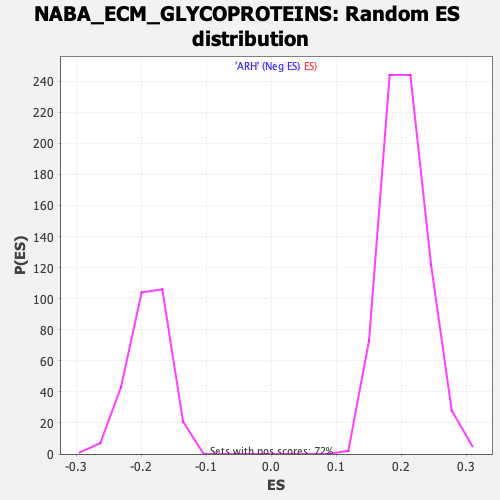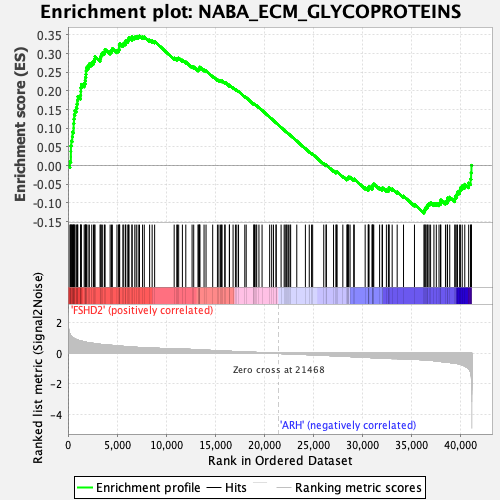
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | FSHD2 |
| GeneSet | NABA_ECM_GLYCOPROTEINS |
| Enrichment Score (ES) | 0.34757543 |
| Normalized Enrichment Score (NES) | 1.6885211 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.16407658 |
| FWER p-Value | 0.994 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | NELL2 | ENSG00000184613.10 | 198 | 1.241 | 0.0112 | Yes | ||
| 2 | LAMA1 | ENSG00000101680.15 | 292 | 1.136 | 0.0236 | Yes | ||
| 3 | TECTA | ENSG00000109927.10 | 293 | 1.135 | 0.0383 | Yes | ||
| 4 | BGLAP | ENSG00000242252.2 | 299 | 1.133 | 0.0528 | Yes | ||
| 5 | VWCE | ENSG00000167992.13 | 337 | 1.099 | 0.0661 | Yes | ||
| 6 | VWA5A | ENSG00000110002.16 | 433 | 1.042 | 0.0773 | Yes | ||
| 7 | MFAP4 | ENSG00000166482.11 | 456 | 1.033 | 0.0901 | Yes | ||
| 8 | GLDN | ENSG00000186417.14 | 573 | 0.990 | 0.1001 | Yes | ||
| 9 | EFEMP1 | ENSG00000115380.20 | 575 | 0.988 | 0.1128 | Yes | ||
| 10 | RSPO1 | ENSG00000169218.14 | 606 | 0.975 | 0.1247 | Yes | ||
| 11 | SLIT1 | ENSG00000187122.17 | 635 | 0.962 | 0.1365 | Yes | ||
| 12 | FBLN1 | ENSG00000077942.19 | 705 | 0.938 | 0.1469 | Yes | ||
| 13 | NTN4 | ENSG00000074527.12 | 840 | 0.901 | 0.1553 | Yes | ||
| 14 | MGP | ENSG00000111341.10 | 913 | 0.879 | 0.1649 | Yes | ||
| 15 | TNN | ENSG00000120332.16 | 965 | 0.867 | 0.1749 | Yes | ||
| 16 | NTN5 | ENSG00000142233.11 | 1001 | 0.859 | 0.1851 | Yes | ||
| 17 | TSPEAR | ENSG00000175894.18 | 1291 | 0.804 | 0.1885 | Yes | ||
| 18 | RELN | ENSG00000189056.14 | 1303 | 0.802 | 0.1986 | Yes | ||
| 19 | PCOLCE2 | ENSG00000163710.9 | 1312 | 0.800 | 0.2087 | Yes | ||
| 20 | TINAG | ENSG00000137251.16 | 1358 | 0.793 | 0.2179 | Yes | ||
| 21 | VWA5B2 | ENSG00000145198.14 | 1649 | 0.754 | 0.2205 | Yes | ||
| 22 | EDIL3 | ENSG00000164176.13 | 1748 | 0.740 | 0.2277 | Yes | ||
| 23 | NTNG1 | ENSG00000162631.18 | 1784 | 0.736 | 0.2364 | Yes | ||
| 24 | LTBP4 | ENSG00000090006.17 | 1825 | 0.729 | 0.2448 | Yes | ||
| 25 | EMID1 | ENSG00000186998.16 | 1845 | 0.725 | 0.2537 | Yes | ||
| 26 | SNED1 | ENSG00000162804.14 | 1875 | 0.721 | 0.2624 | Yes | ||
| 27 | LAMB4 | ENSG00000091128.13 | 2043 | 0.699 | 0.2673 | Yes | ||
