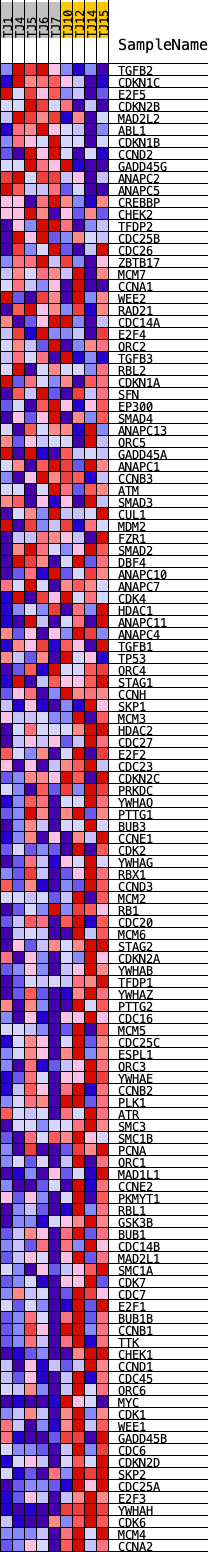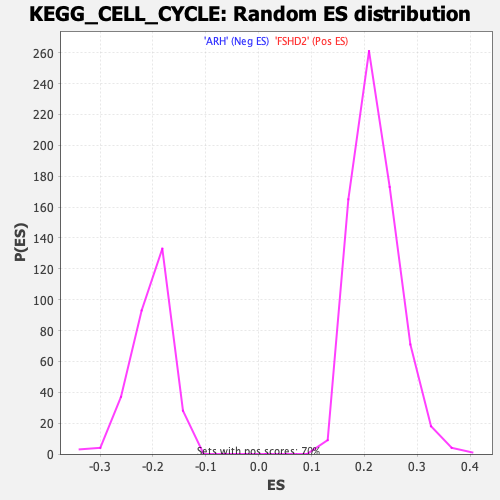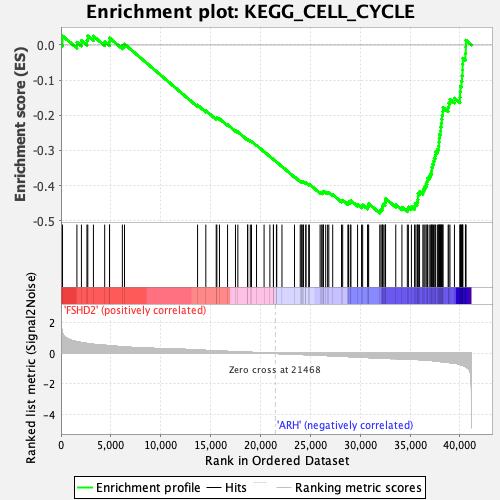
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | ARH |
| GeneSet | KEGG_CELL_CYCLE |
| Enrichment Score (ES) | -0.47864425 |
| Normalized Enrichment Score (NES) | -2.3486214 |
| Nominal p-value | 0.0 |
| FDR q-value | 1.2875805E-4 |
| FWER p-Value | 0.002 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | TGFB2 | ENSG00000092969.12 | 152 | 1.317 | 0.0259 | No | ||
| 2 | CDKN1C | ENSG00000129757.13 | 1596 | 0.759 | 0.0078 | No | ||
| 3 | E2F5 | ENSG00000133740.11 | 2048 | 0.699 | 0.0126 | No | ||
| 4 | CDKN2B | ENSG00000147883.12 | 2601 | 0.644 | 0.0136 | No | ||
| 5 | MAD2L2 | ENSG00000116670.15 | 2687 | 0.635 | 0.0258 | No | ||
| 6 | ABL1 | ENSG00000097007.19 | 3251 | 0.588 | 0.0253 | No | ||
| 7 | CDKN1B | ENSG00000111276.11 | 4380 | 0.521 | 0.0095 | No | ||
| 8 | CCND2 | ENSG00000118971.8 | 4862 | 0.500 | 0.0091 | No | ||
| 9 | GADD45G | ENSG00000130222.11 | 4871 | 0.500 | 0.0201 | No | ||
| 10 | ANAPC2 | ENSG00000176248.9 | 6153 | 0.426 | -0.0015 | No | ||
| 11 | ANAPC5 | ENSG00000089053.13 | 6368 | 0.414 | 0.0026 | No | ||
| 12 | CREBBP | ENSG00000005339.14 | 13694 | 0.216 | -0.1711 | No | ||
| 13 | CHEK2 | ENSG00000183765.22 | 14522 | 0.190 | -0.1869 | No | ||
| 14 | TFDP2 | ENSG00000114126.17 | 15564 | 0.157 | -0.2088 | No | ||
| 15 | CDC25B | ENSG00000101224.17 | 15629 | 0.156 | -0.2068 | No | ||
| 16 | CDC26 | ENSG00000176386.9 | 15890 | 0.148 | -0.2099 | No | ||
| 17 | ZBTB17 | ENSG00000116809.11 | 16702 | 0.125 | -0.2268 | No | ||
| 18 | MCM7 | ENSG00000166508.17 | 17495 | 0.104 | -0.2438 | No | ||
| 19 | CCNA1 | ENSG00000133101.10 | 17734 | 0.098 | -0.2474 | No | ||
| 20 | WEE2 | ENSG00000214102.7 | 18705 | 0.073 | -0.2693 | No | ||
| 21 | RAD21 | ENSG00000164754.14 | 18730 | 0.073 | -0.2683 | No | ||
| 22 | CDC14A | ENSG00000079335.20 | 18982 | 0.065 | -0.2729 | No | ||
| 23 | E2F4 | ENSG00000205250.9 | 19102 | 0.062 | -0.2744 | No | ||
| 24 | ORC2 | ENSG00000115942.9 | 19607 | 0.048 | -0.2856 | No | ||
| 25 | TGFB3 | ENSG00000119699.7 | 20364 | 0.029 | -0.3034 | No | ||
| 26 | RBL2 | ENSG00000103479.16 | 20946 | 0.014 | -0.3172 | No | ||
| 27 | CDKN1A | ENSG00000124762.13 | 21297 | 0.004 | -0.3257 | No | ||
