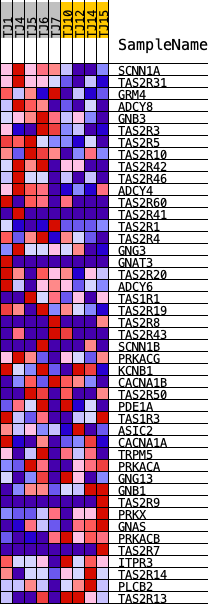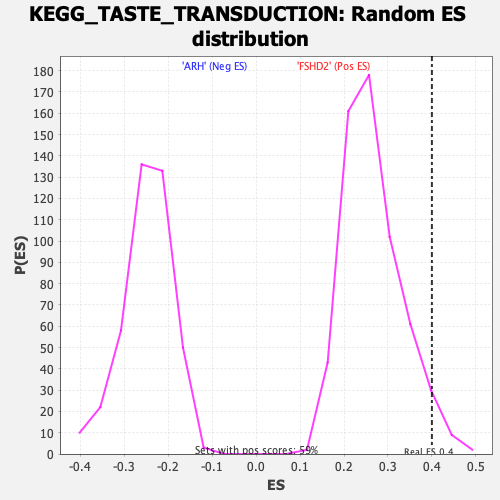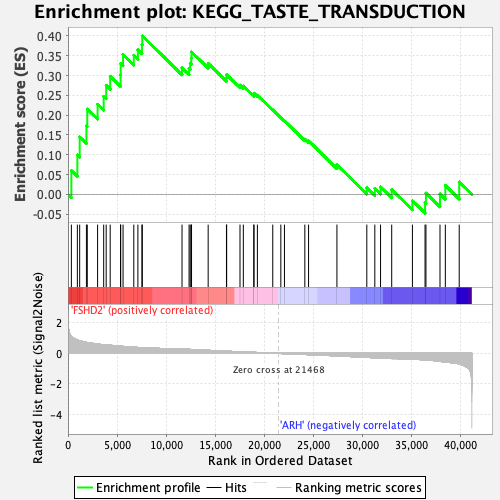
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | FSHD2 |
| GeneSet | KEGG_TASTE_TRANSDUCTION |
| Enrichment Score (ES) | 0.4000573 |
| Normalized Enrichment Score (NES) | 1.5136575 |
| Nominal p-value | 0.027210884 |
| FDR q-value | 0.238648 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SCNN1A | ENSG00000111319.13 | 344 | 1.095 | 0.0599 | Yes | ||
| 2 | TAS2R31 | ENSG00000256436.1 | 954 | 0.870 | 0.0994 | Yes | ||
| 3 | GRM4 | ENSG00000124493.13 | 1190 | 0.820 | 0.1448 | Yes | ||
| 4 | ADCY8 | ENSG00000155897.10 | 1877 | 0.721 | 0.1731 | Yes | ||
| 5 | GNB3 | ENSG00000111664.10 | 1964 | 0.709 | 0.2153 | Yes | ||
| 6 | TAS2R3 | ENSG00000127362.2 | 3017 | 0.605 | 0.2274 | Yes | ||
| 7 | TAS2R5 | ENSG00000127366.5 | 3637 | 0.562 | 0.2475 | Yes | ||
| 8 | TAS2R10 | ENSG00000121318.2 | 3892 | 0.545 | 0.2753 | Yes | ||
| 9 | TAS2R42 | ENSG00000186136.1 | 4300 | 0.524 | 0.2980 | Yes | ||
| 10 | TAS2R46 | ENSG00000226761.3 | 5356 | 0.468 | 0.3016 | Yes | ||
| 11 | ADCY4 | ENSG00000129467.13 | 5370 | 0.467 | 0.3304 | Yes | ||
| 12 | TAS2R60 | ENSG00000185899.1 | 5612 | 0.453 | 0.3528 | Yes | ||
| 13 | TAS2R41 | ENSG00000221855.1 | 6706 | 0.400 | 0.3512 | Yes | ||
| 14 | TAS2R1 | ENSG00000169777.6 | 7124 | 0.384 | 0.3650 | Yes | ||
| 15 | TAS2R4 | ENSG00000127364.3 | 7536 | 0.369 | 0.3780 | Yes | ||
| 16 | GNG3 | ENSG00000162188.6 | 7573 | 0.368 | 0.4001 | Yes | ||
| 17 | GNAT3 | ENSG00000214415.4 | 11622 | 0.285 | 0.3194 | No | ||
| 18 | TAS2R20 | ENSG00000255837.1 | 12341 | 0.259 | 0.3181 | No | ||
| 19 | ADCY6 | ENSG00000174233.11 | 12456 | 0.256 | 0.3313 | No | ||
| 20 | TAS1R1 | ENSG00000173662.21 | 12567 | 0.252 | 0.3443 | No | ||