| 28 | LTBP1 | ENSG00000049323.16 | 2175 | 0.686 | 0.2730 | Yes | ||
| 29 | IGFBP3 | ENSG00000146674.15 | 2420 | 0.661 | 0.2756 | Yes | ||
| 30 | SMOC2 | ENSG00000112562.18 | 2598 | 0.644 | 0.2796 | Yes | ||
| 31 | NTNG2 | ENSG00000196358.11 | 2685 | 0.635 | 0.2857 | Yes | ||
| 32 | CRISPLD1 | ENSG00000121005.9 | 2756 | 0.628 | 0.2921 | Yes | ||
| 33 | RSPO3 | ENSG00000146374.14 | 3267 | 0.586 | 0.2872 | Yes | ||
| 34 | SBSPON | ENSG00000164764.11 | 3341 | 0.581 | 0.2930 | Yes | ||
| 35 | LAMB1 | ENSG00000091136.14 | 3394 | 0.576 | 0.2992 | Yes | ||
| 36 | SPON1 | ENSG00000262655.4 | 3535 | 0.565 | 0.3030 | Yes | ||
| 37 | IGFBP4 | ENSG00000141753.7 | 3723 | 0.556 | 0.3057 | Yes | ||
| 38 | MMRN2 | ENSG00000173269.14 | 3779 | 0.551 | 0.3115 | Yes | ||
| 39 | CRELD1 | ENSG00000163703.17 | 4296 | 0.524 | 0.3056 | Yes | ||
| 40 | LGI3 | ENSG00000168481.9 | 4431 | 0.519 | 0.3091 | Yes | ||
| 41 | SSPO | ENSG00000197558.12 | 4521 | 0.518 | 0.3136 | Yes | ||
| 42 | LAMC3 | ENSG00000050555.18 | 4992 | 0.492 | 0.3085 | Yes | ||
| 43 | LAMA3 | ENSG00000053747.16 | 5156 | 0.480 | 0.3107 | Yes | ||
| 44 | CCN5 | ENSG00000064205.10 | 5220 | 0.476 | 0.3153 | Yes | ||
| 45 | VWF | ENSG00000110799.13 | 5222 | 0.476 | 0.3215 | Yes | ||
| 46 | ANOS1 | ENSG00000011201.12 | 5269 | 0.473 | 0.3265 | Yes | ||
| 47 | NID2 | ENSG00000087303.18 | 5592 | 0.454 | 0.3245 | Yes | ||
| 48 | LAMA5 | ENSG00000130702.15 | 5685 | 0.448 | 0.3280 | Yes | ||
| 49 | EYS | ENSG00000188107.15 | 5877 | 0.440 | 0.3291 | Yes | ||
| 50 | PCOLCE | ENSG00000106333.13 | 5878 | 0.440 | 0.3347 | Yes | ||
| 51 | COLQ | ENSG00000206561.13 | 6095 | 0.428 | 0.3350 | Yes | ||
| 52 | SPARCL1 | ENSG00000152583.12 | 6166 | 0.425 | 0.3388 | Yes | ||
| 53 | COCH | ENSG00000100473.18 | 6222 | 0.421 | 0.3429 | Yes | ||
| 54 | LTBP3 | ENSG00000168056.16 | 6522 | 0.407 | 0.3409 | Yes | ||
| 55 | LGI4 | ENSG00000153902.14 | 6529 | 0.407 | 0.3460 | Yes | ||
| 56 | NTN3 | ENSG00000162068.2 | 6808 | 0.396 | 0.3443 | Yes | ||
| 57 | CCN3 | ENSG00000136999.5 | 6958 | 0.391 | 0.3458 | Yes | ||
| 58 | RSPO2 | ENSG00000147655.12 | 7153 | 0.382 | 0.3460 | Yes | ||
| 59 | FGG | ENSG00000171557.17 | 7287 | 0.376 | 0.3476 | Yes | ||