| 28 | SFN | ENSG00000175793.12 | 21306 | 0.004 | -0.3258 | No | ||
| 29 | EP300 | ENSG00000100393.13 | 21602 | -0.004 | -0.3329 | No | ||
| 30 | SMAD4 | ENSG00000141646.13 | 21636 | -0.005 | -0.3336 | No | ||
| 31 | ANAPC13 | ENSG00000129055.12 | 22157 | -0.017 | -0.3459 | No | ||
| 32 | ORC5 | ENSG00000164815.11 | 23413 | -0.050 | -0.3753 | No | ||
| 33 | GADD45A | ENSG00000116717.13 | 23989 | -0.064 | -0.3879 | No | ||
| 34 | ANAPC1 | ENSG00000153107.13 | 24092 | -0.067 | -0.3889 | No | ||
| 35 | CCNB3 | ENSG00000147082.17 | 24164 | -0.069 | -0.3891 | No | ||
| 36 | ATM | ENSG00000149311.18 | 24258 | -0.072 | -0.3897 | No | ||
| 37 | SMAD3 | ENSG00000166949.16 | 24321 | -0.074 | -0.3896 | No | ||
| 38 | CUL1 | ENSG00000055130.17 | 24550 | -0.079 | -0.3934 | No | ||
| 39 | MDM2 | ENSG00000135679.25 | 24559 | -0.079 | -0.3918 | No | ||
| 40 | FZR1 | ENSG00000105325.14 | 24842 | -0.087 | -0.3967 | No | ||
| 41 | SMAD2 | ENSG00000175387.16 | 24905 | -0.088 | -0.3962 | No | ||
| 42 | DBF4 | ENSG00000006634.8 | 25980 | -0.120 | -0.4197 | No | ||
| 43 | ANAPC10 | ENSG00000164162.14 | 26104 | -0.124 | -0.4199 | No | ||
| 44 | ANAPC7 | ENSG00000196510.12 | 26263 | -0.128 | -0.4209 | No | ||
| 45 | CDK4 | ENSG00000135446.17 | 26296 | -0.129 | -0.4188 | No | ||
| 46 | HDAC1 | ENSG00000116478.12 | 26306 | -0.129 | -0.4161 | No | ||
| 47 | ANAPC11 | ENSG00000141552.17 | 26543 | -0.136 | -0.4188 | No | ||
| 48 | ANAPC4 | ENSG00000053900.10 | 26737 | -0.142 | -0.4203 | No | ||
| 49 | TGFB1 | ENSG00000105329.10 | 26855 | -0.145 | -0.4199 | No | ||
| 50 | TP53 | ENSG00000141510.17 | 27245 | -0.156 | -0.4259 | No | ||
| 51 | ORC4 | ENSG00000115947.13 | 28129 | -0.185 | -0.4432 | No | ||
| 52 | STAG1 | ENSG00000118007.13 | 28245 | -0.188 | -0.4418 | No | ||
| 53 | CCNH | ENSG00000134480.15 | 28769 | -0.204 | -0.4500 | No | ||
| 54 | SKP1 | ENSG00000113558.18 | 28827 | -0.206 | -0.4467 | No | ||
| 55 | MCM3 | ENSG00000112118.20 | 29007 | -0.212 | -0.4463 | No | ||
| 56 | HDAC2 | ENSG00000196591.12 | 29075 | -0.214 | -0.4431 | No | ||
| 57 | CDC27 | ENSG00000004897.12 | 29738 | -0.236 | -0.4540 | No | ||
| 58 | E2F2 | ENSG00000007968.7 | 30138 | -0.249 | -0.4581 | No | ||
| 59 | CDC23 | ENSG00000094880.11 | 30249 | -0.252 | -0.4551 | No | ||
| 60 | CDKN2C | ENSG00000123080.12 | 30743 | -0.267 | -0.4611 | No | ||