| 21 | TAS2R19 | ENSG00000212124.2 | 12592 | 0.252 | 0.3595 | No | ||
| 22 | TAS2R8 | ENSG00000121314.2 | 14287 | 0.197 | 0.3306 | No | ||
| 23 | TAS2R43 | ENSG00000255374.3 | 16152 | 0.141 | 0.2940 | No | ||
| 24 | SCNN1B | ENSG00000168447.11 | 16172 | 0.141 | 0.3023 | No | ||
| 25 | PRKACG | ENSG00000165059.7 | 17532 | 0.103 | 0.2757 | No | ||
| 26 | KCNB1 | ENSG00000158445.10 | 17871 | 0.094 | 0.2734 | No | ||
| 27 | CACNA1B | ENSG00000148408.13 | 18944 | 0.067 | 0.2514 | No | ||
| 28 | TAS2R50 | ENSG00000212126.3 | 18954 | 0.066 | 0.2554 | No | ||
| 29 | PDE1A | ENSG00000115252.18 | 19305 | 0.056 | 0.2504 | No | ||
| 30 | TAS1R3 | ENSG00000169962.5 | 20872 | 0.016 | 0.2133 | No | ||
| 31 | ASIC2 | ENSG00000108684.14 | 21694 | -0.006 | 0.1937 | No | ||
| 32 | CACNA1A | ENSG00000141837.20 | 22062 | -0.016 | 0.1857 | No | ||
| 33 | TRPM5 | ENSG00000070985.13 | 24141 | -0.068 | 0.1394 | No | ||
| 34 | PRKACA | ENSG00000072062.14 | 24521 | -0.077 | 0.1350 | No | ||
| 35 | GNG13 | ENSG00000127588.5 | 27412 | -0.161 | 0.0748 | No | ||
| 36 | GNB1 | ENSG00000078369.18 | 30458 | -0.259 | 0.0169 | No | ||
| 37 | TAS2R9 | ENSG00000121381.4 | 31274 | -0.285 | 0.0149 | No | ||
| 38 | PRKX | ENSG00000183943.6 | 31858 | -0.292 | 0.0189 | No | ||
| 39 | GNAS | ENSG00000087460.25 | 32996 | -0.336 | 0.0122 | No | ||
| 40 | PRKACB | ENSG00000142875.19 | 35106 | -0.369 | -0.0161 | No | ||
| 41 | TAS2R7 | ENSG00000121377.2 | 36394 | -0.421 | -0.0211 | No | ||
| 42 | ITPR3 | ENSG00000096433.11 | 36486 | -0.426 | 0.0032 | No | ||
| 43 | TAS2R14 | ENSG00000212127.5 | 37918 | -0.522 | 0.0010 | No | ||
| 44 | PLCB2 | ENSG00000137841.12 | 38462 | -0.558 | 0.0226 | No | ||
| 45 | TAS2R13 | ENSG00000212128.2 | 39877 | -0.690 | 0.0312 | No |