| 60 | VWA7 | ENSG00000204396.11 | 7599 | 0.367 | 0.3447 | No | ||
| 61 | ADIPOQ | ENSG00000181092.10 | 7776 | 0.363 | 0.3451 | No | ||
| 62 | ZP2 | ENSG00000103310.10 | 8318 | 0.352 | 0.3365 | No | ||
| 63 | MEPE | ENSG00000152595.16 | 8577 | 0.348 | 0.3347 | No | ||
| 64 | VWA2 | ENSG00000165816.13 | 8831 | 0.342 | 0.3329 | No | ||
| 65 | FNDC1 | ENSG00000164694.17 | 10822 | 0.298 | 0.2882 | No | ||
| 66 | PXDNL | ENSG00000147485.13 | 11090 | 0.288 | 0.2854 | No | ||
| 67 | DSPP | ENSG00000152591.15 | 11156 | 0.285 | 0.2875 | No | ||
| 68 | TNR | ENSG00000116147.17 | 11282 | 0.285 | 0.2882 | No | ||
| 69 | CTHRC1 | ENSG00000164932.13 | 11661 | 0.284 | 0.2826 | No | ||
| 70 | VWA3A | ENSG00000175267.15 | 12007 | 0.271 | 0.2777 | No | ||
| 71 | NID1 | ENSG00000116962.15 | 12648 | 0.250 | 0.2653 | No | ||
| 72 | USH2A | ENSG00000042781.13 | 12814 | 0.244 | 0.2644 | No | ||
| 73 | CCN6 | ENSG00000112761.20 | 13264 | 0.230 | 0.2564 | No | ||
| 74 | VTN | ENSG00000109072.14 | 13279 | 0.229 | 0.2591 | No | ||
| 75 | IGFBPL1 | ENSG00000137142.5 | 13388 | 0.226 | 0.2593 | No | ||
| 76 | IGFBP2 | ENSG00000115457.10 | 13426 | 0.224 | 0.2613 | No | ||
| 77 | MATN4 | ENSG00000124159.15 | 13437 | 0.223 | 0.2640 | No | ||
| 78 | MATN1 | ENSG00000162510.6 | 13868 | 0.210 | 0.2562 | No | ||
| 79 | TNFAIP6 | ENSG00000123610.5 | 14070 | 0.204 | 0.2539 | No | ||
| 80 | LAMA2 | ENSG00000196569.12 | 14748 | 0.182 | 0.2398 | No | ||
| 81 | IGFALS | ENSG00000099769.5 | 15231 | 0.167 | 0.2302 | No | ||
| 82 | VWA3B | ENSG00000168658.19 | 15379 | 0.162 | 0.2287 | No | ||
| 83 | SPON2 | ENSG00000159674.12 | 15536 | 0.158 | 0.2269 | No | ||
| 84 | HMCN2 | ENSG00000148357.16 | 15639 | 0.155 | 0.2264 | No | ||
| 85 | MFGE8 | ENSG00000140545.15 | 15705 | 0.153 | 0.2268 | No | ||
| 86 | EMILIN3 | ENSG00000183798.5 | 15977 | 0.146 | 0.2221 | No | ||
| 87 | IGSF10 | ENSG00000152580.8 | 16016 | 0.145 | 0.2230 | No | ||
| 88 | EMILIN1 | ENSG00000138080.14 | 16451 | 0.132 | 0.2142 | No | ||
| 89 | LGI1 | ENSG00000108231.13 | 16455 | 0.132 | 0.2158 | No | ||
| 90 | SMOC1 | ENSG00000198732.10 | 16826 | 0.122 | 0.2083 | No | ||
| 91 | NTN1 | ENSG00000065320.9 | 17067 | 0.116 | 0.2040 | No | ||
| 92 | SVEP1 | ENSG00000165124.18 | 17205 | 0.112 | 0.2021 | No | ||