| 61 | PRKDC | ENSG00000253729.7 | 30793 | -0.269 | -0.4562 | No | ||
| 62 | YWHAQ | ENSG00000134308.14 | 30852 | -0.271 | -0.4515 | No | ||
| 63 | PTTG1 | ENSG00000164611.13 | 31965 | -0.295 | -0.4720 | Yes | ||
| 64 | BUB3 | ENSG00000154473.18 | 32087 | -0.300 | -0.4682 | Yes | ||
| 65 | CCNE1 | ENSG00000105173.14 | 32197 | -0.305 | -0.4640 | Yes | ||
| 66 | CDK2 | ENSG00000123374.11 | 32246 | -0.307 | -0.4583 | Yes | ||
| 67 | YWHAG | ENSG00000170027.7 | 32308 | -0.310 | -0.4528 | Yes | ||
| 68 | RBX1 | ENSG00000100387.9 | 32483 | -0.316 | -0.4499 | Yes | ||
| 69 | CCND3 | ENSG00000112576.12 | 32532 | -0.317 | -0.4440 | Yes | ||
| 70 | MCM2 | ENSG00000073111.14 | 32546 | -0.318 | -0.4371 | Yes | ||
| 71 | RB1 | ENSG00000139687.15 | 33569 | -0.339 | -0.4544 | Yes | ||
| 72 | CDC20 | ENSG00000117399.14 | 34187 | -0.349 | -0.4616 | Yes | ||
| 73 | MCM6 | ENSG00000076003.5 | 34728 | -0.357 | -0.4667 | Yes | ||
| 74 | STAG2 | ENSG00000101972.18 | 34834 | -0.363 | -0.4611 | Yes | ||
| 75 | CDKN2A | ENSG00000147889.17 | 35138 | -0.371 | -0.4602 | Yes | ||
| 76 | YWHAB | ENSG00000166913.13 | 35457 | -0.386 | -0.4592 | Yes | ||
| 77 | TFDP1 | ENSG00000198176.13 | 35512 | -0.389 | -0.4518 | Yes | ||
| 78 | YWHAZ | ENSG00000164924.18 | 35696 | -0.397 | -0.4473 | Yes | ||
| 79 | PTTG2 | ENSG00000250254.2 | 35781 | -0.401 | -0.4403 | Yes | ||
| 80 | CDC16 | ENSG00000130177.16 | 35800 | -0.402 | -0.4317 | Yes | ||
| 81 | MCM5 | ENSG00000100297.16 | 35805 | -0.402 | -0.4228 | Yes | ||
| 82 | CDC25C | ENSG00000158402.20 | 35954 | -0.403 | -0.4173 | Yes | ||
| 83 | ESPL1 | ENSG00000135476.11 | 36287 | -0.415 | -0.4160 | Yes | ||
| 84 | ORC3 | ENSG00000135336.14 | 36389 | -0.421 | -0.4090 | Yes | ||
| 85 | YWHAE | ENSG00000108953.17 | 36492 | -0.426 | -0.4019 | Yes | ||
| 86 | CCNB2 | ENSG00000157456.8 | 36613 | -0.432 | -0.3952 | Yes | ||
| 87 | PLK1 | ENSG00000166851.15 | 36707 | -0.436 | -0.3876 | Yes | ||
| 88 | ATR | ENSG00000175054.16 | 36761 | -0.438 | -0.3791 | Yes | ||
| 89 | SMC3 | ENSG00000108055.9 | 36921 | -0.447 | -0.3729 | Yes | ||
| 90 | SMC1B | ENSG00000077935.17 | 37069 | -0.456 | -0.3662 | Yes | ||
| 91 | PCNA | ENSG00000132646.11 | 37166 | -0.461 | -0.3582 | Yes | ||
| 92 | ORC1 | ENSG00000085840.13 | 37179 | -0.462 | -0.3480 | Yes | ||
