | ENSG00000102882.12 | 23490 | -0.052 | -0.4756 | No | ||
| 10 | VEGFC | ENSG00000150630.4 | 23777 | -0.060 | -0.4779 | No | ||
| 11 | FGFR3 | ENSG00000068078.18 | 24034 | -0.065 | -0.4790 | No | ||
| 12 | TYMP | ENSG00000025708.14 | 24222 | -0.070 | -0.4780 | No | ||
| 13 | VEGFD | ENSG00000165197.5 | 24461 | -0.076 | -0.4778 | No | ||
| 14 | MDM2 | ENSG00000135679.25 | 24559 | -0.079 | -0.4740 | No | ||
| 15 | MMP9 | ENSG00000100985.7 | 25181 | -0.096 | -0.4816 | No | ||
| 16 | CDK4 | ENSG00000135446.17 | 26296 | -0.129 | -0.4986 | No | ||
| 17 | BRAF | ENSG00000157764.13 | 26610 | -0.138 | -0.4954 | No | ||
| 18 | TP53 | ENSG00000141510.17 | 27245 | -0.156 | -0.4985 | No | ||
| 19 | MAP2K2 | ENSG00000126934.13 | 27294 | -0.158 | -0.4873 | No | ||
| 20 | DAPK3 | ENSG00000167657.14 | 29918 | -0.241 | -0.5321 | Yes | ||
| 21 | ERBB2 | ENSG00000141736.13 | 30053 | -0.246 | -0.5161 | Yes | ||
| 22 | E2F2 | ENSG00000007968.7 | 30138 | -0.249 | -0.4986 | Yes | ||
| 23 | DAPK2 | ENSG00000035664.11 | 30624 | -0.263 | -0.4898 | Yes | ||
| 24 | RAF1 | ENSG00000132155.11 | 30802 | -0.270 | -0.4729 | Yes | ||
| 25 | HRAS | ENSG00000174775.17 | 30941 | -0.275 | -0.4547 | Yes | ||
| 26 | EGFR | ENSG00000146648.18 | 32910 | -0.332 | -0.4765 | Yes | ||
| 27 | RB1 | ENSG00000139687.15 | 33569 | -0.339 | -0.4659 | Yes | ||
| 28 | PGF | ENSG00000119630.14 | 34094 | -0.345 | -0.4515 | Yes | ||
| 29 | VEGFB | ENSG00000173511.9 | 34699 | -0.356 | -0.4383 | Yes | ||
| 30 | CDKN2A | ENSG00000147889.17 | 35138 | -0.371 | -0.4198 | Yes | ||
| 31 | MAP2K1 | ENSG00000169032.9 | 37608 | -0.494 | -0.4411 | Yes | ||
| 32 | RASSF1 | ENSG00000068028.17 | 37838 | -0.515 | -0.4062 | Yes | ||
| 33 | E2F1 | ENSG00000101412.13 | 38085 | -0.533 | -0.3703 | Yes | ||
| 34 | MAPK1 | ENSG00000100030.14 | 38116 | -0.536 | -0.3290 | Yes | ||
| 35 | NRAS | ENSG00000213281.5 | 38236 | -0.545 | -0.2891 | Yes | ||
| 36 | CCND1 | ENSG00000110092.3 | 38307 | -0.552 | -0.2475 | Yes | ||
| 37 | DAPK1 | ENSG00000196730.13 | 38811 | -0.582 | -0.2140 | Yes | ||
| 38 | MYC | ENSG00000136997.19 | 39003 | -0.602 | -0.1714 | Yes | ||
| 39 | KRAS | ENSG00000133703.12 | 39516 | -0.637 | -0.1338 | Yes | ||
| 40 | CDH1 | ENSG00000039068.19 | 40053 | -0.722 | -0.0902 | Yes | ||
| 41 | CXCL8 | ENSG00000169429.11 | 40055 | -0.722 | -0.0335 | Yes | ||
| 42 | E2F3 | ENSG00000112242.15 | 40268 | -0.769 | 0.0217 | Yes |