| 93 | LAMB3 | ENSG00000196878.15 | 17382 | 0.108 | 0.1992 | No | ||
| 94 | AGRN | ENSG00000188157.15 | 18021 | 0.090 | 0.1848 | No | ||
| 95 | FBN1 | ENSG00000166147.13 | 18163 | 0.087 | 0.1825 | No | ||
| 96 | SLIT3 | ENSG00000184347.14 | 18947 | 0.067 | 0.1642 | No | ||
| 97 | KCP | ENSG00000135253.13 | 18949 | 0.066 | 0.1650 | No | ||
| 98 | ZP1 | ENSG00000149506.11 | 19012 | 0.065 | 0.1644 | No | ||
| 99 | CRELD2 | ENSG00000184164.14 | 19149 | 0.061 | 0.1618 | No | ||
| 100 | CILP2 | ENSG00000160161.9 | 19234 | 0.059 | 0.1605 | No | ||
| 101 | THBS2 | ENSG00000186340.15 | 19463 | 0.052 | 0.1557 | No | ||
| 102 | MFAP2 | ENSG00000117122.14 | 19781 | 0.044 | 0.1485 | No | ||
| 103 | VIT | ENSG00000205221.12 | 20556 | 0.023 | 0.1299 | No | ||
| 104 | NELL1 | ENSG00000165973.19 | 20760 | 0.019 | 0.1252 | No | ||
| 105 | FGA | ENSG00000171560.16 | 20943 | 0.014 | 0.1209 | No | ||
| 106 | AMELX | ENSG00000125363.14 | 21206 | 0.007 | 0.1146 | No | ||
| 107 | FBLN2 | ENSG00000163520.14 | 21217 | 0.006 | 0.1144 | No | ||
| 108 | CRISPLD2 | ENSG00000103196.12 | 21721 | -0.007 | 0.1023 | No | ||
| 109 | THBS1 | ENSG00000137801.11 | 22033 | -0.015 | 0.0949 | No | ||
| 110 | INTS14 | ENSG00000138614.15 | 22139 | -0.016 | 0.0925 | No | ||
| 111 | CCN2 | ENSG00000118523.6 | 22253 | -0.019 | 0.0900 | No | ||
| 112 | TSKU | ENSG00000182704.8 | 22329 | -0.022 | 0.0885 | No | ||
| 113 | FNDC8 | ENSG00000073598.6 | 22463 | -0.026 | 0.0855 | No | ||
| 114 | PXDN | ENSG00000130508.11 | 22625 | -0.029 | 0.0820 | No | ||
| 115 | PAPLN | ENSG00000100767.16 | 22695 | -0.031 | 0.0807 | No | ||
| 116 | EMILIN2 | ENSG00000132205.11 | 23319 | -0.047 | 0.0661 | No | ||
| 117 | VWA1 | ENSG00000179403.12 | 24198 | -0.070 | 0.0456 | No | ||
| 118 | OIT3 | ENSG00000138315.13 | 24592 | -0.080 | 0.0370 | No | ||
| 119 | LAMA4 | ENSG00000112769.20 | 24827 | -0.086 | 0.0324 | No | ||
| 120 | EFEMP2 | ENSG00000172638.13 | 24938 | -0.090 | 0.0309 | No | ||
| 121 | DMP1 | ENSG00000152592.14 | 26068 | -0.123 | 0.0049 | No | ||
| 122 | FBN2 | ENSG00000138829.12 | 26300 | -0.129 | 0.0010 | No | ||
| 123 | FNDC7 | ENSG00000143107.10 | 26336 | -0.130 | 0.0018 | No | ||
| 124 | IGFBP5 | ENSG00000115461.5 | 27066 | -0.151 | -0.0140 | No | ||
