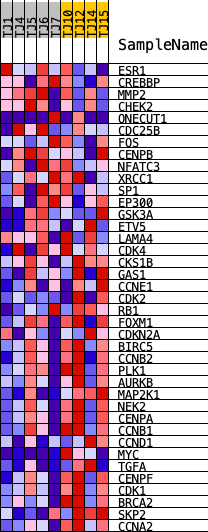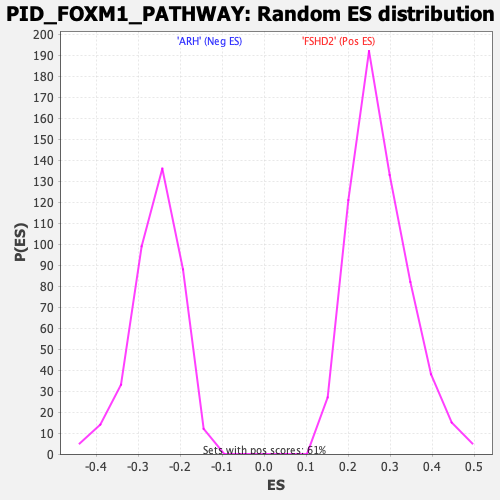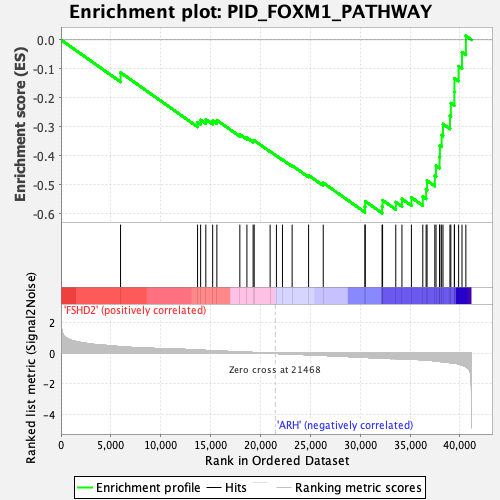
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | ARH |
| GeneSet | PID_FOXM1_PATHWAY |
| Enrichment Score (ES) | -0.59777874 |
| Normalized Enrichment Score (NES) | -2.3221242 |
| Nominal p-value | 0.0 |
| FDR q-value | 1.2446613E-4 |
| FWER p-Value | 0.002 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ESR1 | ENSG00000091831.23 | 5971 | 0.435 | -0.1126 | No | ||
| 2 | CREBBP | ENSG00000005339.14 | 13694 | 0.216 | -0.2842 | No | ||
| 3 | MMP2 | ENSG00000087245.13 | 14001 | 0.206 | -0.2762 | No | ||
| 4 | CHEK2 | ENSG00000183765.22 | 14522 | 0.190 | -0.2746 | No | ||
| 5 | ONECUT1 | ENSG00000169856.9 | 15209 | 0.168 | -0.2787 | No | ||
| 6 | CDC25B | ENSG00000101224.17 | 15629 | 0.156 | -0.2772 | No | ||
| 7 | FOS | ENSG00000170345.10 | 17933 | 0.092 | -0.3263 | No | ||
| 8 | CENPB | ENSG00000125817.8 | 18642 | 0.075 | -0.3378 | No | ||
| 9 | NFATC3 | ENSG00000072736.19 | 19266 | 0.058 | -0.3487 | No | ||
| 10 | XRCC1 | ENSG00000073050.11 | 19372 | 0.054 | -0.3472 | No | ||
| 11 | SP1 | ENSG00000185591.10 | 20970 | 0.013 | -0.3850 | No | ||
| 12 | EP300 | ENSG00000100393.13 | 21602 | -0.004 | -0.4001 | No | ||
| 13 | GSK3A | ENSG00000105723.12 | 22208 | -0.018 | -0.4134 | No | ||
| 14 | ETV5 | ENSG00000244405.8 | 23177 | -0.044 | -0.4336 | No | ||
| 15 | LAMA4 | ENSG00000112769.20 | 24827 | -0.086 | -0.4672 | No | ||
| 16 | CDK4 | ENSG00000135446.17 | 26296 | -0.129 | -0.4933 | No | ||
| 17 | CKS1B | ENSG00000173207.13 | 30473 | -0.259 | -0.5754 | Yes | ||
| 18 | GAS1 | ENSG00000180447.7 | 30508 | -0.260 | -0.5567 | Yes | ||
| 19 | CCNE1 | ENSG00000105173.14 | 32197 | -0.305 | -0.5749 | Yes | ||
| 20 | CDK2 | ENSG00000123374.11 | 32246 | -0.307 | -0.5531 | Yes | ||
| 21 | RB1 | ENSG00000139687.15 | 33569 | -0.339 | -0.5598 | Yes | ||
| 22 | FOXM1 | ENSG00000111206.12 | 34188 | -0.349 | -0.5487 | Yes | ||
| 23 | CDKN2A | ENSG00000147889.17 | 35138 | -0.371 | -0.5439 | Yes | ||
| 24 | BIRC5 | ENSG00000089685.15 | 36279 | -0.415 | -0.5405 | Yes | ||
| 25 | CCNB2 | ENSG00000157456.8 | 36613 | -0.432 | -0.5162 | Yes | ||
| 26 | PLK1 | ENSG00000166851.15 | 36707 | -0.436 | -0.4858 | Yes | ||
| 27 | AURKB | ENSG00000178999.13 | 37485 | -0.484 | -0.4684 | Yes | ||
| 28 | MAP2K1 | ENSG00000169032.9 | 37608 | -0.494 | -0.4343 | Yes | ||
| 29 | NEK2 | ENSG00000117650.13 | 37958 | -0.525 | -0.4035 | Yes | ||
| 30 | CENPA | ENSG00000115163.15 | 37993 | -0.527 | -0.3648 | Yes | ||
| 31 | CCNB1 | ENSG00000134057.15 | 38169 | -0.540 | -0.3285 | Yes | ||
| 32 | CCND1 | ENSG00000110092.3 | 38307 | -0.552 | -0.2905 | Yes | ||
| 33 | MYC | ENSG00000136997.19 | 39003 | -0.602 | -0.2622 | Yes | ||
| 34 | TGFA | ENSG00000163235.16 | 39092 | -0.612 | -0.2185 | Yes | ||
| 35 | CENPF | ENSG00000117724.13 | 39444 | -0.631 | -0.1797 | Yes | ||
| 36 | CDK1 | ENSG00000170312.16 | 39453 | -0.632 | -0.1325 | Yes | ||
| 37 | BRCA2 | ENSG00000139618.15 | 39873 | -0.690 | -0.0909 | Yes | ||
| 38 | SKP2 | ENSG00000145604.16 | 40211 | -0.755 | -0.0425 | Yes | ||
| 39 | CCNA2 | ENSG00000145386.10 | 40598 | -0.874 | 0.0137 | Yes |