| 93 | MAD1L1 | ENSG00000002822.15 | 37265 | -0.468 | -0.3396 | Yes | ||
| 94 | CCNE2 | ENSG00000175305.17 | 37360 | -0.475 | -0.3312 | Yes | ||
| 95 | PKMYT1 | ENSG00000127564.17 | 37418 | -0.479 | -0.3218 | Yes | ||
| 96 | RBL1 | ENSG00000080839.12 | 37521 | -0.487 | -0.3133 | Yes | ||
| 97 | GSK3B | ENSG00000082701.15 | 37577 | -0.491 | -0.3036 | Yes | ||
| 98 | BUB1 | ENSG00000169679.15 | 37763 | -0.508 | -0.2967 | Yes | ||
| 99 | CDC14B | ENSG00000081377.17 | 37879 | -0.518 | -0.2878 | Yes | ||
| 100 | MAD2L1 | ENSG00000164109.14 | 37893 | -0.520 | -0.2765 | Yes | ||
| 101 | SMC1A | ENSG00000072501.17 | 37922 | -0.522 | -0.2654 | Yes | ||
| 102 | CDK7 | ENSG00000134058.12 | 37950 | -0.524 | -0.2543 | Yes | ||
| 103 | CDC7 | ENSG00000097046.13 | 38052 | -0.531 | -0.2448 | Yes | ||
| 104 | E2F1 | ENSG00000101412.13 | 38085 | -0.533 | -0.2335 | Yes | ||
| 105 | BUB1B | ENSG00000156970.12 | 38129 | -0.537 | -0.2225 | Yes | ||
| 106 | CCNB1 | ENSG00000134057.15 | 38169 | -0.540 | -0.2113 | Yes | ||
| 107 | TTK | ENSG00000112742.10 | 38213 | -0.544 | -0.2001 | Yes | ||
| 108 | CHEK1 | ENSG00000149554.13 | 38279 | -0.549 | -0.1894 | Yes | ||
| 109 | CCND1 | ENSG00000110092.3 | 38307 | -0.552 | -0.1776 | Yes | ||
| 110 | CDC45 | ENSG00000093009.10 | 38822 | -0.583 | -0.1770 | Yes | ||
| 111 | ORC6 | ENSG00000091651.9 | 38888 | -0.589 | -0.1654 | Yes | ||
| 112 | MYC | ENSG00000136997.19 | 39003 | -0.602 | -0.1546 | Yes | ||
| 113 | CDK1 | ENSG00000170312.16 | 39453 | -0.632 | -0.1513 | Yes | ||
| 114 | WEE1 | ENSG00000166483.11 | 40002 | -0.714 | -0.1486 | Yes | ||
| 115 | GADD45B | ENSG00000099860.9 | 40019 | -0.716 | -0.1329 | Yes | ||
| 116 | CDC6 | ENSG00000094804.12 | 40060 | -0.723 | -0.1176 | Yes | ||
| 117 | CDKN2D | ENSG00000129355.7 | 40158 | -0.743 | -0.1033 | Yes | ||
| 118 | SKP2 | ENSG00000145604.16 | 40211 | -0.755 | -0.0875 | Yes | ||
| 119 | CDC25A | ENSG00000164045.12 | 40263 | -0.768 | -0.0715 | Yes | ||
| 120 | E2F3 | ENSG00000112242.15 | 40268 | -0.769 | -0.0543 | Yes | ||
| 121 | YWHAH | ENSG00000128245.15 | 40295 | -0.774 | -0.0375 | Yes | ||
| 122 | CDK6 | ENSG00000105810.9 | 40555 | -0.858 | -0.0245 | Yes | ||
| 123 | MCM4 | ENSG00000104738.18 | 40591 | -0.872 | -0.0058 | Yes | ||
| 124 | CCNA2 | ENSG00000145386.10 | 40598 | -0.874 | 0.0137 | Yes |