| 125 | TNXB | ENSG00000168477.19 | 27304 | -0.158 | -0.0178 | No | ||
| 126 | THBS3 | ENSG00000169231.13 | 27375 | -0.160 | -0.0174 | No | ||
| 127 | MATN2 | ENSG00000132561.14 | 27429 | -0.162 | -0.0166 | No | ||
| 128 | SPARC | ENSG00000113140.11 | 28014 | -0.181 | -0.0285 | No | ||
| 129 | MFAP3 | ENSG00000037749.12 | 28424 | -0.194 | -0.0360 | No | ||
| 130 | ECM2 | ENSG00000106823.12 | 28489 | -0.196 | -0.0350 | No | ||
| 131 | ZP3 | ENSG00000188372.15 | 28519 | -0.197 | -0.0332 | No | ||
| 132 | IGFBP1 | ENSG00000146678.10 | 28557 | -0.198 | -0.0315 | No | ||
| 133 | LTBP2 | ENSG00000119681.12 | 28615 | -0.200 | -0.0303 | No | ||
| 134 | LAMC1 | ENSG00000135862.6 | 28753 | -0.203 | -0.0311 | No | ||
| 135 | MFAP1 | ENSG00000140259.7 | 29124 | -0.216 | -0.0373 | No | ||
| 136 | POMZP3 | ENSG00000146707.14 | 29178 | -0.217 | -0.0358 | No | ||
| 137 | CCN1 | ENSG00000142871.17 | 30290 | -0.253 | -0.0596 | No | ||
| 138 | MXRA5 | ENSG00000101825.8 | 30601 | -0.263 | -0.0638 | No | ||
| 139 | INTS6L | ENSG00000165359.15 | 30623 | -0.263 | -0.0609 | No | ||
| 140 | RSPO4 | ENSG00000101282.9 | 30644 | -0.264 | -0.0580 | No | ||
| 141 | EGFLAM | ENSG00000164318.18 | 30681 | -0.265 | -0.0554 | No | ||
| 142 | COMP | ENSG00000105664.11 | 31006 | -0.277 | -0.0598 | No | ||
| 143 | FBN3 | ENSG00000142449.13 | 31009 | -0.277 | -0.0562 | No | ||
| 144 | SRPX | ENSG00000101955.15 | 31035 | -0.278 | -0.0532 | No | ||
| 145 | FBLN5 | ENSG00000140092.14 | 31116 | -0.281 | -0.0515 | No | ||
| 146 | LAMC2 | ENSG00000058085.15 | 31167 | -0.284 | -0.0491 | No | ||
| 147 | THSD4 | ENSG00000187720.14 | 31761 | -0.288 | -0.0599 | No | ||
| 148 | LGI2 | ENSG00000153012.12 | 32016 | -0.297 | -0.0622 | No | ||
| 149 | MFAP5 | ENSG00000197614.11 | 32055 | -0.298 | -0.0593 | No | ||
| 150 | DMBT1 | ENSG00000187908.19 | 32453 | -0.315 | -0.0649 | No | ||
| 151 | FGL2 | ENSG00000127951.7 | 32655 | -0.323 | -0.0656 | No | ||
| 152 | LAMB2 | ENSG00000172037.14 | 32705 | -0.324 | -0.0626 | No | ||
| 153 | VWA5B1 | ENSG00000158816.15 | 32729 | -0.325 | -0.0590 | No | ||
| 154 | FGB | ENSG00000171564.11 | 33043 | -0.336 | -0.0623 | No | ||
| 155 | ABI3BP | ENSG00000154175.17 | 33553 | -0.338 | -0.0703 | No | ||
| 156 | THBS4 | ENSG00000113296.14 | 34200 | -0.350 | -0.0816 | No | ||
| 157 | ELN | ENSG00000049540.17 | 35316 | -0.380 | -0.1039 | No | ||
| 158 | TINAGL1 | ENSG00000142910.16 | 36281 | -0.415 | -0.1220 | No | ||
| 159 | CCN4 | ENSG00000104415.14 | 36365 | -0.419 | -0.1186 | No | ||
| 160 | IGFBP7 | ENSG00000163453.11 | 36410 | -0.421 | -0.1143 | No | ||
| 161 | TNC | ENSG00000041982.16 | 36501 | -0.427 | -0.1109 | No | ||
| 162 | AEBP1 | ENSG00000106624.11 | 36626 | -0.432 | -0.1084 | No | ||
| 163 | FBLN7 | ENSG00000144152.13 | 36684 | -0.435 | -0.1041 | No | ||
| 164 | POSTN | ENSG00000133110.15 | 36812 | -0.441 | -0.1015 | No | ||
| 165 | CILP | ENSG00000138615.6 | 36960 | -0.449 | -0.0993 | No | ||
| 166 | SRPX2 | ENSG00000102359.7 | 37299 | -0.471 | -0.1015 | No | ||
| 167 | CRIM1 | ENSG00000150938.10 | 37543 | -0.488 | -0.1011 | No | ||
| 168 | TGFBI | ENSG00000120708.17 | 37818 | -0.514 | -0.1011 | No | ||
| 169 | ZPLD1 | ENSG00000170044.8 | 37968 | -0.525 | -0.0980 | No | ||
| 170 | SLIT2 | ENSG00000145147.20 | 38005 | -0.528 | -0.0920 | No | ||
| 171 | OTOG | ENSG00000188162.11 | 38476 | -0.558 | -0.0963 | No | ||
| 172 | TECTB | ENSG00000119913.6 | 38653 | -0.566 | -0.0933 | No | ||
| 173 | SPP1 | ENSG00000118785.14 | 38686 | -0.569 | -0.0867 | No | ||
| 174 | FRAS1 | ENSG00000138759.19 | 38912 | -0.591 | -0.0845 | No | ||
| 175 | IGFBP6 | ENSG00000167779.9 | 39424 | -0.628 | -0.0889 | No | ||
| 176 | IBSP | ENSG00000029559.7 | 39472 | -0.633 | -0.0818 | No | ||
| 177 | NPNT | ENSG00000168743.12 | 39626 | -0.650 | -0.0772 | No | ||
| 178 | NDNF | ENSG00000173376.14 | 39706 | -0.662 | -0.0705 | No | ||
| 179 | ECM1 | ENSG00000143369.15 | 39921 | -0.699 | -0.0667 | No | ||
| 180 | GAS6 | ENSG00000183087.15 | 39997 | -0.713 | -0.0593 | No | ||
| 181 | DPT | ENSG00000143196.5 | 40183 | -0.749 | -0.0542 | No | ||
| 182 | HMCN1 | ENSG00000143341.12 | 40444 | -0.822 | -0.0499 | No | ||
| 183 | FN1 | ENSG00000115414.19 | 40842 | -0.991 | -0.0468 | No | ||
| 184 | MATN3 | ENSG00000132031.13 | 41030 | -1.191 | -0.0359 | No | ||
| 185 | BMPER | ENSG00000164619.10 | 41089 | -1.386 | -0.0194 | No | ||
| 186 | VWDE | ENSG00000146530.14 | 41124 | -1.636 | 0.0009 | No |